BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QBF12h04.xg.3.2
(747 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
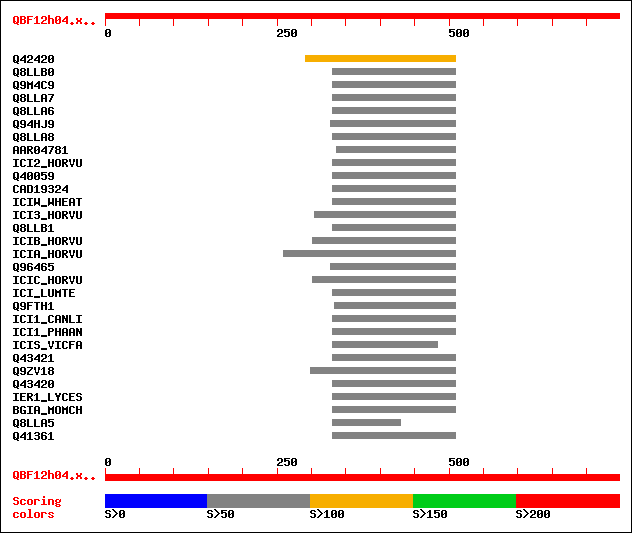
Sequences producing significant alignments: (bits) Value
sptr|Q42420|Q42420 SUBSTILIN /chymotrypsin-like inhibitor. 133 2e-30
sptr|Q8LLB0|Q8LLB0 CI2E. 100 3e-20
sptr|Q9M4C9|Q9M4C9 Hypothetical protein. 96 4e-19
sptr|Q8LLA7|Q8LLA7 CI2C. 96 7e-19
sptr|Q8LLA6|Q8LLA6 CI2B. 90 3e-17
sptr|Q94HJ9|Q94HJ9 Hypothetical protein. 87 2e-16
sptr|Q8LLA8|Q8LLA8 CI2D. 86 4e-16
sptrnew|AAR04781|AAR04781 Serine proteinase inhibitor. 85 9e-16
sw|P01053|ICI2_HORVU Subtilisin-chymotrypsin inhibitor-2A (CI-2A). 84 2e-15
sptr|Q40059|Q40059 Chymotrypsin inhibitor 2. 83 5e-15
sptrnew|CAD19324|CAD19324 WSCI proteinase inhibitor (Fragment). 82 8e-15
sw|P82977|ICIW_WHEAT Subtilisin-chymotrypsin inhibitor WSCI. 82 8e-15
sw|P08626|ICI3_HORVU Subtilisin-chymotrypsin inhibitor-2B (CI-2B... 82 8e-15
sptr|Q8LLB1|Q8LLB1 CI2F. 80 3e-14
sw|P16063|ICIB_HORVU Subtilisin-chymotrypsin inhibitor CI-1B. 76 4e-13
sw|P16062|ICIA_HORVU Subtilisin-chymotrypsin inhibitor CI-1A. 74 2e-12
sptr|Q96465|Q96465 Subtilisin-chymotrypsin inhibitor 2 (Putative... 73 5e-12
sw|P01054|ICIC_HORVU Subtilisin-chymotrypsin inhibitor CI-1C. 70 3e-11
sw|P83472|ICI_LUMTE Chymotrypsin inhibitor (LTCI). 67 3e-10
sptr|Q9FTH1|Q9FTH1 P0410E01.29 protein. 66 4e-10
sw|P81712|ICI1_CANLI Subtilisin inhibitor CLSI-I. 63 4e-09
sw|P16064|ICI1_PHAAN Subtilisin inhibitor I (ASI-I) [Contains: S... 61 2e-08
sw|P08820|ICIS_VICFA Subtilisin inhibitor (Fragment). 59 5e-08
sptr|Q43421|Q43421 Trypsin inhibitor. 58 2e-07
sptr|Q9ZV18|Q9ZV18 Putative protease inhibitor. 55 1e-06
sptr|Q43420|Q43420 Pumpkin fruit chymotrypsin inhibitor. 55 1e-06
sw|P20076|IER1_LYCES Ethylene-responsive proteinase inhibitor I ... 54 2e-06
sw|P24076|BGIA_MOMCH Glu S.griseus protease inhibitor (BGIA). 54 2e-06
sptr|Q8LLA5|Q8LLA5 CI2A. 53 5e-06
sptr|Q41361|Q41361 Pathogenesis-related protein PR-6 type (Fragm... 52 7e-06
>sptr|Q42420|Q42420 SUBSTILIN /chymotrypsin-like inhibitor.
