BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QBTB.068P22F020924.3.1
(714 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
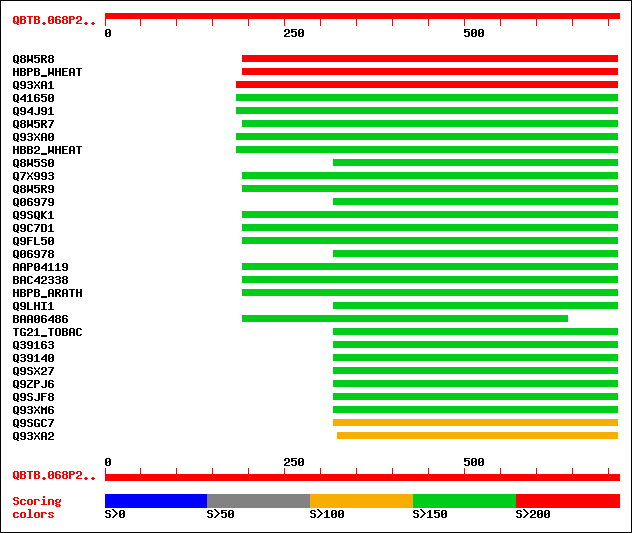
sptr|Q8W5R8|Q8W5R8 bZIP transcription factor. 226 2e-58
sw|P23923|HBPB_WHEAT Transcription factor HBP-1b(c38). 222 3e-57
sptr|Q93XA1|Q93XA1 TGA-type basic leucine zipper protein TGA2.1. 201 1e-50
sptr|Q41650|Q41650 CREB-like protein. 200 1e-50
sptr|Q94J91|Q94J91 Putative transcription factor HBP-1b. 198 7e-50
sptr|Q8W5R7|Q8W5R7 bZIP transcription factor. 196 2e-49
sptr|Q93XA0|Q93XA0 TGA-type basic leucine zipper protein TGA2.2. 194 1e-48
sw|Q41558|HBB2_WHEAT Transcription factor HBP-1b(c1) (Fragment). 191 6e-48
sptr|Q8W5S0|Q8W5S0 bZIP transcription factor. 191 8e-48
sptr|Q7X993|Q7X993 bZIP transcription factor. 191 1e-47
sptr|Q8W5R9|Q8W5R9 bZIP transcription factor. 191 1e-47
sptr|Q06979|Q06979 Ocs-element binding factor 3.2. 190 1e-47
sptr|Q9SQK1|Q9SQK1 bZIP transcription factor. 187 1e-46
sptr|Q9C7D1|Q9C7D1 Transcription factor HBP-1B-like (Transcripti... 187 1e-46
sptr|Q9FL50|Q9FL50 Transcription factor HBP-1b (AT5g06960/MOJ9_13). 184 1e-45
sptr|Q06978|Q06978 Ocs-element binding factor 3.1. 184 1e-45
sptrnew|AAP04119|AAP04119 Putative bZIP transcription factor, HB... 183 2e-45
sptrnew|BAC42338|BAC42338 Putative bZip transcription factor Atb... 183 2e-45
sw|P43273|HBPB_ARATH Transcription factor HBP-1b homolog. 183 2e-45
sptr|Q9LHI1|Q9LHI1 Leucine zipper transcription factor HBP-1b. 180 2e-44
sptrnew|BAA06486|BAA06486 Transcription factor HBP-1b(c38) (Frag... 179 3e-44
sw|O24160|TG21_TOBAC TGACG-sequence specific DNA-binding protein... 179 4e-44
sptr|Q39163|Q39163 Ocs-element binding factor 5 (Fragment). 178 7e-44
sptr|Q39140|Q39140 Leucine zipper. 177 2e-43
sptr|Q9SX27|Q9SX27 F24J5.12 protein (Putative bZIP transcription... 156 2e-37
sptr|Q9ZPJ6|Q9ZPJ6 Transcription factor PERIANTHIA. 155 7e-37
sptr|Q9SJF8|Q9SJF8 T27G7.2. 151 7e-36
sptr|Q93XM6|Q93XM6 bZIP transcription factor. 151 7e-36
sptr|Q9SGC7|Q9SGC7 T23G18.19. 141 1e-32
sptr|Q93XA2|Q93XA2 TGA-type basic leucine zipper protein TGA1.1. 140 2e-32
>sptr|Q8W5R8|Q8W5R8 bZIP transcription factor.
