BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QAO3g02.yg.2.1
(344 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
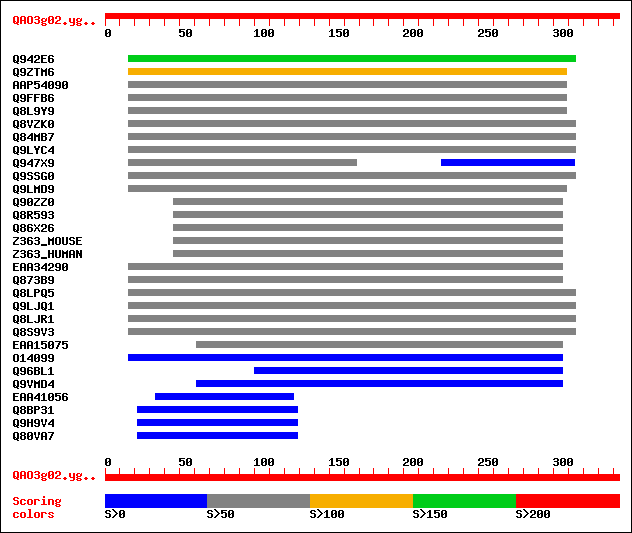
sptr|Q942E6|Q942E6 Putative PGPD14 protein (Pollen germination r... 155 1e-37
sptr|Q9ZTM6|Q9ZTM6 PGPD14. 102 1e-21
sptrnew|AAP54090|AAP54090 Putative PGPD14 protein (Pollen germin... 99 1e-20
sptr|Q9FFB6|Q9FFB6 PGPD14 protein (AT5g22920/MRN17_15). 98 3e-20
sptr|Q8L9Y9|Q8L9Y9 PGPD14 protein. 97 4e-20
sptr|Q8VZK0|Q8VZK0 Hypothetical protein. 87 4e-17
sptr|Q84MB7|Q84MB7 At3g62970. 77 7e-14
sptr|Q9LYC4|Q9LYC4 Hypothetical 30.9 kDa protein. 72 2e-12
sptr|Q947X9|Q947X9 Hypothetical 39.6 kDa protein. 58 3e-11
sptr|Q9SSG0|Q9SSG0 F25A4.27 protein. 64 4e-10
sptr|Q9LMD9|Q9LMD9 F14D16.3. 63 8e-10
sptr|Q90ZZ0|Q90ZZ0 Zinc finger protein. 58 2e-08
sptr|Q8R593|Q8R593 RIKEN cDNA 6720407C15 gene. 57 4e-08
sptr|Q86X26|Q86X26 Similar to zinc finger protein 363. 57 4e-08
sw|Q9CR50|Z363_MOUSE Zinc finger protein 363 (CH-rich interactin... 57 4e-08
sw|Q96PM5|Z363_HUMAN Zinc finger protein 363 (CH-rich interactin... 57 4e-08
sptrnew|EAA34290|EAA34290 Hypothetical protein. 57 7e-08
sptr|Q873B9|Q873B9 Hypothetical protein B24N11.050. 57 7e-08
sptr|Q8LPQ5|Q8LPQ5 AT3g18290/MIE15_8. 56 9e-08
sptr|Q9LJQ1|Q9LJQ1 Gb|AAD55299.1. 56 9e-08
sptr|Q8LJR1|Q8LJR1 Putative zinc finger protein. 54 4e-07
sptr|Q8S9V3|Q8S9V3 Putative zinc finger protein. 53 1e-06
sptrnew|EAA15075|EAA15075 ENSANGP00000012255 (Fragment). 50 7e-06
sptr|O14099|O14099 Zinc finger protein. 48 3e-05
sptr|Q96BL1|Q96BL1 Hypothetical protein (Fragment). 48 3e-05
sptr|Q9VMD4|Q9VMD4 CG16947 protein. 46 1e-04
sptrnew|EAA41056|EAA41056 GLP_447_25082_24141. 33 0.64
sptr|Q8BP31|Q8BP31 Unknown EST. 32 2.4
sptr|Q9H9V4|Q9H9V4 Hypothetical protein FLJ12526. 32 2.4
sptr|Q80VA7|Q80VA7 Hypothetical protein (Fragment). 32 2.4
>sptr|Q942E6|Q942E6 Putative PGPD14 protein (Pollen germination
related protein).
