BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QAZ4d09.yg.2.1
(542 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
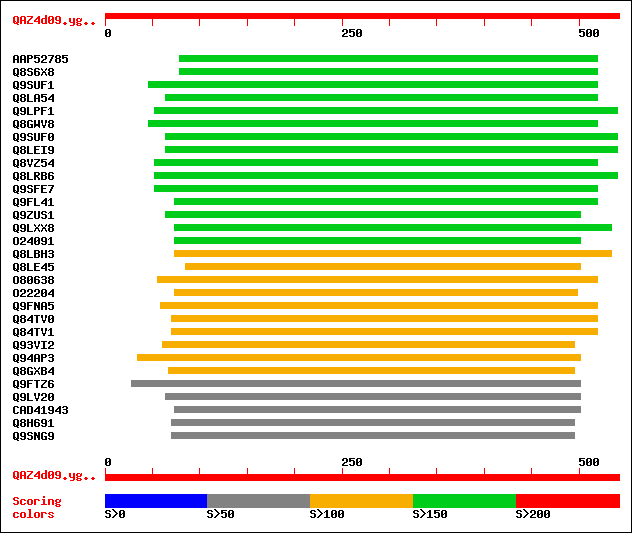
Sequences producing significant alignments: (bits) Value
sptrnew|AAP52785|AAP52785 Putative nodulin-like protein. 182 3e-45
sptr|Q8S6X8|Q8S6X8 Putative nodulin-like protein. 182 3e-45
sptr|Q9SUF1|Q9SUF1 Nodulin-like protein. 168 4e-41
sptr|Q8LA54|Q8LA54 Nodulin-like protein. 166 2e-40
sptr|Q9LPF1|Q9LPF1 T12C22.7 protein (At1g44800/T12C22_7). 165 3e-40
sptr|Q8GWV8|Q8GWV8 Putative nodulin (At4g08290). 164 4e-40
sptr|Q9SUF0|Q9SUF0 Nodulin-like protein. 163 1e-39
sptr|Q8LEI9|Q8LEI9 Nodulin protein, putative. 162 2e-39
sptr|Q8VZ54|Q8VZ54 Putative nodulin protein. 160 6e-39
sptr|Q8LRB6|Q8LRB6 Nodulin-like protein. 158 3e-38
sptr|Q9SFE7|Q9SFE7 T26F17.11. 158 3e-38
sptr|Q9FL41|Q9FL41 MtN21 nodulin protein-like. 155 2e-37
sptr|Q9ZUS1|Q9ZUS1 Nodulin-like protein (Putative nodulin protein). 154 6e-37
sptr|Q9LXX8|Q9LXX8 Nodulin-like protein. 152 2e-36
sptr|O24091|O24091 MtN21 protein. 151 5e-36
sptr|Q8LBH3|Q8LBH3 Nodulin-like protein. 149 3e-35
sptr|Q8LE45|Q8LE45 Nodulin-like protein. 148 4e-35
sptr|O80638|O80638 Nodulin-like protein (At2g39510). 147 7e-35
sptr|O22204|O22204 Putative integral membrane protein nodulin. 144 6e-34
sptr|Q9FNA5|Q9FNA5 Similarity to MtN21 (Nodulin-like protein). 128 4e-29
sptr|Q84TV0|Q84TV0 Putative nodulin protein. 115 3e-25
sptr|Q84TV1|Q84TV1 Nodulin-like protein. 115 4e-25
sptr|Q93VI2|Q93VI2 Putative MtN21. 111 4e-24
sptr|Q94AP3|Q94AP3 Hypothetical protein (Putative nodulin protein). 110 8e-24
sptr|Q8GXB4|Q8GXB4 Putative nodulin protein N21. 101 5e-21
sptr|Q9FTZ6|Q9FTZ6 Putative nodulin (P0494A10.12 protein). 100 1e-20
sptr|Q9LV20|Q9LV20 Nodulin-like protein. 96 3e-19
sptrnew|CAD41943|CAD41943 OSJNBa0070M12.20 protein. 96 3e-19
sptr|Q8H691|Q8H691 OSJNBa0004I20.6 protein. 95 4e-19
sptr|Q9SNG9|Q9SNG9 ESTs AU055729(S20023). 95 4e-19
>sptrnew|AAP52785|AAP52785 Putative nodulin-like protein.