Length = 329
Score = 226 bits (576), Expect = 2e-58
Identities = 119/174 (68%), Positives = 128/174 (73%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNREAARKS 370
MAD SPR N+ LE G+ L H DQKTLRRLAQNREAARKS
Sbjct: 1 MADMSPRTDTSTDDTDDNHMLEPGQLALAAASDSDRSKDKHEDQKTLRRLAQNREAARKS 60
Query: 371 RLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYARW 550
RLRKKAYVQQLENSRLKLT GIFISSSVDQ+HSMSGNGA AFDMEYARW
Sbjct: 61 RLRKKAYVQQLENSRLKLTQLEQELQRARQQGIFISSSVDQTHSMSGNGALAFDMEYARW 120
Query: 551 LEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
LEEHNRQI+ELR+ V+AHA D +LR VVDKIMSHY+EIF+ KGNAAKADVFHVL
Sbjct: 121 LEEHNRQINELRSAVNAHAGDNELRGVVDKIMSHYEEIFKQKGNAAKADVFHVL 174
>sw|P23923|HBPB_WHEAT Transcription factor HBP-1b(c38).
Length = 332
Score = 222 bits (566), Expect = 3e-57
Identities = 119/175 (68%), Positives = 128/175 (73%), Gaps = 1/175 (0%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXX-HGDQKTLRRLAQNREAARK 367
MA+ASPR N LE G L +GDQKT+RRLAQNREAARK
Sbjct: 1 MAEASPRTETSTDDTDENLMLEPGNAALAVVSDSSDRSRDKNGDQKTMRRLAQNREAARK 60
Query: 368 SRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYAR 547
SRLRKKAYVQQLENSRLKLT GIFISSS DQSHSMSGNGA AFD EYAR
Sbjct: 61 SRLRKKAYVQQLENSRLKLTQLEQELQRARQQGIFISSSADQSHSMSGNGALAFDTEYAR 120
Query: 548 WLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
WLEEHNRQ++ELRA V+AHA DT+LRSVV+KIMSHYDEIF+ KGNAAKADVFHVL
Sbjct: 121 WLEEHNRQVNELRAAVNAHAGDTELRSVVEKIMSHYDEIFKQKGNAAKADVFHVL 175
>sptr|Q93XA1|Q93XA1 TGA-type basic leucine zipper protein TGA2.1.
Length = 467
Score = 201 bits (510), Expect = 1e-50
Identities = 108/178 (60%), Positives = 122/178 (68%), Gaps = 1/178 (0%)
Frame = +2
Query: 182 EVTMADASPRXXXXXXXXXXNNGLELGRG-GLXXXXXXXXXXXXHGDQKTLRRLAQNREA 358
E MADASPR + R L DQKTLRRLAQNREA
Sbjct: 134 ESAMADASPRTDISTDVDTDDKNQPFDRNQSLAAVSDSSDRSKDKSDQKTLRRLAQNREA 193
Query: 359 ARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDME 538
ARKSRLRKKAYVQQLE+SRLKLT GIFISSS DQ+H++SGNGA FD E
Sbjct: 194 ARKSRLRKKAYVQQLESSRLKLTQLEQELQRARQQGIFISSSGDQAHTLSGNGAMQFDAE 253
Query: 539 YARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
YARWLEE NRQI+ELRA V++HASDT+LR +VD I++HYDEIFR+KG AAKADVFH+L
Sbjct: 254 YARWLEEQNRQINELRAAVNSHASDTELRMIVDGILAHYDEIFRMKGVAAKADVFHLL 311
>sptr|Q41650|Q41650 CREB-like protein.
Length = 433
Score = 200 bits (509), Expect = 1e-50
Identities = 108/179 (60%), Positives = 124/179 (69%), Gaps = 2/179 (1%)
Frame = +2
Query: 182 EVTMADASPRXXXXXXXXXX--NNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNRE 355
E TMADASPR N + + + DQKTLRRLAQNRE
Sbjct: 132 EFTMADASPRTDISTDVDTDDKNQRFDTNQSLVPVGSDSSDRSKDKSDQKTLRRLAQNRE 191
Query: 356 AARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDM 535
AARKSRLRKKAYVQQLE+SRLKLT G+FISSS +Q+HS+SGNGA FD
Sbjct: 192 AARKSRLRKKAYVQQLESSRLKLTQLEQELQRARQQGVFISSSGEQTHSLSGNGAMQFDA 251
Query: 536 EYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
EYARWLEE NRQI+ELRA V++HASDT+LR +VD I++HYDEIFRLKG AAKADVFH+L
Sbjct: 252 EYARWLEEQNRQINELRAAVNSHASDTELRMIVDGILAHYDEIFRLKGVAAKADVFHLL 310
>sptr|Q94J91|Q94J91 Putative transcription factor HBP-1b.
