BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QAZ3f05.yg.3.1
(1626 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
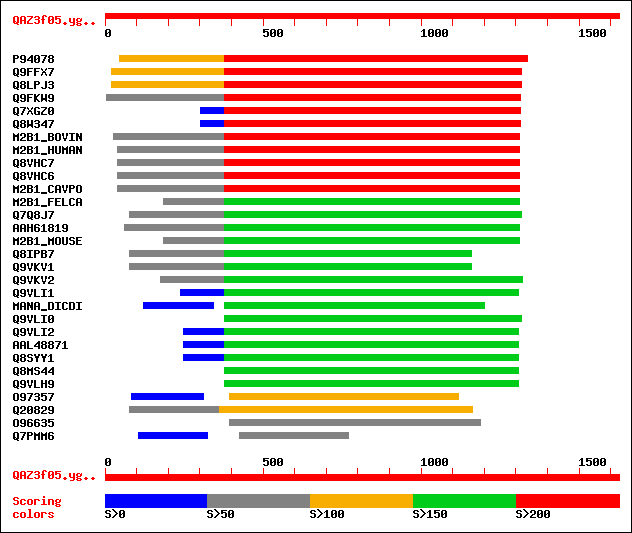
sptr|P94078|P94078 Alpha-mannosidase (AT3G26720/MLJ15_12). 428 e-149
sptr|Q9FFX7|Q9FFX7 Alpha-mannosidase. 431 e-144
sptr|Q8LPJ3|Q8LPJ3 Alpha-mannosidase. 431 e-144
sptr|Q9FKW9|Q9FKW9 Alpha-mannosidase. 375 e-121
sptr|Q7XGZ0|Q7XGZ0 Putative alpha-mannosidase. 309 2e-89
sptr|Q8W347|Q8W347 Putative alpha-mannosidase. 309 2e-89
sw|Q29451|M2B1_BOVIN Lysosomal alpha-mannosidase precursor (EC 3... 206 5e-62
sw|O00754|M2B1_HUMAN Lysosomal alpha-mannosidase precursor (EC 3... 201 8e-61
sptr|Q8VHC7|Q8VHC7 Lysosomal alpha-mannosidase (EC 3.2.1.24). 205 3e-60
sptr|Q8VHC6|Q8VHC6 Lysosomal alpha-mannosidase (EC 3.2.1.24). 205 3e-60
sw|Q8VHC8|M2B1_CAVPO Lysosomal alpha-mannosidase precursor (EC 3... 205 3e-60
sw|O46432|M2B1_FELCA Lysosomal alpha-mannosidase precursor (EC 3... 198 2e-59
sptr|Q7Q8J7|Q7Q8J7 AgCP15371 (Fragment). 190 3e-58
sptrnew|AAH61819|AAH61819 Similar to mannosidase 2, alpha B1. 188 5e-57
sw|O09159|M2B1_MOUSE Lysosomal alpha-mannosidase precursor (EC 3... 191 1e-56
sptr|Q8IPB7|Q8IPB7 CG6206-PB. 176 3e-54
sptr|Q9VKV1|Q9VKV1 CG6206 protein (BCDNA.GH02419). 176 3e-54
sptr|Q9VKV2|Q9VKV2 CG5322 protein. 165 4e-49
sptr|Q9VLI1|Q9VLI1 CG9465 protein. 171 5e-47
sw|P34098|MANA_DICDI Lysosomal alpha-mannosidase precursor (EC 3... 169 1e-44
sptr|Q9VLI0|Q9VLI0 CG9466 protein (GH02475P). 181 4e-44
sptr|Q9VLI2|Q9VLI2 CG9463 protein. 165 8e-44
sptrnew|AAL48871|AAL48871 RE28991p (Fragment). 162 4e-43
sptr|Q8SYY1|Q8SYY1 RE28991p. 162 4e-43
sptr|Q8MS44|Q8MS44 RE08556p. 166 1e-39
sptr|Q9VLH9|Q9VLH9 CG9468 protein. 164 4e-39
sptr|O97357|O97357 Lysosomal acid alpha-mannosidase precursor. 134 2e-33
sptr|Q20829|Q20829 Hypothetical protein. 121 3e-26
sptr|O96635|O96635 Lysosomal alpha mannosidase (Fragment). 100 7e-20
sptr|Q7PMM6|Q7PMM6 ENSANGP00000014255. 75 2e-15
>sptr|P94078|P94078 Alpha-mannosidase (AT3G26720/MLJ15_12).
Length = 1019
Score = 428 bits (1100), Expect(2) = e-149
Identities = 207/324 (63%), Positives = 260/324 (80%), Gaps = 4/324 (1%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VYK K+ E EF +GPIP DDG KE+ T++ T M TN TFYTDS+GRDFIKRIRD+R++
Sbjct: 693 VYKGKNHAEIEFTIGPIPADDGISKEIITKLTTTMKTNGTFYTDSNGRDFIKRIRDFRTD 752
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W ++V+QP+AGNYYP+NLGIY++D + ELSVLVDR++GGSS+++GQIELMLHRR+ HDD
Sbjct: 753 WDLQVYQPVAGNYYPLNLGIYMQDKTSELSVLVDRAVGGSSLENGQIELMLHRRMQHDDI 812
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQDG 917
+GV E LNETVCL + C+GL IQGK+YV+ID G+GA+WRRTFGQEIYSPLL+AFTEQ+G
Sbjct: 813 RGVGEILNETVCLPEGCKGLTIQGKFYVQIDKPGDGAKWRRTFGQEIYSPLLIAFTEQEG 872
Query: 918 GNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASID 1097
+W NSH FSA + +YSLP NVA+LTLQELE+G VLLR AHL+E GED + S +A ++
Sbjct: 873 DSWINSHKTTFSAFEPSYSLPKNVALLTLQELENGEVLLRLAHLFEVGEDSEYSVMAKVE 932
Query: 1098 LKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDPSKLVVEL 1277
LK++F KKI +V ETSLS NQE+A MEK+RL WK +GSA +E V RG VD KLVVEL
Sbjct: 933 LKKLFHNKKIREVKETSLSGNQEKAEMEKRRLIWKVEGSAGEE-VKRGEAVDAEKLVVEL 991
Query: 1278 GPMEIRTFIVSFDH----ISDKQQ 1337
PMEIRT ++ FD + DK+Q
Sbjct: 992 VPMEIRTLLIKFDDQIEMVGDKEQ 1015
Score = 125 bits (315), Expect(2) = e-149
Identities = 60/116 (51%), Positives = 83/116 (71%), Gaps = 1/116 (0%)
Frame = +2
Query: 47 STQYSPQGSESSNLQVGQGNLKLQYNEAG-KLSLYSDSKTMVQANFEQKYKYYIGQDGNG 223
S Y GS + N++VGQGNLKL+Y+E G K++ + +K V A EQ Y YYIG +G
Sbjct: 584 SASYVTSGSMNQNVEVGQGNLKLRYSEEGVKITRHLSTKNQVTA--EQSYAYYIGSNGTD 641
Query: 224 SDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
DPQASGAY+FRP+G +PI + + LT+++GP+ DEVHQ++NSWI QITR +G+
Sbjct: 642 KDPQASGAYVFRPDGVLPIKSKEEAQLTIVQGPLFDEVHQELNSWISQITRVYKGK 697
>sptr|Q9FFX7|Q9FFX7 Alpha-mannosidase.
Length = 1030
Score = 431 bits (1109), Expect(2) = e-144
Identities = 209/316 (66%), Positives = 257/316 (81%), Gaps = 3/316 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VYK K+ +E EFIVG IP+DDG GKEV T+I +++ +NKTFYTDSSGRD+IKRIRDYRS+
Sbjct: 702 VYKGKEHVEVEFIVGNIPIDDGIGKEVVTQISSSLKSNKTFYTDSSGRDYIKRIRDYRSD 761
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK++V+QPIAGNYYP+N GIY++D KE SV+VDR+ GGSSI DGQ+ELMLHRRLL DD
Sbjct: 762 WKLDVNQPIAGNYYPINHGIYLQDSKKEFSVMVDRAFGGSSIVDGQVELMLHRRLLLDDS 821
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQDG 917
+GVAE LNETVC+ +C GL IQGKYY +IDP GEGA+WRRTFGQEIYSPLLLAF +QD
Sbjct: 822 RGVAENLNETVCVQDKCTGLTIQGKYYYRIDPYGEGAKWRRTFGQEIYSPLLLAFAQQDD 881
Query: 918 GNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASID 1097
G + A FS +D +YSLPDNVA+LTLQEL+DG+VLLR AHLYE EDK+LS +AS++
Sbjct: 882 GKPMSFGAASFSGIDPSYSLPDNVALLTLQELDDGNVLLRLAHLYEVEEDKELSGVASVE 941
Query: 1098 LKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAA---DEKVVRGGPVDPSKLV 1268
LK++FP KKIGK+ E SLSANQER+ MEKKRL WK +G + ++K RG +DP KL
Sbjct: 942 LKKLFPGKKIGKLTEMSLSANQERSTMEKKRLVWKVEGEGSYGEEKKAKRGREIDPRKLE 1001
Query: 1269 VELGPMEIRTFIVSFD 1316
+EL PMEIRT ++ +
Sbjct: 1002 MELYPMEIRTVLIHLE 1017
Score = 105 bits (263), Expect(2) = e-144
Identities = 57/129 (44%), Positives = 80/129 (62%), Gaps = 5/129 (3%)
Frame = +2
Query: 20 KKSAHISSKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYK 196
KK+ SSKS + E S + +G G+LKL ++ + G Y + +T + +Q +
Sbjct: 578 KKTDGYSSKSYVSNILKGEQSIINIGHGHLKLSFSTDQGTAINYVNGRTSMTEPVKQTFS 637
Query: 197 YYIGQDGNGSD----PQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIY 364
YY +G+ PQ SGAY+FRPNGT PI+ +GQV LTV+ GP++DEVHQQIN WI
Sbjct: 638 YYSAYNGSNDKEPLIPQNSGAYVFRPNGTFPINPEGQVPLTVIHGPLVDEVHQQINPWIS 697
Query: 365 QITRRLQGE 391
QITR +G+
Sbjct: 698 QITRVYKGK 706
>sptr|Q8LPJ3|Q8LPJ3 Alpha-mannosidase.