Length = 330
Score = 182 bits (461), Expect = 3e-45
Identities = 101/147 (68%), Positives = 107/147 (72%)
Frame = +1
Query: 79 MAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMSLP 258
MAMVFLQFGFAG+FLISVASLRQGMSHYVLVVYRN VAA+VMAPFALWFER
Sbjct: 1 MAMVFLQFGFAGLFLISVASLRQGMSHYVLVVYRNAVAAVVMAPFALWFER--------- 51
Query: 259 VFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIXXR 438
PVLDQNF YMG +TSASFSSALTNILPA+TFVNAIILRMERI I R
Sbjct: 52 ------------PVLDQNFFYMGAKNTSASFSSALTNILPAVTFVNAIILRMERISIKER 99
Query: 439 RSQAXXXGTAXTVXXXLLMILXXXXII 519
RSQA GT TV +LMIL +I
Sbjct: 100 RSQAKIAGTLITVGGAMLMILFKGPVI 126
>sptr|Q8S6X8|Q8S6X8 Putative nodulin-like protein.
Length = 330
Score = 182 bits (461), Expect = 3e-45
Identities = 101/147 (68%), Positives = 107/147 (72%)
Frame = +1
Query: 79 MAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMSLP 258
MAMVFLQFGFAG+FLISVASLRQGMSHYVLVVYRN VAA+VMAPFALWFER
Sbjct: 1 MAMVFLQFGFAGLFLISVASLRQGMSHYVLVVYRNAVAAVVMAPFALWFER--------- 51
Query: 259 VFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIXXR 438
PVLDQNF YMG +TSASFSSALTNILPA+TFVNAIILRMERI I R
Sbjct: 52 ------------PVLDQNFFYMGAKNTSASFSSALTNILPAVTFVNAIILRMERISIKER 99
Query: 439 RSQAXXXGTAXTVXXXLLMILXXXXII 519
RSQA GT TV +LMIL +I
Sbjct: 100 RSQAKIAGTLITVGGAMLMILFKGPVI 126
>sptr|Q9SUF1|Q9SUF1 Nodulin-like protein.
Length = 384
Score = 168 bits (425), Expect = 4e-41
Identities = 84/158 (53%), Positives = 117/158 (74%)
Frame = +1
Query: 46 LAGTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWF 225
++ T K+ PY+ M+FLQFG AG +++ +A+L QG + YV++VYRN+VAA+V+APFAL F
Sbjct: 4 VSATMHKLRPYLLMIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAPFALIF 63
Query: 226 ERKTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAII 405
ERK RPKM+L V KI+ALG LEPVLDQ F Y+G+N TSA+++SA+ NILP++TF+ A I
Sbjct: 64 ERKVRPKMTLSVLWKIMALGFLEPVLDQGFGYLGMNMTSATYTSAIMNILPSVTFIIAWI 123
Query: 406 LRMERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
LRME++ I RS+A GT + L+M L +I
Sbjct: 124 LRMEKVNIAEVRSKAKIIGTLVGLGGALVMTLYKGPLI 161
>sptr|Q8LA54|Q8LA54 Nodulin-like protein.
Length = 377
Score = 166 bits (419), Expect = 2e-40
Identities = 83/152 (54%), Positives = 114/152 (75%)
Frame = +1
Query: 64 KVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRP 243
K+ PY+ M+FLQFG AG +++ +A+L QG + YV++VYRN+VAA+V+APFAL FERK RP
Sbjct: 3 KLRPYLLMIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAPFALIFERKVRP 62
Query: 244 KMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERI 423
KM+L V KI+ALG LEPVLDQ F Y+G+N TSA+++SA+ NILP++TF+ A ILRME++
Sbjct: 63 KMTLSVLWKIMALGFLEPVLDQGFGYLGMNMTSATYTSAIMNILPSVTFIIAWILRMEKV 122
Query: 424 VIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
I RS+A GT + L+M L +I
Sbjct: 123 NIAEVRSKAKIIGTLVGLGGALVMTLYKGPLI 154
>sptr|Q9LPF1|Q9LPF1 T12C22.7 protein (At1g44800/T12C22_7).
Length = 370
Score = 165 bits (417), Expect = 3e-40
Identities = 79/163 (48%), Positives = 113/163 (69%)
Frame = +1
Query: 52 GTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFER 231
G+ K+ P +A++ LQFG+AGM++I++ S + GM H+VL YR+VVA +VMAPFAL FER
Sbjct: 4 GSMEKIKPILAIISLQFGYAGMYIITMVSFKHGMDHWVLATYRHVVATVVMAPFALMFER 63
Query: 232 KTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILR 411
K RPKM+L +F ++LALG+LEP++DQN Y+G+ +TSAS++SA TN LPA+TF+ A+I R
Sbjct: 64 KIRPKMTLAIFWRLLALGILEPLMDQNLYYIGLKNTSASYTSAFTNALPAVTFILALIFR 123
Query: 412 MERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXSXH 540
+E + S A GT TV ++M L I + H
Sbjct: 124 LETVNFRKVHSVAKVVGTVITVGGAMIMTLYKGPAIEIVKAAH 166
>sptr|Q8GWV8|Q8GWV8 Putative nodulin (At4g08290).