Length = 472
Score = 198 bits (503), Expect = 7e-50
Identities = 108/179 (60%), Positives = 120/179 (67%), Gaps = 2/179 (1%)
Frame = +2
Query: 182 EVTMADASPRXXXXXX--XXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNRE 355
E T+AD SPR N E G+ D KTLRRLAQNRE
Sbjct: 138 ESTIADTSPRTDTSTDPDTDERNQMFEQGQLAAPTASDSSDRSKDKLDHKTLRRLAQNRE 197
Query: 356 AARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDM 535
AARKSRLRKKAY+Q LE+SRLKLT GIFIS+S DQSHS SGNGA AFDM
Sbjct: 198 AARKSRLRKKAYIQNLESSRLKLTQIEQELQRARQQGIFISTSSDQSHSASGNGALAFDM 257
Query: 536 EYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
EYARWLEEHN+ I+ELRA V+AHA D DL+S VD IM+HY+EIF+LKG AAKADVFHVL
Sbjct: 258 EYARWLEEHNKHINELRAAVNAHAGDNDLKSTVDSIMAHYNEIFKLKGVAAKADVFHVL 316
>sptr|Q8W5R7|Q8W5R7 bZIP transcription factor.
Length = 332
Score = 196 bits (499), Expect = 2e-49
Identities = 107/176 (60%), Positives = 118/176 (67%), Gaps = 2/176 (1%)
Frame = +2
Query: 191 MADASPRXXXXXX--XXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNREAAR 364
MAD SPR N E G+ D KTLRRLAQNREAAR
Sbjct: 1 MADTSPRTDTSTDPDTDERNQMFEQGQLAAPTASDSSDRSKDKLDHKTLRRLAQNREAAR 60
Query: 365 KSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYA 544
KSRLRKKAY+Q LE+SRLKLT GIFIS+S DQSHS SGNGA AFDMEYA
Sbjct: 61 KSRLRKKAYIQNLESSRLKLTQIEQELQRARQQGIFISTSSDQSHSASGNGALAFDMEYA 120
Query: 545 RWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
RWLEEHN+ I+ELRA V+AHA D DL+S VD IM+HY+EIF+LKG AAKADVFHVL
Sbjct: 121 RWLEEHNKHINELRAAVNAHAGDNDLKSTVDSIMAHYNEIFKLKGVAAKADVFHVL 176
>sptr|Q93XA0|Q93XA0 TGA-type basic leucine zipper protein TGA2.2.
Length = 460
Score = 194 bits (493), Expect = 1e-48
Identities = 106/177 (59%), Positives = 121/177 (68%)
Frame = +2
Query: 182 EVTMADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNREAA 361
E MADASPR N L RG DQKTLRRLAQNREAA
Sbjct: 135 ETNMADASPRTDTSTDDTEDKNQLP-ERG------ESSDRSKDKSDQKTLRRLAQNREAA 187
Query: 362 RKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEY 541
RKSRLRKKAYVQQLE+SRLKLT GIFISS+ DQ+ SMSGNGA AFD+EY
Sbjct: 188 RKSRLRKKAYVQQLESSRLKLTQLEQELQRARQQGIFISSTGDQAQSMSGNGAMAFDVEY 247
Query: 542 ARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
ARWLEEHNRQ +ELRA +++HA D +LR++VD M+ +D+IFRLKG AAKADVFH+L
Sbjct: 248 ARWLEEHNRQTNELRAAINSHAGDIELRTIVDNFMAQFDDIFRLKGIAAKADVFHIL 304
>sw|Q41558|HBB2_WHEAT Transcription factor HBP-1b(c1) (Fragment).
Length = 476
Score = 191 bits (486), Expect = 6e-48
Identities = 107/181 (59%), Positives = 116/181 (64%), Gaps = 4/181 (2%)
Frame = +2
Query: 182 EVTMADASPRXXXXXX----XXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQN 349
E +MAD SPR N E G+ D K+LRRLAQN
Sbjct: 140 ESSMADTSPRTDTSTDPDIDIDERNQMFEQGQLAAPTASDSSDKSRDKLDHKSLRRLAQN 199
Query: 350 REAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAF 529
REAARKSRLRKKAY+Q LE+SRLKLT GIFISSS DQS S SGNGA AF
Sbjct: 200 REAARKSRLRKKAYIQNLESSRLKLTQLEQELQRARQQGIFISSSGDQSQSASGNGAVAF 259
Query: 530 DMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHV 709
DMEYARWLEEHN+ I+ELRA +AHA D DLR +VD IMS YDE FRLKG AAKADVFHV
Sbjct: 260 DMEYARWLEEHNKHINELRAAANAHAGDDDLRKIVDSIMSQYDEFFRLKGVAAKADVFHV 319
Query: 710 L 712
L
Sbjct: 320 L 320
>sptr|Q8W5S0|Q8W5S0 bZIP transcription factor.