Length = 73
Score = 133 bits (335), Expect = 2e-30
Identities = 67/73 (91%), Positives = 67/73 (91%)
Frame = +1
Query: 292 MSSTECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVR 471
MSSTEC AKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVR
Sbjct: 1 MSSTECGGGGGGAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVR 60
Query: 472 IFVDIVAQTPHIG 510
IFVDIVAQTPHIG
Sbjct: 61 IFVDIVAQTPHIG 73
>sptr|Q8LLB0|Q8LLB0 CI2E.
Length = 72
Score = 100 bits (248), Expect = 3e-20
Identities = 44/60 (73%), Positives = 55/60 (91%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
KTSWPEVVGL++++AK++ILKDKP+ADIVV+PVGS VT D+RPNRVRIFV V +TPH+G
Sbjct: 13 KTSWPEVVGLTIKEAKEIILKDKPEADIVVVPVGSAVTEDFRPNRVRIFVGTVVETPHVG 72
>sptr|Q9M4C9|Q9M4C9 Hypothetical protein.
Length = 74
Score = 96.3 bits (238), Expect = 4e-19
Identities = 45/60 (75%), Positives = 51/60 (85%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
KTSWPEV G S+E+AK++ILKD P+ADIVVLP GS VT D+R NRVRIFVD VA TPHIG
Sbjct: 15 KTSWPEVAGKSIEEAKEIILKDMPEADIVVLPAGSPVTLDFRTNRVRIFVDTVASTPHIG 74
>sptr|Q8LLA7|Q8LLA7 CI2C.
Length = 74
Score = 95.5 bits (236), Expect = 7e-19
Identities = 45/60 (75%), Positives = 51/60 (85%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
KTSWPEVVG S+E+AK++ILKD P+ADIVVLP GS VT D+R NRVRIFVD VA PHIG
Sbjct: 15 KTSWPEVVGKSIEEAKEIILKDMPEADIVVLPAGSPVTLDFRTNRVRIFVDTVASIPHIG 74
>sptr|Q8LLA6|Q8LLA6 CI2B.
Length = 77
Score = 90.1 bits (222), Expect = 3e-17
Identities = 42/60 (70%), Positives = 50/60 (83%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
K+SWPE+VG S+EDA++VILKD P+ADI VLP SVV AD+R NRVR+ VD VA TPHIG
Sbjct: 18 KSSWPELVGKSIEDAREVILKDMPEADIKVLPANSVVAADWRSNRVRVLVDTVATTPHIG 77
>sptr|Q94HJ9|Q94HJ9 Hypothetical protein.
Length = 71
Score = 87.0 bits (214), Expect = 2e-16
Identities = 43/62 (69%), Positives = 51/62 (82%), Gaps = 1/62 (1%)
Frame = +1
Query: 328 AKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD-IVAQTPH 504
AK SWPEVVG+++E+AK ILKDKPDADIVVLPVG+ +T D RPNRVRIF VA+TP
Sbjct: 10 AKRSWPEVVGMTMEEAKAAILKDKPDADIVVLPVGAPMTRDLRPNRVRIFGSATVAETPR 69
Query: 505 IG 510
+G
Sbjct: 70 VG 71
>sptr|Q8LLA8|Q8LLA8 CI2D.