Length = 384
Score = 164 bits (416), Expect = 4e-40
Identities = 83/158 (52%), Positives = 116/158 (73%)
Frame = +1
Query: 46 LAGTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWF 225
++ T K+ PY+ M+FLQFG AG +++ +A+L QG + YV++VYRN+VAA+V+A FAL F
Sbjct: 4 VSATMHKLRPYLLMIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAHFALIF 63
Query: 226 ERKTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAII 405
ERK RPKM+L V KI+ALG LEPVLDQ F Y+G+N TSA+++SA+ NILP++TF+ A I
Sbjct: 64 ERKVRPKMTLSVLWKIMALGFLEPVLDQGFGYLGMNMTSATYTSAIMNILPSVTFIIAWI 123
Query: 406 LRMERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
LRME++ I RS+A GT + L+M L +I
Sbjct: 124 LRMEKVNIAEVRSKAKIIGTLVGLGGALVMTLYKGPLI 161
>sptr|Q9SUF0|Q9SUF0 Nodulin-like protein.
Length = 368
Score = 163 bits (413), Expect = 1e-39
Identities = 79/159 (49%), Positives = 113/159 (71%)
Frame = +1
Query: 64 KVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRP 243
K+ P +A++ LQFG+AGM++I++ S + GM+H++L YR+VVA IV+APFAL ERK RP
Sbjct: 3 KLKPIIAIISLQFGYAGMYIITMVSFKHGMNHWILATYRHVVATIVIAPFALILERKIRP 62
Query: 244 KMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERI 423
KM+ P+F++ILALG LEP+LDQN Y+G+ +TSA++SSA N LPA+TF+ A+I R+E +
Sbjct: 63 KMTWPLFLRILALGFLEPLLDQNLYYIGMKATSATYSSAFVNALPAITFIMAVIFRIETV 122
Query: 424 VIXXRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXSXH 540
+ RS A GTA TV ++M L I + H
Sbjct: 123 NLKKTRSLAKVIGTAITVGGAMVMTLYKGPAIELFKTAH 161
>sptr|Q8LEI9|Q8LEI9 Nodulin protein, putative.
Length = 365
Score = 162 bits (410), Expect = 2e-39
Identities = 77/159 (48%), Positives = 111/159 (69%)
Frame = +1
Query: 64 KVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRP 243
K+ P +A++ LQFG+AGM++I++ S + GM H+VL YR++VA +VMAPFAL FERK RP
Sbjct: 3 KIKPILAIISLQFGYAGMYIITMVSFKHGMDHWVLATYRHIVATVVMAPFALMFERKIRP 62
Query: 244 KMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERI 423
KM+L +F ++LALG+LEP++DQN Y+G+ +TSAS++SA TN LPA+TF+ A+I R+E +
Sbjct: 63 KMTLAIFWRLLALGILEPLMDQNLYYIGLKNTSASYTSAFTNALPAVTFILALIFRLETV 122
Query: 424 VIXXRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXSXH 540
S A GT TV ++M L I + H
Sbjct: 123 NFRKVHSVAKVVGTVITVGGAMIMTLYKGPAIEIVKAAH 161
>sptr|Q8VZ54|Q8VZ54 Putative nodulin protein.
Length = 389
Score = 160 bits (406), Expect = 6e-39
Identities = 79/156 (50%), Positives = 108/156 (69%)
Frame = +1
Query: 52 GTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFER 231
G + PY+AM+ +QFG+AGM++I++ SL+ GM+HYVL VYR+ +A V+APFAL+ ER
Sbjct: 4 GLMNSLKPYLAMISMQFGYAGMYIITMVSLKHGMNHYVLAVYRHAIATAVIAPFALFHER 63
Query: 232 KTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILR 411
K RPKM+ +F++I LG +EPVLDQN Y+G+ TSA+F+SA N+LPA+TFV AI R
Sbjct: 64 KIRPKMTFRIFLQIALLGFIEPVLDQNLYYVGMTYTSATFASATANVLPAITFVLAINFR 123
Query: 412 MERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
+E + RS A GT TV LLM L I+
Sbjct: 124 LESVNFKKVRSIAKVVGTVITVSGALLMTLYKGPIV 159
>sptr|Q8LRB6|Q8LRB6 Nodulin-like protein.