Length = 333
Score = 191 bits (485), Expect = 8e-48
Identities = 96/132 (72%), Positives = 109/132 (82%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQK LRRLAQNREAARKSRLRKKAYVQQLE+S+LKL GI+ISSS DQ+
Sbjct: 47 DQKVLRRLAQNREAARKSRLRKKAYVQQLESSKLKLASLEQEINKARQQGIYISSSGDQT 106
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
H+MSGNGA FD+EYARWLEE N+QI+ELR V+AHASD+DLR +VD IM+HYDEIFRLK
Sbjct: 107 HAMSGNGAMTFDLEYARWLEEQNKQINELRTAVNAHASDSDLRLIVDGIMAHYDEIFRLK 166
Query: 677 GNAAKADVFHVL 712
G AAKADVFH+L
Sbjct: 167 GVAAKADVFHIL 178
>sptr|Q7X993|Q7X993 bZIP transcription factor.
Length = 334
Score = 191 bits (484), Expect = 1e-47
Identities = 104/179 (58%), Positives = 121/179 (67%), Gaps = 5/179 (2%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXX--NNGLELGRGGLXXXXXXXXXXXXHG---DQKTLRRLAQNRE 355
MADAS R N+ LE G G DQKT+RRLAQNRE
Sbjct: 1 MADASSRTDTSIVVDNDDKNHQLENGHSGAVMASNSSDRSDRSDKLMDQKTIRRLAQNRE 60
Query: 356 AARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDM 535
AARKSRLRKKAYVQQLE+S+LKL GIFISSS DQ+H+MSGNGA FD+
Sbjct: 61 AARKSRLRKKAYVQQLESSKLKLAQLEQELQKARQQGIFISSSGDQTHAMSGNGALTFDL 120
Query: 536 EYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
EY RWLEE N+QI+ELR V+AHASD+DLR +VD IM+HYDE+F++KG AAKADVFH+L
Sbjct: 121 EYTRWLEEQNKQINELRTAVNAHASDSDLRLIVDGIMAHYDEVFKVKGVAAKADVFHIL 179
>sptr|Q8W5R9|Q8W5R9 bZIP transcription factor.
Length = 334
Score = 191 bits (484), Expect = 1e-47
Identities = 104/179 (58%), Positives = 121/179 (67%), Gaps = 5/179 (2%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXX--NNGLELGRGGLXXXXXXXXXXXXHG---DQKTLRRLAQNRE 355
MADAS R N+ LE G G DQKT+RRLAQNRE
Sbjct: 1 MADASSRTDTSIVVDNDDKNHQLENGHSGAVMASNSSDRSDRSDKLMDQKTIRRLAQNRE 60
Query: 356 AARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDM 535
AARKSRLRKKAYVQQLE+S+LKL GIFISSS DQ+H+MSGNGA FD+
Sbjct: 61 AARKSRLRKKAYVQQLESSKLKLAQLEQELQKARQQGIFISSSGDQTHAMSGNGALTFDL 120
Query: 536 EYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
EY RWLEE N+QI+ELR V+AHASD+DLR +VD IM+HYDE+F++KG AAKADVFH+L
Sbjct: 121 EYTRWLEEQNKQINELRTAVNAHASDSDLRLIVDGIMAHYDEVFKVKGVAAKADVFHIL 179
>sptr|Q06979|Q06979 Ocs-element binding factor 3.2.
Length = 468
Score = 190 bits (483), Expect = 1e-47
Identities = 97/132 (73%), Positives = 106/132 (80%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAY+Q LE+SRLKLT GIFIS+S DQ
Sbjct: 181 DQKTLRRLAQNREAARKSRLRKKAYIQNLESSRLKLTQLEQELQRARQQGIFISTSGDQP 240
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
S SGNGA AFDMEYARWLEEHN+ ++ELRA V+AHA D DLR +VD IM HYDEIF+LK
Sbjct: 241 QSTSGNGALAFDMEYARWLEEHNKHVNELRAAVNAHAGDNDLRCIVDSIMVHYDEIFKLK 300
Query: 677 GNAAKADVFHVL 712
G AAKADVFHVL
Sbjct: 301 GVAAKADVFHVL 312
>sptr|Q9SQK1|Q9SQK1 bZIP transcription factor.