Length = 299
Score = 155 bits (391), Expect = 1e-37
Identities = 72/100 (72%), Positives = 78/100 (78%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSD 195
YLF+STNDVSVLP HTIHVKCL+EMEEHCQFACPLCS M ERL +SD
Sbjct: 200 YLFESTNDVSVLPCGHTIHVKCLREMEEHCQFACPLCSKSVCDMSKAWERLDEELATISD 259
Query: 196 SCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
+CD+KMVRILCNDCGA S+VQFHLIAHKCQ SYNTR I
Sbjct: 260 TCDNKMVRILCNDCGATSEVQFHLIAHKCQKCKSYNTRQI 299
>sptr|Q9ZTM6|Q9ZTM6 PGPD14.
Length = 285
Score = 102 bits (254), Expect = 1e-21
Identities = 45/100 (45%), Positives = 66/100 (66%), Gaps = 2/100 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERL--XXXXXXL 189
Y+FD+T +++VLP HT+H++C+ +ME+H Q++CP+CS M E+L +
Sbjct: 168 YVFDTTKNITVLPCGHTMHLECVMQMEQHNQYSCPVCSRSYCDMSRVWEKLDQEVASTPM 227
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTR 309
+ +KMV ILCNDCG S+V FH++A KC N SYNTR
Sbjct: 228 PEMYQNKMVWILCNDCGETSEVNFHIVARKCPNCKSYNTR 267
>sptrnew|AAP54090|AAP54090 Putative PGPD14 protein (Pollen
germination related protein).
Length = 266
Score = 99.4 bits (246), Expect = 1e-20
Identities = 48/100 (48%), Positives = 60/100 (60%), Gaps = 2/100 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERL--XXXXXXL 189
YLFDST D+SVL HTIH++CL M H FACP+CS M ++L +
Sbjct: 152 YLFDSTKDISVLHCGHTIHLECLNVMRAHHHFACPVCSRSACDMSDAWKKLDEEVAATPM 211
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTR 309
+ KM+ ILCNDCGA S+V FH++A KC SYNTR
Sbjct: 212 PEFYQKKMIWILCNDCGATSNVNFHVLAQKCPGCSSYNTR 251
>sptr|Q9FFB6|Q9FFB6 PGPD14 protein (AT5g22920/MRN17_15).
Length = 291
Score = 97.8 bits (242), Expect = 3e-20
Identities = 46/100 (46%), Positives = 65/100 (65%), Gaps = 2/100 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--L 189
YLFDST D++VL HT+H++C K+M H ++ CP+CS M ++L +
Sbjct: 169 YLFDSTRDITVLRCGHTMHLECTKDMGLHNRYTCPVCSKSICDMSNLWKKLDEEVAAYPM 228
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTR 309
++KMV ILCNDCG+ ++V+FHLIAHKC + SYNTR
Sbjct: 229 PKMYENKMVWILCNDCGSNTNVRFHLIAHKCSSCGSYNTR 268
>sptr|Q8L9Y9|Q8L9Y9 PGPD14 protein.
Length = 274
Score = 97.4 bits (241), Expect = 4e-20
Identities = 46/100 (46%), Positives = 65/100 (65%), Gaps = 2/100 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--L 189
YLFDST D++VL HT+H++C K+M H ++ CP+CS M ++L +
Sbjct: 152 YLFDSTRDITVLRCGHTMHLECTKDMGLHNRYTCPVCSKSICDMSNLWKKLDEEVAAYPM 211
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTR 309
++KMV ILCNDCG+ ++V+FHLIAHKC + SYNTR
Sbjct: 212 PKLYENKMVWILCNDCGSNTNVRFHLIAHKCSSCGSYNTR 251
>sptr|Q8VZK0|Q8VZK0 Hypothetical protein.