Length = 1024
Score = 431 bits (1109), Expect(2) = e-144
Identities = 209/316 (66%), Positives = 257/316 (81%), Gaps = 3/316 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VYK K+ +E EFIVG IP+DDG GKEV T+I +++ +NKTFYTDSSGRD+IKRIRDYRS+
Sbjct: 696 VYKGKEHVEVEFIVGNIPIDDGIGKEVVTQISSSLKSNKTFYTDSSGRDYIKRIRDYRSD 755
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK++V+QPIAGNYYP+N GIY++D KE SV+VDR+ GGSSI DGQ+ELMLHRRLL DD
Sbjct: 756 WKLDVNQPIAGNYYPINHGIYLQDSKKEFSVMVDRAFGGSSIVDGQVELMLHRRLLLDDS 815
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQDG 917
+GVAE LNETVC+ +C GL IQGKYY +IDP GEGA+WRRTFGQEIYSPLLLAF +QD
Sbjct: 816 RGVAENLNETVCVQDKCTGLTIQGKYYYRIDPYGEGAKWRRTFGQEIYSPLLLAFAQQDD 875
Query: 918 GNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASID 1097
G + A FS +D +YSLPDNVA+LTLQEL+DG+VLLR AHLYE EDK+LS +AS++
Sbjct: 876 GKPMSFGAASFSGIDPSYSLPDNVALLTLQELDDGNVLLRLAHLYEVEEDKELSGVASVE 935
Query: 1098 LKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAA---DEKVVRGGPVDPSKLV 1268
LK++FP KKIGK+ E SLSANQER+ MEKKRL WK +G + ++K RG +DP KL
Sbjct: 936 LKKLFPGKKIGKLTEMSLSANQERSTMEKKRLVWKVEGEGSYGEEKKAKRGREIDPRKLE 995
Query: 1269 VELGPMEIRTFIVSFD 1316
+EL PMEIRT ++ +
Sbjct: 996 MELYPMEIRTVLIHLE 1011
Score = 105 bits (263), Expect(2) = e-144
Identities = 57/129 (44%), Positives = 80/129 (62%), Gaps = 5/129 (3%)
Frame = +2
Query: 20 KKSAHISSKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYK 196
KK+ SSKS + E S + +G G+LKL ++ + G Y + +T + +Q +
Sbjct: 572 KKTDGYSSKSYVSNILKGEQSIINIGHGHLKLSFSTDQGTAINYVNGRTSMTEPVKQTFS 631
Query: 197 YYIGQDGNGSD----PQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIY 364
YY +G+ PQ SGAY+FRPNGT PI+ +GQV LTV+ GP++DEVHQQIN WI
Sbjct: 632 YYSAYNGSNDKEPLIPQNSGAYVFRPNGTFPINPEGQVPLTVIHGPLVDEVHQQINPWIS 691
Query: 365 QITRRLQGE 391
QITR +G+
Sbjct: 692 QITRVYKGK 700
>sptr|Q9FKW9|Q9FKW9 Alpha-mannosidase.
Length = 1047
Score = 375 bits (964), Expect(2) = e-121
Identities = 186/315 (59%), Positives = 238/315 (75%), Gaps = 3/315 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGN--GKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYR 551
+YKEK+ E EF +GPI V G+ GKE+ T +VT+M T K FYTDS+GRDF+KR+RD R
Sbjct: 722 LYKEKEHAEFEFTIGPISVGKGHLTGKEIITRMVTDMTTAKEFYTDSNGRDFLKRVRDNR 781
Query: 552 SEWKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHD 731
++W +EV++PIAGNYYP+NLG+Y++D ELSVLVDR+ GG+SIKDG+IELMLHRR D
Sbjct: 782 TDWHLEVNEPIAGNYYPLNLGMYIKDEKAELSVLVDRATGGASIKDGEIELMLHRRTSMD 841
Query: 732 DGKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQ 911
D +GV E+L ETVC++ C GL I+G YYV I+ GEG RWRR GQEIYSPLL+AF +
Sbjct: 842 DSRGVEESLVETVCVNDTCAGLTIRGNYYVSINKVGEGGRWRRETGQEIYSPLLMAFAHE 901
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
+ W S+ K AMD Y+LP N+A++TL+EL+ G+VLLR AHLYEAGED D S +A
Sbjct: 902 NKEKWKASNTVKGYAMDHLYTLPQNIALITLEELDLGNVLLRLAHLYEAGEDSDYSKIAK 961
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAAD-EKVVRGGPVDPSKLV 1268
++LK++F K I +V E SLSANQE+ M K+++KWK +G A +RGGPVD S LV
Sbjct: 962 VELKKLFSGKMIKEVTEMSLSANQEKVKM-KEKMKWKVEGEAEQPSSPLRGGPVDKSTLV 1020
Query: 1269 VELGPMEIRTFIVSF 1313
VELGPMEIRTF+V F
Sbjct: 1021 VELGPMEIRTFVVQF 1035
Score = 84.3 bits (207), Expect(2) = e-121
Identities = 47/141 (33%), Positives = 78/141 (55%), Gaps = 17/141 (12%)
Frame = +2
Query: 5 ASANLKKSAHISSKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANF 181
+ A+ + S + S SP + + ++G GNLK+ ++ ++G+L +S+T
Sbjct: 584 SKASAQGSNNHKHSSVMLSPMNNTT---EIGPGNLKMVFSSDSGRLERMYNSRTGADIKV 640
Query: 182 EQKYKYYIGQDGNGSDPQASGAYIFRPNGTV--PIST--------------DGQVSLTVL 313
+Q Y +Y G+ DPQ SGAYIFRPNG++ P+S+ + Q L ++
Sbjct: 641 DQNYFWYASNVGDAKDPQVSGAYIFRPNGSLAYPVSSSKICTVTSAFIGNGNVQSKLQIV 700
Query: 314 RGPILDEVHQQINSWIYQITR 376
RGP++DEVHQQ + W+ Q+ R
Sbjct: 701 RGPLIDEVHQQFSPWVAQVVR 721
>sptr|Q7XGZ0|Q7XGZ0 Putative alpha-mannosidase.
Length = 438
Score = 309 bits (792), Expect(2) = 2e-89
Identities = 156/315 (49%), Positives = 213/315 (67%), Gaps = 3/315 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGN--GKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYR 551
+YK K+ E E+ +GPIPVDD + GKEV T + TNM TNK FYTDS+GRDF++R+R++R
Sbjct: 163 LYKNKEHAEVEYTIGPIPVDDDDDIGKEVVTRLTTNMATNKIFYTDSNGRDFLERVRNHR 222
Query: 552 SEWKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHD 731
+W + + QP+AGNYYPVN GIYV DG ELSVLVD ++G SSI+DGQIE+MLHRRL D
Sbjct: 223 DDWDLNLSQPVAGNYYPVNQGIYVADGKYELSVLVDHAVGASSIQDGQIEVMLHRRLSAD 282
Query: 732 DGKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQ 911
DG+GV E LNE VC+D++C+GL+
Sbjct: 283 DGRGVGEPLNEVVCVDQKCDGLV------------------------------------- 305
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
D +W ++++AK S ++ YSLPDNVA++TLQ L+DG+ LLR AHL++A ED S +A
Sbjct: 306 DERSWKSNNIAKASTVEGNYSLPDNVAIITLQSLDDGTTLLRLAHLFQAQEDTQYSVMAK 365
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQG-SAADEKVVRGGPVDPSKLV 1268
++L+++F ++ I + ETSLSANQ+++ E K+L W+ G S D ++GGPVD LV
Sbjct: 366 VELRKLFGKRIIKDLTETSLSANQKKS--EMKKLNWRVTGESKTDPAPLKGGPVDSHALV 423
Query: 1269 VELGPMEIRTFIVSF 1313
VELGPMEIRTF++ F
Sbjct: 424 VELGPMEIRTFLLKF 438
Score = 44.3 bits (103), Expect(2) = 2e-89
Identities = 17/25 (68%), Positives = 22/25 (88%)
Frame = +2
Query: 302 LTVLRGPILDEVHQQINSWIYQITR 376
L V+ GP++DEVHQQ +SWIYQ+TR
Sbjct: 138 LKVIHGPLVDEVHQQFSSWIYQVTR 162
>sptr|Q8W347|Q8W347 Putative alpha-mannosidase.