Length = 325
Score = 187 bits (475), Expect = 1e-46
Identities = 101/174 (58%), Positives = 118/174 (67%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHGDQKTLRRLAQNREAARKS 370
MAD SP N L + G DQKTLRRLAQNREAARKS
Sbjct: 1 MADISPSTSTDADTEDKNRFLNSQQLGAVASDGSDRTR----DQKTLRRLAQNREAARKS 56
Query: 371 RLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYARW 550
RLRKKAYVQQLE+SR+KLT GIFIS S DQS SMSGNGA AFD+EYARW
Sbjct: 57 RLRKKAYVQQLESSRMKLTQLEQELQRARQQGIFISGSGDQSQSMSGNGALAFDVEYARW 116
Query: 551 LEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
LEE NR+I+ELR V++HA D +LR +VD I++HYD+IFR+KG+AAK+DVFH+L
Sbjct: 117 LEEQNRRINELRGAVNSHAGDGELRIIVDGILAHYDDIFRIKGDAAKSDVFHIL 170
>sptr|Q9C7D1|Q9C7D1 Transcription factor HBP-1B-like (Transcription
factor TGA6).
Length = 330
Score = 187 bits (475), Expect = 1e-46
Identities = 99/175 (56%), Positives = 118/175 (67%), Gaps = 1/175 (0%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHG-DQKTLRRLAQNREAARK 367
MAD S R + L RG + DQKTLRRLAQNREAARK
Sbjct: 1 MADTSSRTDVSTDGDTDHRDLGSDRGHMHAAASDSSDRSKDKLDQKTLRRLAQNREAARK 60
Query: 368 SRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYAR 547
SRLRKKAYVQQLENSRLKLT G+FISSS DQ+HS GNGA AFD E++R
Sbjct: 61 SRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISSSGDQAHSTGGNGALAFDAEHSR 120
Query: 548 WLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
WLEE NRQ++ELR+ ++AHA DT+LR +VD +M+HY+E+FR+K NAAK DVFH+L
Sbjct: 121 WLEEKNRQMNELRSALNAHAGDTELRIIVDGVMAHYEELFRIKSNAAKNDVFHLL 175
>sptr|Q9FL50|Q9FL50 Transcription factor HBP-1b (AT5g06960/MOJ9_13).
Length = 330
Score = 184 bits (467), Expect = 1e-45
Identities = 94/175 (53%), Positives = 118/175 (67%), Gaps = 1/175 (0%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHG-DQKTLRRLAQNREAARK 367
M D SPR +N L G L DQKTLRRLAQNREAARK
Sbjct: 1 MGDTSPRTSVSTDGDTDHNNLMFDEGHLGIGASDSSDRSKSKMDQKTLRRLAQNREAARK 60
Query: 368 SRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYAR 547
SRLRKKAYVQQLENSRLKLT G+FISSS DQ+HS +G+GA AFD+EY R
Sbjct: 61 SRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISSSGDQAHSTAGDGAMAFDVEYRR 120
Query: 548 WLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
W E+ NRQ+ EL + + +HA+D++LR +VD +++HY+E++R+KGNAAK+DVFH+L
Sbjct: 121 WQEDKNRQMKELSSAIDSHATDSELRIIVDGVIAHYEELYRIKGNAAKSDVFHLL 175
>sptr|Q06978|Q06978 Ocs-element binding factor 3.1.
Length = 331
Score = 184 bits (467), Expect = 1e-45
Identities = 92/132 (69%), Positives = 104/132 (78%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAY+Q LE+SRLKLT GIFIS+S DQ
Sbjct: 44 DQKTLRRLAQNREAARKSRLRKKAYIQNLESSRLKLTQLEQELHQTRQQGIFISTSGDQP 103
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
S SGNGA AFDMEYARWLEEHN+ ++ELR V+AHA D DLR +V +M+HYDE FRLK
Sbjct: 104 QSTSGNGALAFDMEYARWLEEHNKHVNELRLAVNAHAGDNDLRGIVGSVMAHYDEFFRLK 163
Query: 677 GNAAKADVFHVL 712
G AA++DVFHVL
Sbjct: 164 GVAARSDVFHVL 175
>sptrnew|AAP04119|AAP04119 Putative bZIP transcription factor,
HBP-1b homolog.
Length = 330
Score = 183 bits (464), Expect = 2e-45
Identities = 97/176 (55%), Positives = 118/176 (67%), Gaps = 2/176 (1%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHG--DQKTLRRLAQNREAAR 364
MAD SPR + L G L G DQKTLRRLAQNREAAR
Sbjct: 1 MADTSPRTDVSTDDDTDHPDLG-SEGALVNTAASDSSDRSKGKMDQKTLRRLAQNREAAR 59
Query: 365 KSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYA 544
KSRLRKKAYVQQLENSRLKLT G+FIS + DQ+HS GNGA AFD E++
Sbjct: 60 KSRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISGTGDQAHSTGGNGALAFDAEHS 119
Query: 545 RWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
RWLEE N+Q++ELR+ ++AHA D++LR +VD +M+HY+E+FR+K NAAK DVFH+L
Sbjct: 120 RWLEEKNKQMNELRSALNAHAGDSELRIIVDGVMAHYEELFRIKSNAAKNDVFHLL 175
>sptrnew|BAC42338|BAC42338 Putative bZip transcription factor
AtbZip20/tga2.