Length = 398
Score = 158 bits (400), Expect = 3e-38
Identities = 81/163 (49%), Positives = 106/163 (65%)
Frame = +1
Query: 52 GTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFER 231
G K PY AM+ LQFG+AGM +I+ SL GMSHYVLVVYR+ A I +APFAL ER
Sbjct: 8 GFLEKAKPYFAMICLQFGYAGMNVITKVSLNHGMSHYVLVVYRHAFATISIAPFALLLER 67
Query: 232 KTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILR 411
K RPKM+ VF++I L LL PV+DQNF Y G+ T +F+ A++NILPA+TFV A+I R
Sbjct: 68 KVRPKMTWSVFLQIFVLALLGPVIDQNFYYAGLKFTGPTFACAMSNILPAMTFVMAVIFR 127
Query: 412 MERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXSXH 540
ME++ + R QA GT TV ++M L ++ + H
Sbjct: 128 MEKVDLKKVRCQAKVAGTLVTVAGAMMMTLYKGPLMQMAWTSH 170
>sptr|Q9SFE7|Q9SFE7 T26F17.11.
Length = 391
Score = 158 bits (400), Expect = 3e-38
Identities = 80/158 (50%), Positives = 109/158 (68%), Gaps = 2/158 (1%)
Frame = +1
Query: 52 GTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFER 231
G + PY+AM+ +QFG+AGM++I++ SL+ GM+HYVL VYR+ +A V+APFAL+ ER
Sbjct: 4 GLMNSLKPYLAMISMQFGYAGMYIITMVSLKHGMNHYVLAVYRHAIATAVIAPFALFHER 63
Query: 232 KTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILR 411
K RPKM+ +F++I LG +EPVLDQN Y+G+ TSA+F+SA N+LPA+TFV AII R
Sbjct: 64 KIRPKMTFRIFLQIALLGFIEPVLDQNLYYVGMTYTSATFASATANVLPAITFVLAIIFR 123
Query: 412 --MERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
+E + RS A GT TV LLM L I+
Sbjct: 124 FKLESVNFKKVRSIAKVVGTVITVSGALLMTLYKGPIV 161
>sptr|Q9FL41|Q9FL41 MtN21 nodulin protein-like.
Length = 402
Score = 155 bits (393), Expect = 2e-37
Identities = 75/149 (50%), Positives = 105/149 (70%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMS 252
PY AM+ LQFG+AGM +I+ SL GMSHYVLVVYR+ +A V+APFA +FERK +PK++
Sbjct: 18 PYFAMISLQFGYAGMNIITKISLNTGMSHYVLVVYRHAIATAVIAPFAFFFERKAQPKIT 77
Query: 253 LPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIX 432
+F+++ LGLL PV+DQNF YMG+ TS +FS A++N+LPA+TF+ A++ RME + +
Sbjct: 78 FSIFMQLFILGLLGPVIDQNFYYMGLKYTSPTFSCAMSNMLPAMTFILAVLFRMEMLDLK 137
Query: 433 XRRSQAXXXGTAXTVXXXLLMILXXXXII 519
QA GT TV +LM + I+
Sbjct: 138 KLWCQAKIAGTVVTVAGAMLMTIYKGPIV 166
>sptr|Q9ZUS1|Q9ZUS1 Nodulin-like protein (Putative nodulin protein).
Length = 380
Score = 154 bits (389), Expect = 6e-37
Identities = 77/146 (52%), Positives = 105/146 (71%)
Frame = +1
Query: 64 KVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRP 243
K P+++MV LQ G AGM ++S A L +GMS+YVLVVYR+ VA IVMAPFA +F++K RP
Sbjct: 12 KARPFISMVVLQVGLAGMDILSKAVLNKGMSNYVLVVYRHAVATIVMAPFAFYFDKKVRP 71
Query: 244 KMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERI 423
KM+L +F KI LGLLEPV+DQN Y+G+ T+A+F++A+ N+LPA+TFV A I +ER+
Sbjct: 72 KMTLMIFFKISLLGLLEPVIDQNLYYLGMKYTTATFATAMYNVLPAITFVLAYIFGLERV 131
Query: 424 VIXXRRSQAXXXGTAXTVXXXLLMIL 501
+ RS GT TV ++M L
Sbjct: 132 KLRCIRSTGKVVGTLATVGGAMIMTL 157
>sptr|Q9LXX8|Q9LXX8 Nodulin-like protein.