Length = 74
Score = 86.3 bits (212), Expect = 4e-16
Identities = 41/60 (68%), Positives = 49/60 (81%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
K+SWPE+VG S+EDA++VILKD P+ DI V P SVV+AD+R NRVRI VD VA TPHIG
Sbjct: 15 KSSWPELVGKSIEDAREVILKDMPEVDIEVHPTDSVVSADWRSNRVRILVDTVATTPHIG 74
>sptrnew|AAR04781|AAR04781 Serine proteinase inhibitor.
Length = 69
Score = 85.1 bits (209), Expect = 9e-16
Identities = 41/58 (70%), Positives = 46/58 (79%)
Frame = +1
Query: 337 SWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
SWPEVVGLS+++AKKVILKDKPDADIV LPVG VT D+ NRVRIFVD + P G
Sbjct: 12 SWPEVVGLSIKEAKKVILKDKPDADIVALPVGGKVTDDFLSNRVRIFVDTGGEIPRAG 69
>sw|P01053|ICI2_HORVU Subtilisin-chymotrypsin inhibitor-2A (CI-2A).
Length = 83
Score = 84.3 bits (207), Expect = 2e-15
Identities = 40/63 (63%), Positives = 52/63 (82%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV---DIVAQTP 501
KT WPE+VG SVE+AKKVIL+DKP+A I+VLPVG++VT +YR +RVR+FV D +AQ P
Sbjct: 21 KTEWPELVGKSVEEAKKVILQDKPEAQIIVLPVGTIVTMEYRIDRVRLFVDKLDNIAQVP 80
Query: 502 HIG 510
+G
Sbjct: 81 RVG 83
>sptr|Q40059|Q40059 Chymotrypsin inhibitor 2.
Length = 84
Score = 82.8 bits (203), Expect = 5e-15
Identities = 40/63 (63%), Positives = 51/63 (80%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV---DIVAQTP 501
KT WPE+VG SVE+AKKVIL+DKP A I+VLPVG++VT +YR +RVR+FV D +AQ P
Sbjct: 22 KTEWPELVGKSVEEAKKVILQDKPAAQIIVLPVGTIVTMEYRIDRVRLFVDRLDNIAQVP 81
Query: 502 HIG 510
+G
Sbjct: 82 RVG 84
>sptrnew|CAD19324|CAD19324 WSCI proteinase inhibitor (Fragment).
Length = 72
Score = 82.0 bits (201), Expect = 8e-15
Identities = 40/63 (63%), Positives = 51/63 (80%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDI---VAQTP 501
KT WPE+VG SVE+AKKVIL+DK +A IVVLPVG++VT +YR +RVR+FVD +AQ P
Sbjct: 10 KTEWPELVGKSVEEAKKVILQDKSEAQIVVLPVGTIVTMEYRIDRVRLFVDSLDKIAQVP 69
Query: 502 HIG 510
+G
Sbjct: 70 RVG 72
>sw|P82977|ICIW_WHEAT Subtilisin-chymotrypsin inhibitor WSCI.
Length = 72
Score = 82.0 bits (201), Expect = 8e-15
Identities = 40/63 (63%), Positives = 51/63 (80%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDI---VAQTP 501
KT WPE+VG SVE+AKKVIL+DK +A IVVLPVG++VT +YR +RVR+FVD +AQ P
Sbjct: 10 KTEWPELVGKSVEEAKKVILQDKSEAQIVVLPVGTIVTMEYRIDRVRLFVDSLDKIAQVP 69
Query: 502 HIG 510
+G
Sbjct: 70 RVG 72
>sw|P08626|ICI3_HORVU Subtilisin-chymotrypsin inhibitor-2B (CI-2B)
(Fragment).