Length = 330
Score = 183 bits (464), Expect = 2e-45
Identities = 97/176 (55%), Positives = 118/176 (67%), Gaps = 2/176 (1%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHG--DQKTLRRLAQNREAAR 364
MAD SPR + L G L G DQKTLRRLAQNREAAR
Sbjct: 1 MADTSPRTDVSTDDDTDHPDLG-SEGALVNTAASDSSDRSKGKMDQKTLRRLAQNREAAR 59
Query: 365 KSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYA 544
KSRLRKKAYVQQLENSRLKLT G+FIS + DQ+HS GNGA AFD E++
Sbjct: 60 KSRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISGTGDQAHSTGGNGALAFDAEHS 119
Query: 545 RWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
RWLEE N+Q++ELR+ ++AHA D++LR +VD +M+HY+E+FR+K NAAK DVFH+L
Sbjct: 120 RWLEEKNKQMNELRSALNAHAGDSELRIIVDGVMAHYEELFRIKSNAAKNDVFHLL 175
>sw|P43273|HBPB_ARATH Transcription factor HBP-1b homolog.
Length = 330
Score = 183 bits (464), Expect = 2e-45
Identities = 97/176 (55%), Positives = 118/176 (67%), Gaps = 2/176 (1%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXXHG--DQKTLRRLAQNREAAR 364
MAD SPR + L G L G DQKTLRRLAQNREAAR
Sbjct: 1 MADTSPRTDVSTDDDTDHPDLG-SEGALVNTAASDSSDRSKGKMDQKTLRRLAQNREAAR 59
Query: 365 KSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYA 544
KSRLRKKAYVQQLENSRLKLT G+FIS + DQ+HS GNGA AFD E++
Sbjct: 60 KSRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISGTGDQAHSTGGNGALAFDAEHS 119
Query: 545 RWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLKGNAAKADVFHVL 712
RWLEE N+Q++ELR+ ++AHA D++LR +VD +M+HY+E+FR+K NAAK DVFH+L
Sbjct: 120 RWLEEKNKQMNELRSALNAHAGDSELRIIVDGVMAHYEELFRIKSNAAKNDVFHLL 175
>sptr|Q9LHI1|Q9LHI1 Leucine zipper transcription factor HBP-1b.
Length = 325
Score = 180 bits (456), Expect = 2e-44
Identities = 91/133 (68%), Positives = 108/133 (81%), Gaps = 1/133 (0%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLT G+FISSS DQ+
Sbjct: 38 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISSSGDQA 97
Query: 497 HSMSGNG-ASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRL 673
HS GNG A AFD E++RWLEE NRQ++ELR+ ++AHA DT+LR +VD +M+HY+E+FR+
Sbjct: 98 HSTGGNGGALAFDAEHSRWLEEKNRQMNELRSALNAHAGDTELRIIVDGVMAHYEELFRI 157
Query: 674 KGNAAKADVFHVL 712
K NAAK DVFH+L
Sbjct: 158 KSNAAKNDVFHLL 170
>sptrnew|BAA06486|BAA06486 Transcription factor HBP-1b(c38)
(Fragment).
Length = 152
Score = 179 bits (455), Expect = 3e-44
Identities = 98/152 (64%), Positives = 106/152 (69%), Gaps = 1/152 (0%)
Frame = +2
Query: 191 MADASPRXXXXXXXXXXNNGLELGRGGLXXXXXXXXXXXX-HGDQKTLRRLAQNREAARK 367
MA+ASPR N LE G L +GDQKT+RRLAQNREAARK
Sbjct: 1 MAEASPRTETSTDDTDENLMLEPGNAALAVVSDSSDRSRDKNGDQKTMRRLAQNREAARK 60
Query: 368 SRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSHSMSGNGASAFDMEYAR 547
SRLRKKAYVQQLENSRLKLT GIFISSS DQSHSMSGNGA AFD EYAR
Sbjct: 61 SRLRKKAYVQQLENSRLKLTQLEQELQRARQQGIFISSSADQSHSMSGNGALAFDTEYAR 120
Query: 548 WLEEHNRQISELRAGVSAHASDTDLRSVVDKI 643
WLEEHNRQ++ELRA V+AHA DT+LRSVV+KI
Sbjct: 121 WLEEHNRQVNELRAAVNAHAGDTELRSVVEKI 152
>sw|O24160|TG21_TOBAC TGACG-sequence specific DNA-binding protein
TGA-2.1 (TGA2.1).