Length = 377
Score = 152 bits (384), Expect = 2e-36
Identities = 79/154 (51%), Positives = 102/154 (66%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMS 252
PY AMV LQFG+AGM L++ L +GMSHYVLV YRN A +APFAL ERK RPKM+
Sbjct: 11 PYFAMVCLQFGYAGMNLVTKVVLDRGMSHYVLVAYRNAFATAAIAPFALLSERKVRPKMT 70
Query: 253 LPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIX 432
P+F++I L LL P++DQN Y G+ TS +F+ A+TNI+PALTF+ +II RME++ +
Sbjct: 71 FPIFMQIFVLALLGPLIDQNLYYAGLKLTSPTFAGAVTNIVPALTFIISIICRMEKVEMR 130
Query: 433 XRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXS 534
R QA GT V +LMIL +I S
Sbjct: 131 KVRFQAKVVGTLVIVVGAMLMILFKIPLITFLRS 164
>sptr|O24091|O24091 MtN21 protein.
Length = 394
Score = 151 bits (381), Expect = 5e-36
Identities = 77/143 (53%), Positives = 101/143 (70%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMS 252
PY AM+ LQFG+AGM +I+ SL GMSHYVLVVYR+ A I +APFA+ FE K +PK++
Sbjct: 18 PYFAMILLQFGYAGMNIITKLSLNGGMSHYVLVVYRHAFATIAIAPFAIIFEWKDQPKIT 77
Query: 253 LPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIX 432
VF++IL L LL PV+DQNF Y G+ TS +FS A++N+LPA+TFV A++ RME + +
Sbjct: 78 FSVFMQILLLALLGPVIDQNFYYAGLKLTSPTFSCAMSNMLPAMTFVMAVLCRMEIVNLK 137
Query: 433 XRRSQAXXXGTAXTVXXXLLMIL 501
R QA GT TV +LM L
Sbjct: 138 KLRCQAKVIGTILTVAGAMLMTL 160
>sptr|Q8LBH3|Q8LBH3 Nodulin-like protein.
Length = 373
Score = 149 bits (375), Expect = 3e-35
Identities = 78/154 (50%), Positives = 101/154 (65%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMS 252
PY AMV LQFG+AGM L++ L +GMSHYVLV YRN A +APFAL ERK RPKM+
Sbjct: 7 PYFAMVCLQFGYAGMNLVTKVVLDRGMSHYVLVAYRNAFATAAIAPFALLSERKVRPKMT 66
Query: 253 LPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIX 432
P+F++I L LL P++DQN Y + TS +F+ A+TNI+PALTF+ +II RME++ +
Sbjct: 67 FPIFMQIFVLALLGPLIDQNLYYACLKLTSPTFAGAVTNIVPALTFIISIICRMEKVEMR 126
Query: 433 XRRSQAXXXGTAXTVXXXLLMILXXXXIIXXXXS 534
R QA GT V +LMIL +I S
Sbjct: 127 KVRFQAKVVGTLVIVVGAMLMILFKIPLITFLRS 160
>sptr|Q8LE45|Q8LE45 Nodulin-like protein.
Length = 362
Score = 148 bits (373), Expect = 4e-35
Identities = 75/139 (53%), Positives = 100/139 (71%)
Frame = +1
Query: 85 MVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMSLPVF 264
MV LQ G AGM ++S A L +GMS+YVLVVYR+ VA IVMAPFA +F++K RPKM+L +F
Sbjct: 1 MVVLQVGLAGMDILSKAVLNKGMSNYVLVVYRHAVATIVMAPFAFYFDKKVRPKMTLMIF 60
Query: 265 IKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIXXRRS 444
KI LGLLEPV+DQN Y+G+ T+A+F++A+ N+LPA+TFV A I +ER+ + RS
Sbjct: 61 FKISLLGLLEPVIDQNLYYLGMKYTTATFATAMYNVLPAITFVLAYIFGLERVKLRCIRS 120
Query: 445 QAXXXGTAXTVXXXLLMIL 501
GT TV ++M L
Sbjct: 121 TGKVVGTLATVGGAMIMTL 139
>sptr|O80638|O80638 Nodulin-like protein (At2g39510).