Length = 438
Score = 309 bits (792), Expect(2) = 2e-89
Identities = 156/315 (49%), Positives = 213/315 (67%), Gaps = 3/315 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGN--GKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYR 551
+YK K+ E E+ +GPIPVDD + GKEV T + TNM TNK FYTDS+GRDF++R+R++R
Sbjct: 163 LYKNKEHAEVEYTIGPIPVDDDDDIGKEVVTRLTTNMATNKIFYTDSNGRDFLERVRNHR 222
Query: 552 SEWKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHD 731
+W + + QP+AGNYYPVN GIYV DG ELSVLVD ++G SSI+DGQIE+MLHRRL D
Sbjct: 223 DDWDLNLSQPVAGNYYPVNQGIYVADGKYELSVLVDHAVGASSIQDGQIEVMLHRRLSAD 282
Query: 732 DGKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQ 911
DG+GV E LNE VC+D++C+GL+
Sbjct: 283 DGRGVGEPLNEVVCVDQKCDGLV------------------------------------- 305
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
D +W ++++AK S ++ YSLPDNVA++TLQ L+DG+ LLR AHL++A ED S +A
Sbjct: 306 DERSWKSNNIAKASTVEGNYSLPDNVAIITLQSLDDGTTLLRLAHLFQAQEDTQYSVMAK 365
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQG-SAADEKVVRGGPVDPSKLV 1268
++L+++F ++ I + ETSLSANQ+++ E K+L W+ G S D ++GGPVD LV
Sbjct: 366 VELRKLFGKRIIKDLTETSLSANQKKS--EMKKLNWRVTGESKTDPAPLKGGPVDSHALV 423
Query: 1269 VELGPMEIRTFIVSF 1313
VELGPMEIRTF++ F
Sbjct: 424 VELGPMEIRTFLLKF 438
Score = 44.3 bits (103), Expect(2) = 2e-89
Identities = 17/25 (68%), Positives = 22/25 (88%)
Frame = +2
Query: 302 LTVLRGPILDEVHQQINSWIYQITR 376
L V+ GP++DEVHQQ +SWIYQ+TR
Sbjct: 138 LKVIHGPLVDEVHQQFSSWIYQVTR 162
>sw|Q29451|M2B1_BOVIN Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 999
Score = 206 bits (524), Expect(2) = 5e-62
Identities = 124/314 (39%), Positives = 176/314 (56%), Gaps = 3/314 (0%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y + LE E+ VGPIPV DG GKEV + T + T FYTDS+GR+ ++R R+YR
Sbjct: 692 LYPRQRHLELEWTVGPIPVGDGWGKEVISRFDTALATRGLFYTDSNGREILERRRNYRPT 751
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ +P+AGNYYPVN IY+ DG+ +L+VL DRS GGSS++DG +ELM+HRRLL DD
Sbjct: 752 WKLNQTEPVAGNYYPVNSRIYITDGNMQLTVLTDRSQGGSSLRDGSLELMVHRRLLKDDA 811
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKID-PQGEGARWRRTFGQEIYSPLLLAFTEQD 914
+GV E LN K+ GL ++G++ V +D + AR R E+ +P ++ +
Sbjct: 812 RGVGEPLN------KEGSGLWVRGRHLVLLDKKETAAARHRLQAEMEVLAPQVV-LAQGG 864
Query: 915 GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDLSSLA 1088
G + + LP +V +LTL ++LLR H + GED ++LSS
Sbjct: 865 GARYRLEKAPRTQFSGLRRELPPSVRLLTLARWGPETLLLRLEHQFAVGEDSGRNLSSPV 924
Query: 1089 SIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDPSKLV 1268
++DL +F I + ET+L+ANQ A RL+W P P
Sbjct: 925 TLDLTNLFSAFTITNLRETTLAANQLLA--YASRLQWTTDTGPTPHP----SPSRPVSAT 978
Query: 1269 VELGPMEIRTFIVS 1310
+ L PMEIRTF+ S
Sbjct: 979 ITLQPMEIRTFLAS 992
Score = 55.8 bits (133), Expect(2) = 5e-62
Identities = 33/118 (27%), Positives = 55/118 (46%), Gaps = 1/118 (0%)
Frame = +2
Query: 26 SAHISSKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYY 202
S + S+ PQ S S +L + L+ +++ G L + + + Q + +Y
Sbjct: 574 SIYSVSQMPNQRPQKSWSRDLVIQNEYLRARFDPNTGLLMELENLEQNLLLPVRQAFYWY 633
Query: 203 IGQDGNGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITR 376
GN QASGAYIFRPN P+ +++ ++ EVHQ ++W Q+ R
Sbjct: 634 NASTGNNLSSQASGAYIFRPNQNKPLFVSHWAQTHLVKASLVQEVHQNFSAWCSQVVR 691
>sw|O00754|M2B1_HUMAN Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1010
Score = 201 bits (510), Expect(2) = 8e-61
Identities = 129/315 (40%), Positives = 182/315 (57%), Gaps = 4/315 (1%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y + LE E+ VGPIPV D GKEV + T + T FYTDS+GR+ ++R RDYR
Sbjct: 702 LYPGQRHLELEWSVGPIPVGDTWGKEVISRFDTPLETKGRFYTDSNGREILERRRDYRPT 761
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ +P+AGNYYPVN IY+ DG+ +L+VL DRS GGSS++DG +ELM+HRRLL DDG
Sbjct: 762 WKLNQTEPVAGNYYPVNTRIYITDGNMQLTVLTDRSQGGSSLRDGSLELMVHRRLLKDDG 821
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKID-PQGEGARWRRTFGQEIYSP-LLLAFTEQ 911
+GV+E L E G ++G++ V +D Q A R QE+ +P ++LA
Sbjct: 822 RGVSEPLME------NGSGAWVRGRHLVLLDTAQAAAAGHRLLAEQEVLAPQVVLAPGGG 875
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDLSSL 1085
N +FS + LP +V +LTL VLLR H + GED ++LS+
Sbjct: 876 AAYNLGAPPRTQFSGL--RRDLPPSVHLLTLASWGPEMVLLRLEHQFAVGEDSGRNLSAP 933
Query: 1086 ASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDPSKL 1265
+++L+ +F I ++ ET+L ANQ R A RLKW + +DP+ +
Sbjct: 934 VTLNLRDLFSTFTITRLQETTLVANQLREA--ASRLKWTTNTGPTPHQTPY--QLDPANI 989
Query: 1266 VVELGPMEIRTFIVS 1310
+E PMEIRTF+ S
Sbjct: 990 TLE--PMEIRTFLAS 1002
Score = 57.4 bits (137), Expect(2) = 8e-61
Identities = 31/118 (26%), Positives = 58/118 (49%), Gaps = 1/118 (0%)
Frame = +2
Query: 41 SKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYYIGQDG 217
+++ Q P+ S S L + +++ ++ + G L + + Q + +Y G
Sbjct: 589 ARAPQPIPRRSWSPALTIENEHIRATFDPDTGLLMEIMNMNQQLLLPVRQTFFWYNASIG 648
Query: 218 NGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
+ QASGAYIFRPN P+ + +++ P++ EVHQ ++W Q+ R G+
Sbjct: 649 DNESDQASGAYIFRPNQQKPLPVSRWAQIHLVKTPLVQEVHQNFSAWCSQVVRLYPGQ 706
>sptr|Q8VHC7|Q8VHC7 Lysosomal alpha-mannosidase (EC 3.2.1.24).
Length = 1007
Score = 205 bits (521), Expect(2) = 3e-60
Identities = 128/318 (40%), Positives = 184/318 (57%), Gaps = 7/318 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y + LE E+ VGPIPV D GKE+ + T + T F+TDS+GR+ ++R RDYR
Sbjct: 697 LYSGQRHLELEWTVGPIPVGDKWGKEIISRFDTPLETGGVFFTDSNGREVLERRRDYRPS 756
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ +P+AGNYYPVN IY+ DG +L+VL DRS GGSS+ DG +ELM+HRRLL DDG
Sbjct: 757 WKLNQTEPVAGNYYPVNSRIYITDGKMQLTVLTDRSQGGSSMSDGSLELMVHRRLLKDDG 816
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQ-EIYSPLLLAFTEQD 914
+GV EAL E G ++G++ + +D E A R + E+ +P L+ Q
Sbjct: 817 RGVGEALQE------PGSGGWVRGRHLLLLDTAREAAAEHRLLAEKELLAPQLVLAPGQG 870
Query: 915 GGNWANSHVA----KFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDL 1076
+ H A +FS + LP +V +LTL ++LLR H + GED ++L
Sbjct: 871 PSYHHDHHEAVPRKQFSGL--RRQLPPSVRLLTLARWGPDTLLLRLEHQFALGEDSSRNL 928
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDP 1256
S ++DL+ +F I ++ ET+L+ANQ RA+ RLKW + V +DP
Sbjct: 929 SLPVTLDLQDLFSTFTITRLQETTLAANQLRAS--ASRLKWTTEIDPISRPAV--PRLDP 984
Query: 1257 SKLVVELGPMEIRTFIVS 1310
S + ++ PMEIRTF+ S
Sbjct: 985 SSITLQ--PMEIRTFVAS 1000
Score = 51.2 bits (121), Expect(2) = 3e-60
Identities = 31/118 (26%), Positives = 52/118 (44%), Gaps = 1/118 (0%)
Frame = +2
Query: 41 SKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYYIGQDG 217
++S PQ S L + L+ ++ + G LS+ + Q + +Y G
Sbjct: 584 TRSQHSRPQKYSSPVLSIKNEYLRASFHPDTGLLSMIEVLDRKLTLPVNQAFFWYNASVG 643
Query: 218 NGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
+ QASGAYIFRP+ P +++ ++ EVHQ +W Q+ R G+
Sbjct: 644 DKRSSQASGAYIFRPSQQWPFPVSHLARTRLVKTALVQEVHQNFTAWCSQVVRLYSGQ 701
>sptr|Q8VHC6|Q8VHC6 Lysosomal alpha-mannosidase (EC 3.2.1.24).