Length = 456
Score = 179 bits (453), Expect = 4e-44
Identities = 88/132 (66%), Positives = 105/132 (79%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKL+ G +IS+ DQS
Sbjct: 166 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLSQLEQDLQRARQQGKYISNIADQS 225
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
+ + NG AFD EY+RWLEEHN+ I+ELR V+AHASD +LRS+V+ + +H+DE+FR+K
Sbjct: 226 NGVGANGPLAFDAEYSRWLEEHNKHINELRTAVNAHASDPELRSIVNNVTAHFDEVFRVK 285
Query: 677 GNAAKADVFHVL 712
GNAAKADVFHVL
Sbjct: 286 GNAAKADVFHVL 297
>sptr|Q39163|Q39163 Ocs-element binding factor 5 (Fragment).
Length = 324
Score = 178 bits (451), Expect = 7e-44
Identities = 85/132 (64%), Positives = 108/132 (81%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLT G+FISSS DQ+
Sbjct: 38 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTQLEQELQRARQQGVFISSSGDQA 97
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
HS +G+GA AFD+EY RW E+ NRQ+ EL + + +HA+D++LR +VD +++HY+E++R+K
Sbjct: 98 HSTAGDGAMAFDVEYRRWQEDKNRQMKELSSAIDSHATDSELRIIVDGVIAHYEELYRIK 157
Query: 677 GNAAKADVFHVL 712
GNAAK+DVFH+L
Sbjct: 158 GNAAKSDVFHLL 169
>sptr|Q39140|Q39140 Leucine zipper.
Length = 325
Score = 177 bits (448), Expect = 2e-43
Identities = 89/133 (66%), Positives = 108/133 (81%), Gaps = 1/133 (0%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
DQKTLRRLAQNREAARKSRLRKKAYVQQLE+SRLKLT G+FISSS DQ+
Sbjct: 38 DQKTLRRLAQNREAARKSRLRKKAYVQQLEDSRLKLTQVEQELQRARQQGVFISSSGDQA 97
Query: 497 HSMSGNG-ASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRL 673
HS GNG A AFD E++RWLEE NRQ++ELR+ ++AHA DT+LR +VD +M+HY+E+FR+
Sbjct: 98 HSTGGNGGALAFDAEHSRWLEEKNRQMNELRSALNAHAGDTELRIIVDGVMAHYEELFRI 157
Query: 674 KGNAAKADVFHVL 712
K NA+K DVFH+L
Sbjct: 158 KSNASKNDVFHLL 170
>sptr|Q9SX27|Q9SX27 F24J5.12 protein (Putative bZIP transcription
factor, PERIANTHIA).
Length = 452
Score = 156 bits (395), Expect = 2e-37
Identities = 78/134 (58%), Positives = 95/134 (70%), Gaps = 2/134 (1%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSV--D 490
DQ+TLRRLAQNREAARKSRLRKKAYVQQLENSR++L G + V D
Sbjct: 164 DQRTLRRLAQNREAARKSRLRKKAYVQQLENSRIRLAQLEEELKRARQQGSLVERGVSAD 223
Query: 491 QSHSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFR 670
+H +GNG +F++EY RW EEH R I++LR+GV++ D DLR +VD +MSHYDEIFR
Sbjct: 224 HTHLAAGNGVFSFELEYTRWKEEHQRMINDLRSGVNSQLGDNDLRVLVDAVMSHYDEIFR 283
Query: 671 LKGNAAKADVFHVL 712
LKG K DVFH+L
Sbjct: 284 LKGIGTKVDVFHML 297
>sptr|Q9ZPJ6|Q9ZPJ6 Transcription factor PERIANTHIA.
Length = 451
Score = 155 bits (391), Expect = 7e-37
Identities = 77/134 (57%), Positives = 95/134 (70%), Gaps = 2/134 (1%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSV--D 490
DQ+TLRRLAQNREAARKSRLRKKAYVQQLENSR++L G + V D
Sbjct: 163 DQRTLRRLAQNREAARKSRLRKKAYVQQLENSRIRLAQLEEELKRARQQGSLVERGVSAD 222
Query: 491 QSHSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFR 670
+H +GNG +F++EY RW EEH R I++LR+GV++ D DLR +VD +MSHYDEIFR
Sbjct: 223 HTHLAAGNGVFSFELEYTRWKEEHQRMINDLRSGVNSQLGDNDLRVLVDAVMSHYDEIFR 282
Query: 671 LKGNAAKADVFHVL 712
LKG K +VFH+L
Sbjct: 283 LKGIGTKVEVFHML 296
>sptr|Q9SJF8|Q9SJF8 T27G7.2.