Length = 374
Score = 147 bits (371), Expect = 7e-35
Identities = 74/155 (47%), Positives = 106/155 (68%)
Frame = +1
Query: 55 TWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERK 234
TW+ P++ +V LQFG+AG+ +I+ +L QGMS +VL YR++VA I +APFA + +RK
Sbjct: 5 TWK---PFITVVSLQFGYAGLSIIAKFALNQGMSPHVLASYRHIVATIFIAPFAYFLDRK 61
Query: 235 TRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRM 414
RPKM+L +F KIL LGLLEP +DQN Y G+ TSA+F++A+TN+LPA F+ A I R+
Sbjct: 62 IRPKMTLSIFFKILLLGLLEPTIDQNLYYTGMKYTSATFTAAMTNVLPAFAFIMAWIFRL 121
Query: 415 ERIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
E++ + SQA GT TV +LM + +I
Sbjct: 122 EKVNVKKIHSQAKILGTIVTVGGAMLMTVVKGPLI 156
>sptr|O22204|O22204 Putative integral membrane protein nodulin.
Length = 400
Score = 144 bits (363), Expect = 6e-34
Identities = 73/142 (51%), Positives = 96/142 (67%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKMS 252
PY AMV LQFG+AGM L++ L +GMSHYVLV YRN A +APFAL ERK R KM+
Sbjct: 11 PYFAMVCLQFGYAGMNLVTKTVLDRGMSHYVLVAYRNAFATAAIAPFALLSERKVRSKMT 70
Query: 253 LPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVIX 432
P+F++I L LL PV+DQN Y+G+ TS +FSSA++NI+PA+T + A + RME++ +
Sbjct: 71 FPIFMRIFLLALLGPVIDQNLYYIGLKLTSPTFSSAVSNIVPAITIILATLFRMEKVEMR 130
Query: 433 XRRSQAXXXGTAXTVXXXLLMI 498
R GT TV +LMI
Sbjct: 131 KVRCLVKVMGTLVTVVGSILMI 152
>sptr|Q9FNA5|Q9FNA5 Similarity to MtN21 (Nodulin-like protein).
Length = 377
Score = 128 bits (322), Expect = 4e-29
Identities = 62/154 (40%), Positives = 99/154 (64%)
Frame = +1
Query: 58 WRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKT 237
+ + P++A+VF+Q +A M +++ +L +GMS +VLV YR VA+ ++ PFAL ER T
Sbjct: 3 FERARPFIAIVFIQCLYALMSIVAKLALNKGMSPHVLVAYRMAVASALITPFALILERNT 62
Query: 238 RPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRME 417
RPK++ + ++I L L EPV++QN Y G+ T+A+F+SAL N LPA+TF+ A + ++E
Sbjct: 63 RPKLTFKILLQIAILSLFEPVVEQNLYYSGMKLTTATFTSALCNALPAMTFIMACVFKLE 122
Query: 418 RIVIXXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
++ I R SQA GT + +LM +I
Sbjct: 123 KVTIERRHSQAKLVGTMVAIGGAMLMTFVKGNVI 156
>sptr|Q84TV0|Q84TV0 Putative nodulin protein.
Length = 374
Score = 115 bits (288), Expect = 3e-25
Identities = 59/150 (39%), Positives = 87/150 (58%)
Frame = +1
Query: 70 MPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKM 249
MP++A V LQ G+AGM + S +L GM +LV YR + A + +APFA + ERKTRPK+
Sbjct: 6 MPFLANVLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLERKTRPKL 65
Query: 250 SLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVI 429
++P+ +I L +Q F ++G+ +T+A+ + AL N+LPA TF+ A I R E + I
Sbjct: 66 TMPILFQIFLCSLTGATANQVFYFVGLQNTTATIACALNNVLPAATFLLAAICRQEAVGI 125
Query: 430 XXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
QA GT V +L+ II
Sbjct: 126 KKASGQAKVIGTLACVGGAMLLSFYHGHII 155
>sptr|Q84TV1|Q84TV1 Nodulin-like protein.
Length = 376
Score = 115 bits (287), Expect = 4e-25
Identities = 59/150 (39%), Positives = 87/150 (58%)
Frame = +1
Query: 70 MPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKM 249
MP++A V LQ G+AGM + S +L GM +LV YR + A + +APFA + ERKTRPK+
Sbjct: 6 MPFLANVLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLERKTRPKL 65
Query: 250 SLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVI 429
++P+ +I L +Q F ++G+ +T+A+ + AL N+LPA TF+ A I R E + I
Sbjct: 66 TMPILFQIFLCSLTGATANQVFYFVGLQNTTATIACALNNVLPAATFLLAAICRQEAVGI 125
Query: 430 XXRRSQAXXXGTAXTVXXXLLMILXXXXII 519
QA GT V +L+ II
Sbjct: 126 KKASGQAKVIGTLVCVGGAMLLSFYHGHII 155
>sptr|Q93VI2|Q93VI2 Putative MtN21.