Length = 1007
Score = 205 bits (521), Expect(2) = 3e-60
Identities = 128/318 (40%), Positives = 184/318 (57%), Gaps = 7/318 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y + LE E+ VGPIPV D GKE+ + T + T F+TDS+GR+ ++R RDYR
Sbjct: 697 LYSGQRHLELEWTVGPIPVGDKWGKEIISRFDTPLETGGVFFTDSNGREVLERRRDYRPS 756
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ +P+AGNYYPVN IY+ DG +L+VL DRS GGSS+ DG +ELM+HRRLL DDG
Sbjct: 757 WKLNQTEPVAGNYYPVNSRIYITDGKMQLTVLTDRSQGGSSMSDGSLELMVHRRLLKDDG 816
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQ-EIYSPLLLAFTEQD 914
+GV EAL E G ++G++ + +D E A R + E+ +P L+ Q
Sbjct: 817 RGVGEALQE------PGSGGWVRGRHLLLLDTAREAAAEHRLLAEKELLAPQLVLAPGQG 870
Query: 915 GGNWANSHVA----KFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDL 1076
+ H A +FS + LP +V +LTL ++LLR H + GED ++L
Sbjct: 871 PSYHHDHHEAVPRKQFSGL--RRQLPPSVRLLTLARWGPDTLLLRLEHQFALGEDSSRNL 928
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDP 1256
S ++DL+ +F I ++ ET+L+ANQ RA+ RLKW + V +DP
Sbjct: 929 SLPVTLDLQDLFSTFTITRLQETTLAANQLRAS--ASRLKWTTEIDPISRPAV--PRLDP 984
Query: 1257 SKLVVELGPMEIRTFIVS 1310
S + ++ PMEIRTF+ S
Sbjct: 985 SSITLQ--PMEIRTFVAS 1000
Score = 51.2 bits (121), Expect(2) = 3e-60
Identities = 31/118 (26%), Positives = 52/118 (44%), Gaps = 1/118 (0%)
Frame = +2
Query: 41 SKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYYIGQDG 217
++S PQ S L + L+ ++ + G LS+ + Q + +Y G
Sbjct: 584 TRSQHSRPQKYSSPVLSIKNEYLRASFHPDTGLLSMIEVLDRKLTLPVNQAFFWYNASVG 643
Query: 218 NGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
+ QASGAYIFRP+ P +++ ++ EVHQ +W Q+ R G+
Sbjct: 644 DKRSSQASGAYIFRPSQQWPFPVSHLARTRLVKTALVQEVHQNFTAWCSQVVRLYSGQ 701
>sw|Q8VHC8|M2B1_CAVPO Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1007
Score = 205 bits (521), Expect(2) = 3e-60
Identities = 128/318 (40%), Positives = 184/318 (57%), Gaps = 7/318 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y + LE E+ VGPIPV D GKE+ + T + T F+TDS+GR+ ++R RDYR
Sbjct: 697 LYSGQRHLELEWTVGPIPVGDKWGKEIISRFDTPLETGGVFFTDSNGREVLERRRDYRPS 756
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ +P+AGNYYPVN IY+ DG +L+VL DRS GGSS+ DG +ELM+HRRLL DDG
Sbjct: 757 WKLNQTEPVAGNYYPVNSRIYITDGKMQLTVLTDRSQGGSSMSDGSLELMVHRRLLKDDG 816
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQ-EIYSPLLLAFTEQD 914
+GV EAL E G ++G++ + +D E A R + E+ +P L+ Q
Sbjct: 817 RGVGEALQE------PGSGGWVRGRHLLLLDTAREAAAEHRLLAEKELLAPQLVLAPGQG 870
Query: 915 GGNWANSHVA----KFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDL 1076
+ H A +FS + LP +V +LTL ++LLR H + GED ++L
Sbjct: 871 PSYHHDHHEAVPRKQFSGL--RRQLPPSVRLLTLARWGPDTLLLRLEHQFALGEDSSRNL 928
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDP 1256
S ++DL+ +F I ++ ET+L+ANQ RA+ RLKW + V +DP
Sbjct: 929 SLPVTLDLQDLFSTFTITRLQETTLAANQLRAS--ASRLKWTTEIDPISRPAV--PRLDP 984
Query: 1257 SKLVVELGPMEIRTFIVS 1310
S + ++ PMEIRTF+ S
Sbjct: 985 SSITLQ--PMEIRTFVAS 1000
Score = 51.2 bits (121), Expect(2) = 3e-60
Identities = 31/118 (26%), Positives = 52/118 (44%), Gaps = 1/118 (0%)
Frame = +2
Query: 41 SKSTQYSPQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYYIGQDG 217
++S PQ S L + L+ ++ + G LS+ + Q + +Y G
Sbjct: 584 TRSQHSRPQKYSSPVLSIKNEYLRASFHPDTGLLSMIEVLDRKLTLPVNQAFFWYNASVG 643
Query: 218 NGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
+ QASGAYIFRP+ P +++ ++ EVHQ +W Q+ R G+
Sbjct: 644 DKRSSQASGAYIFRPSQQWPFPVSHLARTRLVKTALVQEVHQNFTAWCSQVVRLYSGQ 701
>sw|O46432|M2B1_FELCA Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1007
Score = 198 bits (504), Expect(2) = 2e-59
Identities = 129/316 (40%), Positives = 184/316 (58%), Gaps = 5/316 (1%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ + LE E+ VGPIPV DG GKE+ + T + T FYTDS+GR+ ++R RDYR
Sbjct: 701 LYRGQRHLELEWTVGPIPVGDGWGKEIISRFDTVLETKGLFYTDSNGREILERRRDYRPT 760
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
WK+ + +AGNYYPVN IY+ DG+ +L+VL DRS GGSS++DG +ELM+HRRLL DDG
Sbjct: 761 WKLNQTETVAGNYYPVNSRIYIRDGNMQLTVLTDRSQGGSSLRDGSMELMVHRRLLKDDG 820
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQ-EIYSPLLLAFTEQD 914
+GV EAL E G ++G++ V +D A R + E+ +P ++
Sbjct: 821 RGVGEALLEDGL------GRWVRGRHLVLLDKVRTAATGHRLQAEKEVLTPQVV--LAPG 872
Query: 915 GGNWANSHVA---KFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDK-DLSS 1082
GG + VA +FS + LP +V +LTL + ++LLR H + GED +LSS
Sbjct: 873 GGAPYHLKVAPRKQFSGL--RRELPPSVHLLTLARWDQKTLLLRLEHQFAVGEDSGNLSS 930
Query: 1083 LASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVDPSK 1262
++DL +F I + ET+L ANQ RA+ RLKW + + +DP+
Sbjct: 931 PVTLDLTDLFSAFTITYLQETTLVANQLRAS--ASRLKWTP--NTGPTPLPSPSRLDPA- 985
Query: 1263 LVVELGPMEIRTFIVS 1310
+ L PMEIRTF+ S
Sbjct: 986 -TITLQPMEIRTFLAS 1000
Score = 55.1 bits (131), Expect(2) = 2e-59
Identities = 24/69 (34%), Positives = 37/69 (53%)
Frame = +2
Query: 185 QKYKYYIGQDGNGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIY 364
Q + +Y GN Q SGAYIFRPN P+ +++ P++ EVHQ ++W
Sbjct: 637 QAFYWYNASVGNNLSTQVSGAYIFRPNQEKPLMVSHWAQTRLVKTPLVQEVHQNFSAWCS 696
Query: 365 QITRRLQGE 391
Q+ R +G+
Sbjct: 697 QVVRLYRGQ 705
>sptr|Q7Q8J7|Q7Q8J7 AgCP15371 (Fragment).
Length = 1175
Score = 190 bits (483), Expect(2) = 3e-58
Identities = 124/347 (35%), Positives = 188/347 (54%), Gaps = 33/347 (9%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VY ++ +E E++VGPIPV+DG GKE+ + T ++ F+TD++GR+ ++R+R++R
Sbjct: 835 VYADESHVEFEWMVGPIPVEDGVGKEIVSRFYTAAQSSGVFWTDANGREMMRRVRNHRDT 894
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W +++ + I+GNYYPV I +ED + L+VL DR+ GGSS++DG +ELM+HRRLLHDD
Sbjct: 895 WNVDLEEKISGNYYPVTAKIALEDENLRLAVLNDRAQGGSSLEDGSLELMVHRRLLHDDA 954
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKI---DPQGEGARWRRTFGQ-EIYSPLLLAFT 905
GV EALNE +GL+ +GK++V P R F Q + P L F+
Sbjct: 955 FGVEEALNEKAF----GQGLVARGKHWVVFGAKKPTSPTPEARERFLQNRVLLPNWLFFS 1010
Query: 906 ---EQDGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDL 1076
E +W + +SA+ + SLP NV +LT + ++ S+L+RF HL E ED
Sbjct: 1011 DVGEVKYEDWQKQYTNIYSAL--SLSLPLNVHLLTFEPWKENSILVRFEHLLEKDEDPMY 1068
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGS-------AADEKVV 1235
S +++ VF + I +V E +L+ANQ R + RLK+K S D
Sbjct: 1069 SKPVRFNIQDVFRQFSIEEVREMTLAANQLRE--DSTRLKFKPDPSYIMYSSIKRDVSTP 1126
Query: 1236 RGGPVDPSKLV-------------------VELGPMEIRTFIVSFDH 1319
P P+ L + L PMEIRTF+ ++
Sbjct: 1127 LPSPSPPNVLAGRSPLMDELSRNVADDGFEIMLKPMEIRTFVFQLEY 1173
Score = 58.9 bits (141), Expect(2) = 3e-58
Identities = 37/102 (36%), Positives = 53/102 (51%), Gaps = 2/102 (1%)
Frame = +2
Query: 77 SSNLQVGQGNLKLQYNEAGKLSLYSDSKTMVQANFEQKYKYYIGQDGNGSD--PQASGAY 250
S + +G L + ++ G LS + V Q + YY G GN + ++SGAY
Sbjct: 736 SQEVTIGNKYLNVSFDSNGFLSTITIDG--VTNRLRQTFVYYEGALGNNEEFRNRSSGAY 793
Query: 251 IFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITR 376
IFRPNGT T+ V L V++G + EVHQ + WI Q+ R
Sbjct: 794 IFRPNGTEKTVTEN-VQLKVVKGGTVQEVHQVFSEWISQVVR 834
>sptrnew|AAH61819|AAH61819 Similar to mannosidase 2, alpha B1.