Length = 267
Score = 87.4 bits (215), Expect = 4e-17
Identities = 43/102 (42%), Positives = 57/102 (55%), Gaps = 2/102 (1%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSD 195
YLFDS D +V+ HT+HV+C EM + +F CP+CS M +RL +
Sbjct: 159 YLFDSLKDTNVMKCGHTMHVECYNEMIKRDKFCCPICSRSVIDMSKTWQRLDEEIEATAM 218
Query: 196 SCDH--KMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
D+ K V ILCNDC ++V FH+I KC + SYNTR I
Sbjct: 219 PSDYRDKKVWILCNDCNDTTEVHFHIIGQKCGHCRSYNTRAI 260
>sptr|Q84MB7|Q84MB7 At3g62970.
Length = 276
Score = 76.6 bits (187), Expect = 7e-14
Identities = 37/101 (36%), Positives = 50/101 (49%), Gaps = 1/101 (0%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSD 195
YLFDS V+ HT+H+ C ++M Q+ CP+C+ M L
Sbjct: 165 YLFDSVKAAHVMKCGHTMHMDCFEQMINENQYRCPICAKSMVDMSPSWHLLDFEISATEM 224
Query: 196 SCDHKM-VRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
++K V ILCNDC S FH++ HKC + SYNTR I
Sbjct: 225 PVEYKFEVSILCNDCNKGSKAMFHILGHKCSDCGSYNTRRI 265
>sptr|Q9LYC4|Q9LYC4 Hypothetical 30.9 kDa protein.
Length = 274
Score = 71.6 bits (174), Expect = 2e-12
Identities = 36/100 (36%), Positives = 47/100 (47%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSD 195
YLFDS V+ HT+H+ C ++M Q+ CP+C+ M L
Sbjct: 176 YLFDSVKAAHVMKCGHTMHMDCFEQMINENQYRCPICAKSMVDMSPSWHLLDFE------ 229
Query: 196 SCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
V ILCNDC S FH++ HKC + SYNTR I
Sbjct: 230 ------VSILCNDCNKGSKAMFHILGHKCSDCGSYNTRRI 263
>sptr|Q947X9|Q947X9 Hypothetical 39.6 kDa protein.
Length = 349
Score = 58.2 bits (139), Expect(2) = 3e-11
Identities = 25/51 (49%), Positives = 32/51 (62%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERL 168
YLFDST D+S L HTIH++CL EM H QF+CP+C M ++L
Sbjct: 218 YLFDSTKDISALHCGHTIHLECLYEMRSHQQFSCPVCLRSACDMSHAWQKL 268
Score = 29.6 bits (65), Expect(2) = 3e-11
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = +3
Query: 225 MQRLRGNIRCAVPFDCAQVPEXXXIQHPXD 314
MQRL +I A+P QVP +QHP D
Sbjct: 303 MQRLWDDIERAIPHIGTQVPRMQLLQHPAD 332
>sptr|Q9SSG0|Q9SSG0 F25A4.27 protein.
Length = 255
Score = 63.9 bits (154), Expect = 4e-10
Identities = 34/103 (33%), Positives = 49/103 (47%), Gaps = 3/103 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
Y+F S++ V LP H +H C +E C + CP+CS M + L
Sbjct: 155 YIFTSSSPVKALPCGHLMHSTCFQEYT--CSHYTCPVCSKSLGDMQVYFKMLDALLAEEK 212
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
+ D +K ILCNDCG + +H + HKC SYN+R +
Sbjct: 213 MPDEYSNKTQVILCNDCGRKGNAPYHWLYHKCTTCGSYNSRLL 255
>sptr|Q9LMD9|Q9LMD9 F14D16.3.