Length = 609
Score = 151 bits (382), Expect = 7e-36
Identities = 77/132 (58%), Positives = 97/132 (73%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
D KTLRRLAQNREAARKSRLRKKAYVQQLE+SR+KL+ G+F+
Sbjct: 24 DAKTLRRLAQNREAARKSRLRKKAYVQQLESSRIKLSQLEQELQRARSQGLFMGGCGPPG 83
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
+++ +GA+ FDMEY RWLE+ NR +SE+R G+ AH SD DLR +VD ++H+DEIFRLK
Sbjct: 84 PNIT-SGAAIFDMEYGRWLEDDNRHMSEIRTGLQAHLSDNDLRLIVDGYIAHFDEIFRLK 142
Query: 677 GNAAKADVFHVL 712
AAKADVFH++
Sbjct: 143 AVAAKADVFHLI 154
>sptr|Q93XM6|Q93XM6 bZIP transcription factor.
Length = 481
Score = 151 bits (382), Expect = 7e-36
Identities = 77/132 (58%), Positives = 97/132 (73%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQS 496
D KTLRRLAQNREAARKSRLRKKAYVQQLE+SR+KL+ G+F+
Sbjct: 176 DAKTLRRLAQNREAARKSRLRKKAYVQQLESSRIKLSQLEQELQRARSQGLFMGGCGPPG 235
Query: 497 HSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFRLK 676
+++ +GA+ FDMEY RWLE+ NR +SE+R G+ AH SD DLR +VD ++H+DEIFRLK
Sbjct: 236 PNIT-SGAAIFDMEYGRWLEDDNRHMSEIRTGLQAHLSDNDLRLIVDGYIAHFDEIFRLK 294
Query: 677 GNAAKADVFHVL 712
AAKADVFH++
Sbjct: 295 AVAAKADVFHLI 306
>sptr|Q9SGC7|Q9SGC7 T23G18.19.
Length = 709
Score = 141 bits (355), Expect = 1e-32
Identities = 76/145 (52%), Positives = 97/145 (66%), Gaps = 13/145 (8%)
Frame = +2
Query: 317 DQKTLRRLAQNREAARKSRLRKK-------------AYVQQLENSRLKLTXXXXXXXXXX 457
+ KTLRRLAQNREAARKSRLRKK AYVQQLE+SR+KL+
Sbjct: 111 EDKTLRRLAQNREAARKSRLRKKYENNKCIYIWFEQAYVQQLESSRIKLSQLEQELQRAR 170
Query: 458 XXGIFISSSVDQSHSMSGNGASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVD 637
G+F+ +++ +GA+ FDMEY RWLE+ NR +SE+R G+ AH SD DLR +VD
Sbjct: 171 SQGLFMGGCGPPGPNIT-SGAAIFDMEYGRWLEDDNRHMSEIRTGLQAHLSDNDLRLIVD 229
Query: 638 KIMSHYDEIFRLKGNAAKADVFHVL 712
++H+DEIFRLK AAKADVFH++
Sbjct: 230 GYIAHFDEIFRLKAVAAKADVFHLI 254
>sptr|Q93XA2|Q93XA2 TGA-type basic leucine zipper protein TGA1.1.
Length = 362
Score = 140 bits (352), Expect = 2e-32
Identities = 74/134 (55%), Positives = 93/134 (69%), Gaps = 4/134 (2%)
Frame = +2
Query: 323 KTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTXXXXXXXXXXXXGIFISSSVDQSH- 499
K RRLAQNREAARKSRLRKKAYVQQLE+SRLKL G++I +D +H
Sbjct: 78 KIQRRLAQNREAARKSRLRKKAYVQQLESSRLKLMQLEQELERARHQGMYIGGGLDSNHM 137
Query: 500 SMSGN---GASAFDMEYARWLEEHNRQISELRAGVSAHASDTDLRSVVDKIMSHYDEIFR 670
SG+ G + F+MEY W+ E NRQI+ELR ++AH D +LR +VD +M+HY EIFR
Sbjct: 138 GFSGSVNSGITTFEMEYGHWVNEQNRQITELRTALNAHIGDIELRILVDGMMNHYAEIFR 197
Query: 671 LKGNAAKADVFHVL 712
+K AAKADVF+V+
Sbjct: 198 MKSAAAKADVFYVM 211
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 473,061,664
Number of Sequences: 1381838
Number of extensions: 8650403
Number of successful extensions: 20073
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 19099
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 19978
length of database: 439,479,560
effective HSP length: 120
effective length of database: 273,659,000
effective search space used: 32018103000
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