Length = 1009
Score = 188 bits (477), Expect(2) = 5e-57
Identities = 125/320 (39%), Positives = 178/320 (55%), Gaps = 9/320 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ + LE E+ VGPIPV D GKEV + T M T F+TDS+GR+ +KR D+R
Sbjct: 703 LYEGQRHLELEWTVGPIPVKDDWGKEVISRFNTPMRTRGQFFTDSNGREILKRRDDFRPT 762
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W + +P+AGNYYPVN IY+ DG +L+VL DRS GGSS+ DG +ELM+HRRLL DD
Sbjct: 763 WTLNQTEPVAGNYYPVNTRIYITDGHMQLTVLTDRSQGGSSLLDGSLELMVHRRLLVDDE 822
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGE-GARWRRTFGQEIYSPLLLAFTEQD 914
+GVAE L ET DK ++G++ V + + AR R QE+ +P ++
Sbjct: 823 RGVAEPLLETDTGDK------VRGRHLVILSSVSDAAARHRLLAEQEVLAPQVV--LAHG 874
Query: 915 GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGED--KDLSSLA 1088
G + +S K LP V +LTL +LLR H + ED ++LSS
Sbjct: 875 GSSPYHSQAPKMQFSALRRELPPQVHLLTLARWGPKMLLLRLEHQFAVKEDSNRNLSSPV 934
Query: 1089 SIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPVD---PS 1259
+++L+ +F I + ET+L+ANQ + + RLKW + GP+ PS
Sbjct: 935 TLNLQNLFKTFTINYLQETTLAANQPLSRV--SRLKW----------MTDTGPISYPAPS 982
Query: 1260 KL---VVELGPMEIRTFIVS 1310
+L + L PM+IRTF+ S
Sbjct: 983 RLDPTSITLQPMQIRTFLAS 1002
Score = 57.4 bits (137), Expect(2) = 5e-57
Identities = 30/111 (27%), Positives = 57/111 (51%), Gaps = 1/111 (0%)
Frame = +2
Query: 62 PQGSESSNLQVGQGNLKLQYN-EAGKLSLYSDSKTMVQANFEQKYKYYIGQDGNGSDPQA 238
P+ S+S L + ++ ++ + G L + + + Q + +Y G+ PQA
Sbjct: 597 PKKSKSRVLVIENKYIRATFDSDTGLLRKIENLEQNISLPVRQGFFWYNASAGDEESPQA 656
Query: 239 SGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
SGAYIFRP+ P+ +T+++ ++ EVHQ ++W Q+ R +G+
Sbjct: 657 SGAYIFRPSHRKPLPVSHWAQVTLVKTNLVQEVHQNFSAWCSQVIRLYEGQ 707
>sw|O09159|M2B1_MOUSE Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1013
Score = 191 bits (486), Expect(2) = 1e-56
Identities = 129/321 (40%), Positives = 180/321 (56%), Gaps = 10/321 (3%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+YK + LE E+ VGPIPV D GKEV + T M T F+TDS+GR+ +KR DYR
Sbjct: 704 LYKGQRHLELEWTVGPIPVRDDWGKEVISRFDTPMKTKGQFFTDSNGREILKRRDDYRPT 763
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W + +P+AGNYYPVN IY+ DG +L+VL DRS GGSS++DG +ELM+HRRLL DD
Sbjct: 764 WTLNQTEPVAGNYYPVNTRIYITDGQMQLTVLTDRSQGGSSLQDGSLELMVHRRLLVDDD 823
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGE-GARWRRTFGQEIYSP-LLLAFTEQ 911
+GV+E L ET DK ++G++ V + + AR R QE+ +P ++L+
Sbjct: 824 RGVSEPLLETDTGDK------VRGRHLVLLSSVSDAAARHRLLAEQEVLAPQVVLSLGGS 877
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKD--LSSL 1085
+ + +FS + LP V +LTL +LLR H + ED D LSS
Sbjct: 878 SPYHSRATPKTQFSGL--RQELPPQVHLLTLARWGPKMLLLRLEHQFALKEDSDRNLSSP 935
Query: 1086 ASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKVVRGGPV---DP 1256
+++++ +F I + ET+L+ANQ + RLKW + GP +P
Sbjct: 936 VTLNVQNLFQTFTINYLQETTLAANQPLS--RASRLKW----------MTNTGPTSFPEP 983
Query: 1257 SKL---VVELGPMEIRTFIVS 1310
SKL V L PMEIRTF+ S
Sbjct: 984 SKLDPTSVTLKPMEIRTFLAS 1004
Score = 52.4 bits (124), Expect(2) = 1e-56
Identities = 24/69 (34%), Positives = 39/69 (56%)
Frame = +2
Query: 185 QKYKYYIGQDGNGSDPQASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIY 364
Q + +Y G+ QASGAYIFRPN PI +++++ ++ EVHQ ++W
Sbjct: 640 QGFFWYNASVGDEESSQASGAYIFRPNVGKPIPVSRWAQISLVKTALVQEVHQNFSAWCS 699
Query: 365 QITRRLQGE 391
Q+ R +G+
Sbjct: 700 QVIRLYKGQ 708
>sptr|Q8IPB7|Q8IPB7 CG6206-PB.
Length = 1080
Score = 176 bits (447), Expect(2) = 3e-54
Identities = 101/269 (37%), Positives = 154/269 (57%), Gaps = 8/269 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VY + + E E++VGPIP+DDG GKEV T +++ ++ F TDS+GR+ IKR ++R
Sbjct: 699 VYNKDSYAEFEWLVGPIPIDDGIGKEVITRFNSDIASDGIFRTDSNGREMIKRKINHRDT 758
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W +++++ +AGNYYP+ I VED + +++L DR+ GGSS+KDG +ELM+HRRLL DD
Sbjct: 759 WSVKINEAVAGNYYPITTKIDVEDDTARMAILTDRAQGGSSLKDGSLELMVHRRLLKDDA 818
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYV----KIDPQGEGARWRRTFGQ-EIYSPLLLAF 902
GV EALNET + +GLI +GK+++ D +G + Q E P F
Sbjct: 819 FGVGEALNET----EYGDGLIARGKHHLFFGKSTDREGVSLKGIERLTQLEKLLPTWKFF 874
Query: 903 TEQD---GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKD 1073
+ + W + FS + + LP V +LTL+ + +L+RF H+ E GED
Sbjct: 875 SNMEDYSADEWQTAFTNIFSGI--SLVLPKPVHLLTLEPWHENQLLVRFEHIMENGEDAS 932
Query: 1074 LSSLASIDLKRVFPEKKIGKVIETSLSAN 1160
S ++K V + + ET+L N
Sbjct: 933 YSQPVQFNVKNVLSAFDVEGIRETTLDGN 961
Score = 59.7 bits (143), Expect(2) = 3e-54
Identities = 38/104 (36%), Positives = 63/104 (60%), Gaps = 4/104 (3%)
Frame = +2
Query: 77 SSNLQVGQGNLKLQYNEAGKLS-LYSDSKTMVQANFEQKYKYYIGQDGNGSD--PQASGA 247
SS +G +++L ++ G LS + +D T + + Q++ +Y G GN ++ ++SGA
Sbjct: 599 SSVTVIGNSHIQLGFDTNGFLSEVTADGLTRLVS---QEFLFYEGAVGNNAEFLNRSSGA 655
Query: 248 YIFRPN-GTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITR 376
YIFRPN + +TD QV + V +G ++ EVHQ+ N WI Q+ R
Sbjct: 656 YIFRPNENKIHFATD-QVEIEVYKGDLVHEVHQKFNDWISQVVR 698
>sptr|Q9VKV1|Q9VKV1 CG6206 protein (BCDNA.GH02419).
Length = 1080
Score = 176 bits (447), Expect(2) = 3e-54
Identities = 101/269 (37%), Positives = 154/269 (57%), Gaps = 8/269 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
VY + + E E++VGPIP+DDG GKEV T +++ ++ F TDS+GR+ IKR ++R
Sbjct: 699 VYNKDSYAEFEWLVGPIPIDDGIGKEVITRFNSDIASDGIFRTDSNGREMIKRKINHRDT 758
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
W +++++ +AGNYYP+ I VED + +++L DR+ GGSS+KDG +ELM+HRRLL DD
Sbjct: 759 WSVKINEAVAGNYYPITTKIDVEDDTARMAILTDRAQGGSSLKDGSLELMVHRRLLKDDA 818
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYV----KIDPQGEGARWRRTFGQ-EIYSPLLLAF 902
GV EALNET + +GLI +GK+++ D +G + Q E P F
Sbjct: 819 FGVGEALNET----EYGDGLIARGKHHLFFGKSTDREGVSLKGIERLTQLEKLLPTWKFF 874
Query: 903 TEQD---GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKD 1073
+ + W + FS + + LP V +LTL+ + +L+RF H+ E GED
Sbjct: 875 SNMEDYSADEWQTAFTNIFSGI--SLVLPKPVHLLTLEPWHENQLLVRFEHIMENGEDAS 932
Query: 1074 LSSLASIDLKRVFPEKKIGKVIETSLSAN 1160
S ++K V + + ET+L N
Sbjct: 933 YSQPVQFNVKNVLSAFDVEGIRETTLDGN 961
Score = 59.7 bits (143), Expect(2) = 3e-54
Identities = 38/104 (36%), Positives = 63/104 (60%), Gaps = 4/104 (3%)
Frame = +2
Query: 77 SSNLQVGQGNLKLQYNEAGKLS-LYSDSKTMVQANFEQKYKYYIGQDGNGSD--PQASGA 247
SS +G +++L ++ G LS + +D T + + Q++ +Y G GN ++ ++SGA
Sbjct: 599 SSVTVIGNSHIQLGFDTNGFLSEVTADGLTRLVS---QEFLFYEGAVGNNAEFLNRSSGA 655
Query: 248 YIFRPN-GTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITR 376
YIFRPN + +TD QV + V +G ++ EVHQ+ N WI Q+ R
Sbjct: 656 YIFRPNENKIHFATD-QVEIEVYKGDLVHEVHQKFNDWISQVVR 698
>sptr|Q9VKV2|Q9VKV2 CG5322 protein.