Length = 72
Score = 82.0 bits (201), Expect = 8e-15
Identities = 40/72 (55%), Positives = 53/72 (73%), Gaps = 3/72 (4%)
Frame = +1
Query: 304 ECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV- 480
+C KT WPE+V SVE+AKKVIL+DKP+A I+VLPVG++VT +YR +RVR+FV
Sbjct: 1 DCLCDCQNQKTEWPELVEKSVEEAKKVILQDKPEAQIIVLPVGTIVTMEYRIDRVRLFVD 60
Query: 481 --DIVAQTPHIG 510
D +AQ P +G
Sbjct: 61 RLDNIAQVPRVG 72
>sptr|Q8LLB1|Q8LLB1 CI2F.
Length = 84
Score = 80.1 bits (196), Expect = 3e-14
Identities = 37/63 (58%), Positives = 51/63 (80%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV---DIVAQTP 501
KT WPE+VG SV++AKKVIL+DKP+A I+VLP+G++VT +YR + VR+FV D +AQ P
Sbjct: 22 KTEWPELVGKSVQEAKKVILQDKPEAQIIVLPMGTIVTIEYRIDHVRLFVDRLDNIAQVP 81
Query: 502 HIG 510
+G
Sbjct: 82 RVG 84
>sw|P16063|ICIB_HORVU Subtilisin-chymotrypsin inhibitor CI-1B.
Length = 83
Score = 76.3 bits (186), Expect = 4e-13
Identities = 38/70 (54%), Positives = 49/70 (70%)
Frame = +1
Query: 301 TECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV 480
TE AK SWPEVVG+S E AK++IL+DKPDA I V+PV ++V D+ PNR+ I V
Sbjct: 15 TEGSIGASGAKRSWPEVVGMSAEKAKEIILRDKPDAQIEVIPVDAMVPLDFNPNRIFILV 74
Query: 481 DIVAQTPHIG 510
VA+TP +G
Sbjct: 75 -AVARTPTVG 83
>sw|P16062|ICIA_HORVU Subtilisin-chymotrypsin inhibitor CI-1A.
Length = 83
Score = 73.9 bits (180), Expect = 2e-12
Identities = 39/84 (46%), Positives = 55/84 (65%)
Frame = +1
Query: 259 ISSTGPAVRDTMSSTECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSV 438
+SS +V TE AKTSWPEVVG+S E AK++IL+DKP+A + V+PV ++
Sbjct: 1 MSSMEGSVLKYPEPTEGSIGASSAKTSWPEVVGMSAEKAKEIILRDKPNAQVEVIPVDAM 60
Query: 439 VTADYRPNRVRIFVDIVAQTPHIG 510
V ++ PNRV + V VA+TP +G
Sbjct: 61 VHLNFDPNRVFVLV-AVARTPTVG 83
>sptr|Q96465|Q96465 Subtilisin-chymotrypsin inhibitor 2 (Putative
protease inhibitor).
Length = 68
Score = 72.8 bits (177), Expect = 5e-12
Identities = 32/61 (52%), Positives = 47/61 (77%)
Frame = +1
Query: 328 AKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHI 507
AKT WPE+VG ++++AK+ I D+PD +V++PVGS+VT + NRVR++VD VA+ P I
Sbjct: 8 AKTEWPELVGCTIKEAKEKIKADRPDLKVVIVPVGSIVTQEIDLNRVRVWVDKVAKVPKI 67
Query: 508 G 510
G
Sbjct: 68 G 68
>sw|P01054|ICIC_HORVU Subtilisin-chymotrypsin inhibitor CI-1C.
Length = 77
Score = 70.1 bits (170), Expect = 3e-11
Identities = 35/73 (47%), Positives = 48/73 (65%), Gaps = 3/73 (4%)
Frame = +1
Query: 301 TECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV 480
TE AKTSWPEVVG+S E AK++IL+DKP+A I V+PV ++V ++ PNRV + V
Sbjct: 5 TEGSIGASGAKTSWPEVVGMSAEKAKEIILRDKPNAQIEVIPVDAMVPLNFNPNRVFVLV 64
Query: 481 ---DIVAQTPHIG 510
VA+ +G
Sbjct: 65 HKATTVAZVSRVG 77
>sw|P83472|ICI_LUMTE Chymotrypsin inhibitor (LTCI).