Length = 380
Score = 111 bits (278), Expect = 4e-24
Identities = 59/147 (40%), Positives = 89/147 (60%), Gaps = 2/147 (1%)
Frame = +1
Query: 61 RKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTR 240
R+ +P +AMV +Q GFAGM ++S +L GMS YVL+ YRN++AA+ +APFA +FER T+
Sbjct: 3 RESLPTLAMVMVQLGFAGMNVVSKLALDTGMSPYVLIAYRNIIAAVFLAPFAYYFERYTK 62
Query: 241 PKMSL--PVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRM 414
M + V ++I + L+Q ++G+ ST+ + + AL+N LPALTF A RM
Sbjct: 63 SGMVITKKVLVQIFFSSIFGATLNQVLYFVGLKSTTPTVACALSNTLPALTFAMAAAFRM 122
Query: 415 ERIVIXXRRSQAXXXGTAXTVXXXLLM 495
E + + QA GT V ++M
Sbjct: 123 ESVRLSAAAGQAKVFGTVVCVGGSMIM 149
>sptr|Q94AP3|Q94AP3 Hypothetical protein (Putative nodulin protein).
Length = 389
Score = 110 bits (276), Expect = 8e-24
Identities = 57/156 (36%), Positives = 97/156 (62%)
Frame = +1
Query: 34 DRMELAGTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPF 213
+R L G K+ ++AM+ LQFG+AG ++S A+L G+S V VYRN++A +++ PF
Sbjct: 7 NRRSLWGVPEKLQLHIAMLTLQFGYAGFHVVSRAALNMGISKLVFPVYRNIIALLLLLPF 66
Query: 214 ALWFERKTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFV 393
A + E+K RP ++L I+ L L+ +Q F +G+++TS +F+S++ N +PA+TF+
Sbjct: 67 AYFLEKKERPAITLNFLIQFFFLALIGITANQGFYLLGLDNTSPTFASSMQNSVPAITFL 126
Query: 394 NAIILRMERIVIXXRRSQAXXXGTAXTVXXXLLMIL 501
A +LR+E++ I R + GTA V ++ L
Sbjct: 127 MAALLRIEKVRINRRDGISKILGTALCVAGASVITL 162
>sptr|Q8GXB4|Q8GXB4 Putative nodulin protein N21.
Length = 374
Score = 101 bits (252), Expect = 5e-21
Identities = 50/143 (34%), Positives = 84/143 (58%)
Frame = +1
Query: 67 VMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPK 246
++P++AMV +Q G+AGM + S ++ GM +LV YR + A I P A + ERKTRPK
Sbjct: 6 MLPFLAMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLERKTRPK 65
Query: 247 MSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIV 426
++L + +++ + +Q ++G+ ++S + + ALTN+LPA+TF+ A I R E +
Sbjct: 66 ITLRILVQVFFCSITGATGNQVLYFVGLQNSSPTIACALTNLLPAVTFLLAAIFRQETVG 125
Query: 427 IXXRRSQAXXXGTAXTVXXXLLM 495
I QA GT V +++
Sbjct: 126 IKKASGQAKVIGTLVCVIGAMVL 148
>sptr|Q9FTZ6|Q9FTZ6 Putative nodulin (P0494A10.12 protein).
Length = 374
Score = 100 bits (248), Expect = 1e-20
Identities = 50/158 (31%), Positives = 90/158 (56%)
Frame = +1
Query: 28 GSDRMELAGTWRKVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMA 207
G D+ A KV ++ ++ LQF AG ++S A+L G+S V +VYRN+++ ++A
Sbjct: 2 GRDQAAAAVMHEKVKLFIGVLALQFLLAGFHIVSRAALNMGISKIVFIVYRNLISLALLA 61
Query: 208 PFALWFERKTRPKMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALT 387
PFA + E+K RP ++ + ++ L L +Q F +G+ S +++SA+ N +PA+T
Sbjct: 62 PFAYFLEKKDRPPLTFSLLVEFFLLALCGITANQGFYLLGLYHLSPTYASAIQNTVPAIT 121
Query: 388 FVNAIILRMERIVIXXRRSQAXXXGTAXTVXXXLLMIL 501
F A +LR+E++ + R A GT ++ ++ L
Sbjct: 122 FAMAAVLRLEQVDLGKRHGVAKVVGTVVSIGGATVITL 159
>sptr|Q9LV20|Q9LV20 Nodulin-like protein.