Length = 950
Score = 165 bits (418), Expect(2) = 4e-49
Identities = 105/326 (32%), Positives = 178/326 (54%), Gaps = 11/326 (3%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y++ + +E E++VGPIP DD GKE+ T +N+ + FYTDS+GR+ ++R R+ R
Sbjct: 641 IYEDVNRVEFEWLVGPIPTDDDVGKEIITRFSSNISSKGKFYTDSNGREILERERNQREH 700
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDG 737
+ ++ + I+GNYYPV I ++D K +++L DR+ GG+S+KDG++ELMLHRRLL+DD
Sbjct: 701 FTPDMSEAISGNYYPVTGQISLQDDEKRITLLNDRAQGGTSLKDGELELMLHRRLLNDDA 760
Query: 738 KGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQDG 917
GV EALNET + GLI +GK Y+ +D +G +R ++ F++ +G
Sbjct: 761 FGVGEALNET----QYGTGLIARGKIYLILDAV-DGKPNQRLLQHQLDQHFWKFFSKSNG 815
Query: 918 GNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASID 1097
N ++ + + +P++V +L+L+ +L+R + G ++ S +
Sbjct: 816 VASVNRNM-----IPDFFGIPESVELLSLEPYSKDQILIRLENFNTEG------NVVSFN 864
Query: 1098 LKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGS-----------AADEKVVRGG 1244
+ +F ++ ET+L N + KR K+ G+ A +
Sbjct: 865 IYPLFESLDGYQIWETTLDGNM--LLEDVKRFKFAQDGTGSIPSSVEYYHAPHNPLTANS 922
Query: 1245 PVDPSKLVVELGPMEIRTFIVSFDHI 1322
++ S VV L PM+IRTFI+ +I
Sbjct: 923 TMNASGFVVTLVPMQIRTFIIQQKYI 948
Score = 53.5 bits (127), Expect(2) = 4e-49
Identities = 27/69 (39%), Positives = 40/69 (57%), Gaps = 2/69 (2%)
Frame = +2
Query: 176 NFEQKYKYYIGQDGNGSDPQ--ASGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQI 349
N +Q + Y G GN + + +SGAY+FRP G + I + +V L+ G + EVHQ +
Sbjct: 573 NIQQTFGIYKGYRGNNGESKNRSSGAYVFRPYGDIEI-VNNKVELSFYNGTKVKEVHQHV 631
Query: 350 NSWIYQITR 376
N WI Q+ R
Sbjct: 632 NEWISQVIR 640
>sptr|Q9VLI1|Q9VLI1 CG9465 protein.
Length = 942
Score = 171 bits (434), Expect(2) = 5e-47
Identities = 111/323 (34%), Positives = 177/323 (54%), Gaps = 13/323 (4%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ +E E++VGPIP+DD G E T + + +N FYTDS+GR+ +KR++D R +
Sbjct: 632 IYEGVSRVELEWMVGPIPIDDKLGSEFVTNFKSEISSNGVFYTDSNGRELMKRVKDKRED 691
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ ++ QP++GN+YPV I ++D +K L +L DRS GG+S++DG +E+++HRR L +D
Sbjct: 692 FVSDLSRQPVSGNFYPVTSRIALQDDTKRLVLLNDRSQGGASLEDGALEMLIHRRHLFND 751
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEG-ARWRRTFGQEIYSPLLLAFTEQ 911
G GV EALNET + +GLI +GK Y+ +D +G R +E++ P F++
Sbjct: 752 GGGVGEALNET----QYGKGLIARGKLYLILDSATDGDTVTERKTEKELFLPFWKFFSKT 807
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
G + ++ LP +V +LTL+ + +L+RF H DK + S
Sbjct: 808 GG-----VEITPSKSLPDFNDLPQSVHLLTLEFFSEQEILIRFEHFL----DKSEGRVIS 858
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQG----------SAADEKVVRG 1241
+++ +F + ET+L N + M KR K+ AQ S A K +
Sbjct: 859 FNIRDIFDSLGGLAIRETTLDGNMPLSDM--KRFKFHAQESGTKPSSVEYSTAQHKPLEA 916
Query: 1242 GPVDPSKL-VVELGPMEIRTFIV 1307
D + L V L PM+IRTFI+
Sbjct: 917 VKSDEASLFAVTLYPMQIRTFII 939
Score = 40.4 bits (93), Expect(2) = 5e-47
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = +2
Query: 239 SGAYIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQG 388
S AY FR +G + + + TV G ++ EVHQ +N WI Q+ R +G
Sbjct: 587 SCAYTFRQDGDIELF-ENDFEFTVYEGSLVKEVHQHVNDWISQVIRIYEG 635
>sw|P34098|MANA_DICDI Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Alpha-D-mannoside mannohydrolase) (Laman).
Length = 1005
Score = 169 bits (427), Expect(2) = 1e-44
Identities = 97/282 (34%), Positives = 156/282 (55%), Gaps = 7/282 (2%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y D LE E I+GPI + DG GKE+ + T +VT++T+Y+DS G + KRI +YR
Sbjct: 699 LYSNADHLEVEEIIGPIDISDGIGKEIVSRYTTTLVTDQTWYSDSQGMEMQKRITNYRPS 758
Query: 558 WKIEVHQPIAGNYYPVNLGIYVEDGSKEL--SVLVDRSIGGSSIKDGQIELMLHRRLLHD 731
W + V QP +GNY PVN Y++D ++ L +++ DRS G +S++DGQ+++M+HRR L D
Sbjct: 759 WNLTVVQPTSGNYVPVNAIAYIQDPNQSLQFTIVTDRSRGCASLRDGQLDMMMHRRTLKD 818
Query: 732 DGKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAF--T 905
DG+GV + +NE+ I+ + D R + PLL F T
Sbjct: 819 DGRGVGQPMNEST--------QIVTTSKLIFHDISSYAQSHYRPAALSLSHPLLPMFTTT 870
Query: 906 EQDGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQEL--EDGSVLLRFAHLYEA-GEDKDL 1076
+Q +W + + +S + S LP+ + + TLQ L +D ++LLR ++Y+ G+D
Sbjct: 871 QQSSNDWNSQYQGVYSPLTSASPLPNGLKIQTLQWLDNQDNTILLRIENIYQIDGQDSQD 930
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWK 1202
++DL +F I E +L+ Q+ + + RLKWK
Sbjct: 931 PQTITLDLSTIFSTITITSATEMNLTGVQKLSNL--SRLKWK 970
Score = 35.4 bits (80), Expect(2) = 1e-44
Identities = 21/76 (27%), Positives = 40/76 (52%), Gaps = 2/76 (2%)
Frame = +2
Query: 122 NEAGKLSLYSDSKTMVQANFEQKYKYYIGQDGNGSDPQASGAYIFRP--NGTVPISTDGQ 295
++ G + ++ + V ++ Q+Y +Y GN Q SGAYIFRP + P + +
Sbjct: 613 SQDGSILSITNKTSGVTSSITQEYIWYNPSVGNDDSAQCSGAYIFRPVEDFAYPYN-NAT 671
Query: 296 VSLTVLRGPILDEVHQ 343
S++++RG I + +
Sbjct: 672 PSVSIIRGEISSSIRR 687
>sptr|Q9VLI0|Q9VLI0 CG9466 protein (GH02475P).
Length = 982
Score = 181 bits (458), Expect = 4e-44
Identities = 114/325 (35%), Positives = 184/325 (56%), Gaps = 12/325 (3%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+ ++K ++E E++VGPIPV++ G EV T + + +N FYTDS+GR+ I+R +D R +
Sbjct: 673 ISEDKPYVEFEWLVGPIPVEEEFGTEVVTIFSSEIASNGVFYTDSNGRELIRREKDKRED 732
Query: 558 WKIEVH-QPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ E+ QP +GNYYP+ I ++D +K L++L DR+ GG+S+KDGQIELMLHRRL+ DD
Sbjct: 733 FTPELAVQPTSGNYYPITSRIALQDSNKRLAILNDRAQGGTSMKDGQIELMLHRRLVRDD 792
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQD 914
G GV EALNE +K + LI +GK ++ ++ E R +E + PL F++
Sbjct: 793 GYGVGEALNE----EKYGQPLIARGKVFLILNAADESTSAEREAEKEFHLPLWKFFSKNT 848
Query: 915 GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASI 1094
G S A ++ S P +V +LTL+ D +LLR + + E K + S
Sbjct: 849 G-----STTAAAKSVPSFDDFPKSVHLLTLEPFNDDEILLRVENFKDHIEGK----VVSF 899
Query: 1095 DLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGS-----------AADEKVVRG 1241
+++ +F ++ ET+L N + M K+ K+ A+GS ++ + +
Sbjct: 900 NIRPIFDYLNGVEIRETTLDGNLPLSDM--KQFKFHAEGSGIRGSEPEYYTSSHKPLSAN 957
Query: 1242 GPVDPSKLVVELGPMEIRTFIVSFD 1316
D ++ V L PM+IRTFI+ +
Sbjct: 958 QTQDAAEFAVTLYPMQIRTFIIKHE 982
>sptr|Q9VLI2|Q9VLI2 CG9463 protein.