Length = 64
Score = 67.0 bits (162), Expect = 3e-10
Identities = 37/63 (58%), Positives = 46/63 (73%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD---IVAQTP 501
KTSWPE+VG ++E+AK IL+D+PDA I V P S VT DYRP+RV IFV+ VA+TP
Sbjct: 2 KTSWPELVGETLEEAKAQILEDRPDAVIKVQPEHSPVTYDYRPSRVIIFVNKDGNVAETP 61
Query: 502 HIG 510
G
Sbjct: 62 AAG 64
>sptr|Q9FTH1|Q9FTH1 P0410E01.29 protein.
Length = 66
Score = 66.2 bits (160), Expect = 4e-10
Identities = 30/59 (50%), Positives = 41/59 (69%)
Frame = +1
Query: 334 TSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVDIVAQTPHIG 510
T WPE+VGL++E AK I D+PD + VLPVG+++ PNRV ++VD VA+ P IG
Sbjct: 8 TEWPELVGLTIEQAKAKIKADRPDLQVEVLPVGTIILGVVVPNRVILWVDTVAEIPKIG 66
>sw|P81712|ICI1_CANLI Subtilisin inhibitor CLSI-I.
Length = 65
Score = 63.2 bits (152), Expect = 4e-09
Identities = 33/63 (52%), Positives = 46/63 (73%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD---IVAQTP 501
KTSWPE+VG++ E+A+K I ++ +I V+P GS VTADY+P RVR++VD V +TP
Sbjct: 4 KTSWPELVGVTAEEAEK-IKEEMSGVEIQVVPPGSFVTADYKPQRVRLYVDESNKVTRTP 62
Query: 502 HIG 510
IG
Sbjct: 63 GIG 65
>sw|P16064|ICI1_PHAAN Subtilisin inhibitor I (ASI-I) [Contains:
Subtilisin inhibitor II (ASI-II)].
Length = 92
Score = 60.8 bits (146), Expect = 2e-08
Identities = 32/63 (50%), Positives = 41/63 (65%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD---IVAQTP 501
KTSWPE+VG++ E A+ I ++ D I V P S VTADY P RVR++VD V +TP
Sbjct: 30 KTSWPELVGVTAEQAETKIKEEMVDVQIQVSPHDSFVTADYNPKRVRLYVDESNKVTRTP 89
Query: 502 HIG 510
IG
Sbjct: 90 SIG 92
>sw|P08820|ICIS_VICFA Subtilisin inhibitor (Fragment).
Length = 62
Score = 59.3 bits (142), Expect = 5e-08
Identities = 28/51 (54%), Positives = 41/51 (80%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD 483
+TSWPE+VG+S E+A+K I ++ P+A+I V+P S VTADY+ RVR++VD
Sbjct: 1 RTSWPELVGVSAEEARK-IKEEMPEAEIQVVPQDSFVTADYKFQRVRLYVD 50
>sptr|Q43421|Q43421 Trypsin inhibitor.
Length = 67
Score = 57.8 bits (138), Expect = 2e-07
Identities = 30/64 (46%), Positives = 41/64 (64%), Gaps = 4/64 (6%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVG-SVVTADYRPNRVRIFVD---IVAQT 498
K SWPE+VG E+A K+I ++ P D++++P G + T DYRPNRVR+F D V
Sbjct: 4 KLSWPELVGKDGEEAVKIIQQENPSLDVILMPRGQNWATKDYRPNRVRVFNDDSGKVNSI 63
Query: 499 PHIG 510
P IG
Sbjct: 64 PRIG 67
>sptr|Q9ZV18|Q9ZV18 Putative protease inhibitor.