Length = 383
Score = 95.9 bits (237), Expect = 3e-19
Identities = 49/146 (33%), Positives = 86/146 (58%)
Frame = +1
Query: 64 KVMPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRP 243
KV +A++ LQF FAG ++S +L G+S V VYRN++A +++ PFA +FE+K RP
Sbjct: 32 KVKLVVALITLQFCFAGFHIVSRVALNIGVSKVVYPVYRNLLALLLIGPFAYFFEKKERP 91
Query: 244 KMSLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERI 423
+++ + + L L+ +Q F +G+ + +F+SA+ N +PA+TF+ A LR+E I
Sbjct: 92 PLTISLLAQFFFLALIGITANQGFYLLGLYYATPTFASAMQNSVPAITFIMACALRLEHI 151
Query: 424 VIXXRRSQAXXXGTAXTVXXXLLMIL 501
+ + A GT ++ ++ L
Sbjct: 152 DLVRKHGVAKVLGTLVSIGGATVITL 177
>sptrnew|CAD41943|CAD41943 OSJNBa0070M12.20 protein.
Length = 412
Score = 95.9 bits (237), Expect = 3e-19
Identities = 58/144 (40%), Positives = 82/144 (56%), Gaps = 1/144 (0%)
Frame = +1
Query: 73 PYMAMVFLQFGFAGMFLI-SVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKM 249
P AMV +Q FAG+ + +A + GM VLV YR + A+ V+AP A + ERK R KM
Sbjct: 12 PVAAMVVVQVVFAGVNIFYKLAVVCDGMDMRVLVAYRYLFASAVLAPLAYFVERKNRTKM 71
Query: 250 SLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVI 429
+ V + GL L QN G+ TSA+F++A+TN++PA+TFV A++ R ER+ I
Sbjct: 72 TWRVLMLSFVCGLSGGSLAQNLYISGMKLTSATFATAMTNLIPAVTFVLAVLCRYERLAI 131
Query: 430 XXRRSQAXXXGTAXTVXXXLLMIL 501
QA GT V +L+ L
Sbjct: 132 RTVAGQAKVAGTLLGVGGAMLLTL 155
>sptr|Q8H691|Q8H691 OSJNBa0004I20.6 protein.
Length = 354
Score = 95.1 bits (235), Expect = 4e-19
Identities = 46/142 (32%), Positives = 83/142 (58%)
Frame = +1
Query: 70 MPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKM 249
MP ++MV A M + +L QGM+ VL+ +R +VA + + P A + ERKTRPK
Sbjct: 5 MPTVSMVATNVVIAIMTALIKQALNQGMNRLVLITFRQMVATVFLGPIAYFKERKTRPKF 64
Query: 250 SLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVI 429
+ +F+ + G+L PVL Q +++G+ T+A+F++ N+LP +TF+ +++ R E + +
Sbjct: 65 TTEIFVYMFLSGMLGPVLLQYTLFVGLEFTTATFAATFGNLLPVVTFLISLVFRFEALNV 124
Query: 430 XXRRSQAXXXGTAXTVXXXLLM 495
R A GT ++ +++
Sbjct: 125 KSRSGSAKISGTLVSLSGAMML 146
>sptr|Q9SNG9|Q9SNG9 ESTs AU055729(S20023).
Length = 354
Score = 95.1 bits (235), Expect = 4e-19
Identities = 46/142 (32%), Positives = 83/142 (58%)
Frame = +1
Query: 70 MPYMAMVFLQFGFAGMFLISVASLRQGMSHYVLVVYRNVVAAIVMAPFALWFERKTRPKM 249
MP ++MV A M + +L QGM+ VL+ +R +VA + + P A + ERKTRPK
Sbjct: 5 MPTVSMVATNVVIAIMTALIKQALNQGMNRLVLITFRQMVATVFLGPIAYFKERKTRPKF 64
Query: 250 SLPVFIKILALGLLEPVLDQNFIYMGVNSTSASFSSALTNILPALTFVNAIILRMERIVI 429
+ +F+ + G+L PVL Q +++G+ T+A+F++ N+LP +TF+ +++ R E + +
Sbjct: 65 TTEIFVYMFLSGMLGPVLLQYTLFVGLEFTTATFAATFGNLLPVVTFLISLVFRFEALNV 124
Query: 430 XXRRSQAXXXGTAXTVXXXLLM 495
R A GT ++ +++
Sbjct: 125 KSRSGSAKISGTLVSLSGAMML 146
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 364,531,255
Number of Sequences: 1272877
Number of extensions: 7547741
Number of successful extensions: 23715
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 22737
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 23578
length of database: 406,886,216
effective HSP length: 115
effective length of database: 260,505,361
effective search space used: 16932848465
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