Length = 1003
Score = 165 bits (417), Expect(2) = 8e-44
Identities = 108/323 (33%), Positives = 178/323 (55%), Gaps = 13/323 (4%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ + +E E++VGP+P+DD GKE+ T + + + FYTDS+GR+ ++R +D R +
Sbjct: 693 IYEGVNRVEFEWLVGPVPIDDELGKEIVTIFKSGISSGGVFYTDSNGREMMRREKDKRED 752
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ ++ QP++GNYYPV + ++D SK + +L DRS GG+S++DG++E+MLHRR + D
Sbjct: 753 FSPDLSEQPVSGNYYPVTSRMALQDSSKRMVLLNDRSQGGASLEDGRLEMMLHRRHIFAD 812
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGA-RWRRTFGQEIYSPLLLAFTEQ 911
G G AEA+NE + +GLI +GK ++ ++ +GA R +EI+ P F++
Sbjct: 813 GSGAAEAINE----QQFGKGLIARGKLFLYLNAIEDGATASERVAEKEIHLPFWKFFSK- 867
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
N S V K + LP +V +LTL+ +LLR + + E ++ S
Sbjct: 868 --SNNIQSDVTK--TLSDFNDLPQSVHLLTLEPYSKDEILLRLENFLDQTE----GNVVS 919
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKV-----------VR 1238
++++F + ++ ET+L N + M KRLK+ GS V
Sbjct: 920 FSIRQIFDDLGGLEIRETTLDGNLPLSDM--KRLKFHHDGSGPSHSVPEYFTSLHKPLAA 977
Query: 1239 GGPVDPSKLVVELGPMEIRTFIV 1307
D S+ V L PM+IRTFI+
Sbjct: 978 DKTQDASEFSVTLKPMQIRTFII 1000
Score = 36.2 bits (82), Expect(2) = 8e-44
Identities = 18/47 (38%), Positives = 25/47 (53%)
Frame = +2
Query: 248 YIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQG 388
YIFR + + + D TV G + EVHQ +N WI Q+ R +G
Sbjct: 651 YIFRQDADLELVEDA-FDFTVYDGESVKEVHQHVNEWISQVIRIYEG 696
>sptrnew|AAL48871|AAL48871 RE28991p (Fragment).
Length = 1016
Score = 162 bits (411), Expect(2) = 4e-43
Identities = 107/323 (33%), Positives = 177/323 (54%), Gaps = 13/323 (4%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ + +E E++VGP+P+DD GKE+ T + + + FYTDS+GR+ ++R +D R +
Sbjct: 706 IYEGVNRVEFEWLVGPVPIDDELGKEIVTIFKSGISSGGVFYTDSNGREMMRREKDKRED 765
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ ++ QP++GNYY V + ++D SK + +L DRS GG+S++DG++E+MLHRR + D
Sbjct: 766 FSPDLSEQPVSGNYYQVTSRVALQDSSKRMVLLNDRSQGGASLEDGRLEMMLHRRHIFAD 825
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGA-RWRRTFGQEIYSPLLLAFTEQ 911
G G AEA+NE + +GLI +GK ++ ++ +GA R +EI+ P F++
Sbjct: 826 GSGAAEAINE----QQFGKGLIARGKLFLYLNAIEDGATASERVAEKEIHLPFWKFFSK- 880
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
N S V K + LP +V +LTL+ +LLR + + E ++ S
Sbjct: 881 --SNNIQSDVTK--TLSDFNDLPQSVHLLTLEPYSKDEILLRLENFLDQTE----GNVVS 932
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKV-----------VR 1238
++++F + ++ ET+L N + M KRLK+ GS V
Sbjct: 933 FSIRQIFDDLGGLEIRETTLDGNLPLSDM--KRLKFHHDGSGPSHSVPEYFTSLHKPLAA 990
Query: 1239 GGPVDPSKLVVELGPMEIRTFIV 1307
D S+ V L PM+IRTFI+
Sbjct: 991 DKTQDASEFSVTLKPMQIRTFII 1013
Score = 36.2 bits (82), Expect(2) = 4e-43
Identities = 18/47 (38%), Positives = 25/47 (53%)
Frame = +2
Query: 248 YIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQG 388
YIFR + + + D TV G + EVHQ +N WI Q+ R +G
Sbjct: 664 YIFRQDADLELVEDA-FDFTVYDGESVKEVHQHVNEWISQVIRIYEG 709
>sptr|Q8SYY1|Q8SYY1 RE28991p.
Length = 1003
Score = 162 bits (411), Expect(2) = 4e-43
Identities = 107/323 (33%), Positives = 177/323 (54%), Gaps = 13/323 (4%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ + +E E++VGP+P+DD GKE+ T + + + FYTDS+GR+ ++R +D R +
Sbjct: 693 IYEGVNRVEFEWLVGPVPIDDELGKEIVTIFKSGISSGGVFYTDSNGREMMRREKDKRED 752
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ ++ QP++GNYY V + ++D SK + +L DRS GG+S++DG++E+MLHRR + D
Sbjct: 753 FSPDLSEQPVSGNYYQVTSRVALQDSSKRMVLLNDRSQGGASLEDGRLEMMLHRRHIFAD 812
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGA-RWRRTFGQEIYSPLLLAFTEQ 911
G G AEA+NE + +GLI +GK ++ ++ +GA R +EI+ P F++
Sbjct: 813 GSGAAEAINE----QQFGKGLIARGKLFLYLNAIEDGATASERVAEKEIHLPFWKFFSK- 867
Query: 912 DGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLAS 1091
N S V K + LP +V +LTL+ +LLR + + E ++ S
Sbjct: 868 --SNNIQSDVTK--TLSDFNDLPQSVHLLTLEPYSKDEILLRLENFLDQTE----GNVVS 919
Query: 1092 IDLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQGSAADEKV-----------VR 1238
++++F + ++ ET+L N + M KRLK+ GS V
Sbjct: 920 FSIRQIFDDLGGLEIRETTLDGNLPLSDM--KRLKFHHDGSGPSHSVPEYFTSLHKPLAA 977
Query: 1239 GGPVDPSKLVVELGPMEIRTFIV 1307
D S+ V L PM+IRTFI+
Sbjct: 978 DKTQDASEFSVTLKPMQIRTFII 1000
Score = 36.2 bits (82), Expect(2) = 4e-43
Identities = 18/47 (38%), Positives = 25/47 (53%)
Frame = +2
Query: 248 YIFRPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQG 388
YIFR + + + D TV G + EVHQ +N WI Q+ R +G
Sbjct: 651 YIFRQDADLELVEDA-FDFTVYDGESVKEVHQHVNEWISQVIRIYEG 696
>sptr|Q8MS44|Q8MS44 RE08556p.
Length = 1007
Score = 166 bits (420), Expect = 1e-39
Identities = 107/320 (33%), Positives = 172/320 (53%), Gaps = 10/320 (3%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ K+ +E E+ VGPI ++ G+EV + + +N YTDS+GR+ IKR++D R
Sbjct: 698 IYEGKNLVEIEWQVGPIEREEEFGREVVIIFNSTIASNGVSYTDSNGREMIKRVKDQRET 757
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ + QP A NYYPV I ++D K ++VL DR+ GG+S+ +GQ+ELMLHRRL+ DD
Sbjct: 758 FTPGLDRQPTAANYYPVTSRIALQDSKKRIAVLNDRAQGGASMLNGQLELMLHRRLVRDD 817
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQD 914
G GV EALNE +K + +I +GK Y+ + P E R +EI+ P F++
Sbjct: 818 GYGVGEALNE----EKYGQPMIARGKVYLILSPSDESTAAEREAEKEIHLPFWKFFSKNT 873
Query: 915 GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASI 1094
G S A ++ S P +V +LTL+ D VLLR + + E + S
Sbjct: 874 G-----STTAAAKSVPSFNDFPKSVHLLTLEPFNDDEVLLRVENFLDHTE----GQVVSF 924
Query: 1095 DLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQG---------SAADEKVVRGGP 1247
+++ +F ++ ET+L N + M++ + + G ++A + +
Sbjct: 925 NIRPIFDYLNGVEIRETTLDGNLPLSDMKRFKFHHDSSGQKPDAVEYFTSAHKPLAAEQS 984
Query: 1248 VDPSKLVVELGPMEIRTFIV 1307
+ S+ V L PM+IRTFI+
Sbjct: 985 QEASEFSVTLHPMQIRTFII 1004
>sptr|Q9VLH9|Q9VLH9 CG9468 protein.
Length = 1007
Score = 164 bits (415), Expect = 4e-39
Identities = 106/320 (33%), Positives = 172/320 (53%), Gaps = 10/320 (3%)
Frame = +3
Query: 378 VYKEKDFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSE 557
+Y+ K+ +E E+ VGPI ++ G+EV + + ++ YTDS+GR+ IKR++D R
Sbjct: 698 IYEGKNLVEIEWQVGPIEREEEFGREVVIIFNSTIASDGVSYTDSNGREMIKRVKDQRET 757
Query: 558 WKIEV-HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDD 734
+ + QP A NYYPV I ++D K ++VL DR+ GG+S+ +GQ+ELMLHRRL+ DD
Sbjct: 758 FTPGLDRQPTAANYYPVTSRIALQDSKKRIAVLNDRAQGGASMLNGQLELMLHRRLVRDD 817
Query: 735 GKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLLAFTEQD 914
G GV EALNE +K + +I +GK Y+ + P E R +EI+ P F++
Sbjct: 818 GYGVGEALNE----EKYGQPMIARGKVYLILSPSDESTAAEREAEKEIHLPFWKFFSKNT 873
Query: 915 GGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDLSSLASI 1094
G S A ++ S P +V +LTL+ D VLLR + + E + S
Sbjct: 874 G-----STTAAAKSVPSFNDFPKSVHLLTLEPFNDDEVLLRVENFLDHTE----GQVVSF 924
Query: 1095 DLKRVFPEKKIGKVIETSLSANQERAAMEKKRLKWKAQG---------SAADEKVVRGGP 1247
+++ +F ++ ET+L N + M++ + + G ++A + +
Sbjct: 925 NIRPIFDYLNGVEIRETTLDGNLPLSDMKRFKFHHDSSGQKPDAVEYFTSAHKPLAAEQS 984
Query: 1248 VDPSKLVVELGPMEIRTFIV 1307
+ S+ V L PM+IRTFI+
Sbjct: 985 QEASEFSVTLHPMQIRTFII 1004
>sptr|O97357|O97357 Lysosomal acid alpha-mannosidase precursor.