Length = 70
Score = 55.1 bits (131), Expect = 1e-06
Identities = 33/74 (44%), Positives = 44/74 (59%), Gaps = 3/74 (4%)
Frame = +1
Query: 298 STECXXXXXXAKTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIF 477
STEC K SWPE+ G + + A VI ++ P + V+ GS VTAD+R +RVR+F
Sbjct: 2 STECPR-----KNSWPELTGTNGDYAAVVIERENPTVNAAVILDGSPVTADFRCDRVRVF 56
Query: 478 VD---IVAQTPHIG 510
VD IV +TP G
Sbjct: 57 VDGNRIVVKTPKSG 70
>sptr|Q43420|Q43420 Pumpkin fruit chymotrypsin inhibitor.
Length = 67
Score = 54.7 bits (130), Expect = 1e-06
Identities = 29/64 (45%), Positives = 41/64 (64%), Gaps = 4/64 (6%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVG-SVVTADYRPNRVRIFVD---IVAQT 498
K+SWPE+VG E+A K+I ++ P D++++P G + T D RPNRVR+F D V
Sbjct: 4 KSSWPELVGEDGEEAVKIIQQENPSLDVILMPRGQNWATLDCRPNRVRVFNDESGKVNSI 63
Query: 499 PHIG 510
P IG
Sbjct: 64 PRIG 67
>sw|P20076|IER1_LYCES Ethylene-responsive proteinase inhibitor I
precursor.
Length = 119
Score = 53.9 bits (128), Expect = 2e-06
Identities = 30/64 (46%), Positives = 40/64 (62%), Gaps = 4/64 (6%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPD-ADIVVLPVGSVVTADYRPNRVRIFV---DIVAQT 498
K SWPE++G + AK++I K+ P ++ L GS T D R NRVR+FV DIV QT
Sbjct: 56 KESWPELLGTPAKFAKQIIQKENPKLTNVETLLNGSAFTEDLRCNRVRLFVNLLDIVVQT 115
Query: 499 PHIG 510
P +G
Sbjct: 116 PKVG 119
>sw|P24076|BGIA_MOMCH Glu S.griseus protease inhibitor (BGIA).
Length = 68
Score = 53.9 bits (128), Expect = 2e-06
Identities = 30/63 (47%), Positives = 42/63 (66%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFV---DIVAQTP 501
K SWP++VG + AK VI ++ P V++ VGS VTAD+R +RVR++V IVA+ P
Sbjct: 6 KRSWPQLVGSTGAAAKAVIERENPRVRAVIVRVGSPVTADFRCDRVRVWVTERGIVARPP 65
Query: 502 HIG 510
IG
Sbjct: 66 AIG 68
>sptr|Q8LLA5|Q8LLA5 CI2A.
Length = 55
Score = 52.8 bits (125), Expect = 5e-06
Identities = 24/33 (72%), Positives = 29/33 (87%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPV 429
K+SW E+VG S+EDA+KVILKD P+ADI VLPV
Sbjct: 18 KSSWAELVGKSIEDARKVILKDMPEADIKVLPV 50
>sptr|Q41361|Q41361 Pathogenesis-related protein PR-6 type
(Fragment).
Length = 79
Score = 52.4 bits (124), Expect = 7e-06
Identities = 26/63 (41%), Positives = 38/63 (60%), Gaps = 3/63 (4%)
Frame = +1
Query: 331 KTSWPEVVGLSVEDAKKVILKDKPDADIVVLPVGSVVTADYRPNRVRIFVD---IVAQTP 501
K +WPE+ G E+A + + P V++P GS+VT D R +RVR++VD IV + P
Sbjct: 17 KNTWPELCGARGEEAAATVETENPSVTAVIVPEGSIVTTDERCDRVRVWVDENGIVTRVP 76
Query: 502 HIG 510
IG
Sbjct: 77 VIG 79
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 455,258,785
Number of Sequences: 1381838
Number of extensions: 8700378
Number of successful extensions: 34398
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 31327
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 34065
length of database: 439,479,560
effective HSP length: 120
effective length of database: 273,659,000
effective search space used: 35028352000
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