Length = 977
Score = 134 bits (336), Expect(2) = 2e-33
Identities = 79/259 (30%), Positives = 133/259 (51%), Gaps = 17/259 (6%)
Frame = +3
Query: 393 DFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSEWKIEV 572
D ++ EF IP+ D G+E+ T++ + FYTDS+GR+ +R D+RS++
Sbjct: 679 DLVDVEFTSFGIPIHDNFGRELVARFRTSVESGDVFYTDSNGREMQRRRVDHRSDYPFTQ 738
Query: 573 HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDGKGVAE 752
+P+AGN+YPV ++ D + +V D +GG+S++ G++ ++HRRLL DD KGV E
Sbjct: 739 TEPVAGNFYPVTSVFFINDTQTQFNVFPDAPMGGTSLQSGEVLFVVHRRLLRDDFKGVEE 798
Query: 753 ALNETVCLDKQCE---------------GLIIQGKYYVKIDPQGEGA--RWRRTFGQEIY 881
LNET + + L ++G + G A R R + Y
Sbjct: 799 PLNETAFVTSYADCVMANTSNCGHHYGPPLRVRGTLSFSVSRSGPTAMRRVREQQDENYY 858
Query: 882 SPLLLAFTEQDGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAG 1061
+PL++ + A S +A+++ +SLP ++ ++TLQ ++ +LLR H Y
Sbjct: 859 TPLVMFSSSSTD---AASAIARYNT-SFGFSLPPSLQVVTLQLIDQQKLLLRLGHRYAVS 914
Query: 1062 EDKDLSSLASIDLKRVFPE 1118
ED + S +DL +F E
Sbjct: 915 EDPERSLPVEVDLMTIFKE 933
Score = 32.3 bits (72), Expect(2) = 2e-33
Identities = 21/79 (26%), Positives = 35/79 (44%), Gaps = 2/79 (2%)
Frame = +2
Query: 83 NLQVGQGNLKLQYNEAGKLSLYSDSKTMVQANFEQKYKYYIGQDGNGSDPQASGAYIFRP 262
+L++ L L++ G + + + +Q + YYI +G+ GAYI RP
Sbjct: 577 DLEISNEALVLKFGRNGLIESVTVRSSGQTVAVKQDWCYYISNNGDTVSSSPGGAYIMRP 636
Query: 263 --NGTVPISTDGQVSLTVL 313
N T TD V L ++
Sbjct: 637 VSNATCNPITDSPVELRLV 655
>sptr|Q20829|Q20829 Hypothetical protein.
Length = 955
Score = 121 bits (304), Expect = 3e-26
Identities = 86/269 (31%), Positives = 138/269 (51%), Gaps = 2/269 (0%)
Frame = +3
Query: 363 IRLPGVYKEKDFLETEFIVGPIPVDDGNG--KEVATEIVTNMVTNKTFYTDSSGRDFIKR 536
IRLP + K+++E E+IVGPIP + N KE T T + + +TDS+GR ++R
Sbjct: 666 IRLP---RGKNYIEFEWIVGPIPKETKNPITKEFVTRYTTEVHSKNQSFTDSNGRQAMER 722
Query: 537 IRDYRSEWKIEVHQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHR 716
D + ++ +PIAGNYYP+ Y++D + + SV+ DR+ G DG +E+MLHR
Sbjct: 723 FFDGATSFEYSDTEPIAGNYYPITSFGYIKDENAQFSVITDRA-QGMVASDGVVEIMLHR 781
Query: 717 RLLHDDGKGVAEALNETVCLDKQCEGLIIQGKYYVKIDPQGEGARWRRTFGQEIYSPLLL 896
R +DD GV EAL+E K GL+ GK+ V R + + +L
Sbjct: 782 RCFYDDHFGVEEALDEP---GKDGAGLVAMGKHIVLFTDVKASTSQLRPLAIDNFHQPVL 838
Query: 897 AFTEQDGGNWANSHVAKFSAMDSTYSLPDNVAMLTLQELEDGSVLLRFAHLYEAGEDKDL 1076
AF ++ + + + ++S + + LP + +LTL++ S L+RF ++Y E
Sbjct: 839 AF-NKNPEKFEFNRLLEYSGL--KFELPIGIHLLTLEKWHKKSSLIRFENIYHGEEG--- 892
Query: 1077 SSLASIDLKRVFPEKKIGKVIETSLSANQ 1163
+S+ D +F I E L AN+
Sbjct: 893 NSIKEFDPTHIFTNFDIVSFKELLLGANR 921
Score = 60.1 bits (144), Expect = 1e-07
Identities = 33/105 (31%), Positives = 60/105 (57%)
Frame = +2
Query: 77 SSNLQVGQGNLKLQYNEAGKLSLYSDSKTMVQANFEQKYKYYIGQDGNGSDPQASGAYIF 256
++N+++ L ++ G LS ++ T + + Q++ YY G D D Q SGAYIF
Sbjct: 570 NTNVEIQNEYLIAGFDSQGYLSYITEKATKKKRSIRQEFFYYEGIDSK--DDQPSGAYIF 627
Query: 257 RPNGTVPISTDGQVSLTVLRGPILDEVHQQINSWIYQITRRLQGE 391
RP PI+ +++L V+ G I++EV Q+++ ++ Q R +G+
Sbjct: 628 RPKTQQPIALTSKLALEVVPGAIINEVRQKVSPFVSQQIRLPRGK 672
>sptr|O96635|O96635 Lysosomal alpha mannosidase (Fragment).
Length = 393
Score = 100 bits (249), Expect = 7e-20
Identities = 78/289 (26%), Positives = 134/289 (46%), Gaps = 24/289 (8%)
Frame = +3
Query: 393 DFLETEFIVGPIPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSEWKIEV 572
D ++ EF IP+ D G+E+ T++ + FYTDS+GR+ +R D+RS++
Sbjct: 95 DLVDVEFTSFGIPIHDNFGRELVARFRTSVESGDVFYTDSNGREMQRRRVDHRSDYPFTQ 154
Query: 573 HQPIAGNYYPVNLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDGKGVAE 752
+P+AGN+YPV +V D + +V D +GG+S++ G++ L +GV
Sbjct: 155 TEPVAGNFYPVTSVFFVHDTQTQFNVFPDAPMGGTSLQSGEVLLWFTDVFYVTTLEGVEN 214
Query: 753 ALNETVCLDKQCEGLIIQGKYYVKIDP------------------QGEGARWRRTFGQEI 878
L + L + + Q + V I P Q R R +
Sbjct: 215 LLTKPP-LSQATRIVGCQRQVIVAIIPPLPCVFAARCHFQCRGVAQPRMRRVRDPQDENY 273
Query: 879 YSPLLLAFTEQDGGNWANSHVAKFSAMDST---YSLPDNVAMLTLQELEDGSVLLRFAHL 1049
Y+PL++ + +++ A A ++T +SLP ++ ++TLQ ++ +LLR H
Sbjct: 274 YTPLVMFSS-------SSTDAASAIARNNTSFGFSLPPSLQVVTLQLIDQQKLLLRLGHR 326
Query: 1050 YEAGEDKDLSSLASIDLKRVFPE---KKIGKVIETSLSANQERAAMEKK 1187
Y ED + S +DL +F E I + E SL+A + A + K
Sbjct: 327 YAVSEDPERSLPVEVDLMTIFREFHWITILSIDEVSLTAVGDSATPDPK 375
>sptr|Q7PMM6|Q7PMM6 ENSANGP00000014255.
Length = 1128
Score = 74.7 bits (182), Expect(2) = 2e-15
Identities = 37/115 (32%), Positives = 64/115 (55%)
Frame = +3
Query: 426 IPVDDGNGKEVATEIVTNMVTNKTFYTDSSGRDFIKRIRDYRSEWKIEVHQPIAGNYYPV 605
+ + + E+ + TN+ + TFYTD +G IKR R + PI NYYPV
Sbjct: 895 VDIGQRDNTEIVMRLQTNINSGATFYTDLNGMQIIKRKRFQKL--------PIQANYYPV 946
Query: 606 NLGIYVEDGSKELSVLVDRSIGGSSIKDGQIELMLHRRLLHDDGKGVAEALNETV 770
+++ED + L++L + +GGSS+ G++E+M RRL DD +G+ + + + +
Sbjct: 947 PSTMFIEDDNYRLTLLGGQPLGGSSLSSGEMEIMQDRRLTRDDDRGLGQGVQDNL 1001
Score = 31.2 bits (69), Expect(2) = 2e-15
Identities = 22/81 (27%), Positives = 36/81 (44%), Gaps = 8/81 (9%)
Frame = +2
Query: 107 LKLQYNEAGKLSLYSDSKTMV--------QANFEQKYKYYIGQDGNGSDPQASGAYIFRP 262
+ L++ E G S SK ++ QA ++Y + G SGAY+F P
Sbjct: 789 MSLRFEEGGSTSAAFTSKGLLKSLTIENNQATVPVHLEFY--RYGMQLSSGKSGAYLFHP 846
Query: 263 NGTVPISTDGQVSLTVLRGPI 325
G + T Q + V++GP+
Sbjct: 847 AGNATLMTYDQPIVLVMKGPL 867
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,376,294,041
Number of Sequences: 1381838
Number of extensions: 31392275
Number of successful extensions: 93569
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 82827
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 92981
length of database: 439,479,560
effective HSP length: 128
effective length of database: 262,604,296
effective search space used: 108455574248
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