Length = 1260
Score = 63.2 bits (152), Expect = 8e-10
Identities = 34/101 (33%), Positives = 47/101 (46%), Gaps = 3/101 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
Y+F S + V LP H +H C +E C + CP+CS M L
Sbjct: 1160 YIFTSNSPVKALPCGHVMHSTCFQEYT--CSHYTCPICSKSLGDMQVYFRMLDALLAEQK 1217
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTR 309
+ D ++ ILCNDCG + +H + HKC + SYNTR
Sbjct: 1218 MPDEYLNQTQVILCNDCGRKGNAPYHWLYHKCSSCASYNTR 1258
>sptr|Q90ZZ0|Q90ZZ0 Zinc finger protein.
Length = 264
Score = 58.2 bits (139), Expect = 2e-08
Identities = 29/89 (32%), Positives = 44/89 (49%), Gaps = 2/89 (2%)
Frame = +1
Query: 46 VLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSDSCDHK--MVR 219
VLP H +H C +M + + CPLC M +++ +++ V+
Sbjct: 149 VLPCGHLLHGTCFDDMLKTGAYRCPLCMHSAFNMKEYWKQMDEEISQTPMPTEYQDSTVK 208
Query: 220 ILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
I+CNDC A S V FH++ KC + SYNT
Sbjct: 209 IICNDCQARSTVSFHVLGMKCSSCGSYNT 237
>sptr|Q8R593|Q8R593 RIKEN cDNA 6720407C15 gene.
Length = 261
Score = 57.4 bits (137), Expect = 4e-08
Identities = 32/89 (35%), Positives = 43/89 (48%), Gaps = 2/89 (2%)
Frame = +1
Query: 46 VLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSDSCDHKMVR-- 219
VLP H +H C +EM + + CPLC M +L +++ V
Sbjct: 161 VLPCGHLLHRTCYEEMLKE-GYRCPLCMHSALDMTRYWRQLDTEVAQTPMPSEYQNVTVD 219
Query: 220 ILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
ILCNDC S VQFH++ KC+ SYNT
Sbjct: 220 ILCNDCNGRSTVQFHILGMKCKLCDSYNT 248
>sptr|Q86X26|Q86X26 Similar to zinc finger protein 363.
Length = 261
Score = 57.4 bits (137), Expect = 4e-08
Identities = 32/89 (35%), Positives = 42/89 (47%), Gaps = 2/89 (2%)
Frame = +1
Query: 46 VLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--LSDSCDHKMVR 219
VLP H +H C +EM + + CPLC M +L + + V
Sbjct: 161 VLPCGHLLHRTCYEEMLKE-GYRCPLCMHSALDMTRYWRQLDDEVAQTPMPSEYQNMTVD 219
Query: 220 ILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
ILCNDC S VQFH++ KC+ SYNT
Sbjct: 220 ILCNDCNGRSTVQFHILGMKCKICESYNT 248
>sw|Q9CR50|Z363_MOUSE Zinc finger protein 363 (CH-rich interacting
match with PLAG1) (Androgen receptor
N-terminal-interacting protein).
Length = 261
Score = 57.4 bits (137), Expect = 4e-08
Identities = 32/89 (35%), Positives = 43/89 (48%), Gaps = 2/89 (2%)
Frame = +1
Query: 46 VLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSDSCDHKMVR-- 219
VLP H +H C +EM + + CPLC M +L +++ V
Sbjct: 161 VLPCGHLLHRTCYEEMLKE-GYRCPLCMHSALDMTRYWRQLDTEVAQTPMPSEYQNVTVD 219
Query: 220 ILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
ILCNDC S VQFH++ KC+ SYNT
Sbjct: 220 ILCNDCNGRSTVQFHILGMKCKLCDSYNT 248
>sw|Q96PM5|Z363_HUMAN Zinc finger protein 363 (CH-rich interacting
match with PLAG1) (Androgen receptor
N-terminal-interacting protein).
Length = 261
Score = 57.4 bits (137), Expect = 4e-08
Identities = 32/89 (35%), Positives = 42/89 (47%), Gaps = 2/89 (2%)
Frame = +1
Query: 46 VLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--LSDSCDHKMVR 219
VLP H +H C +EM + + CPLC M +L + + V
Sbjct: 161 VLPCGHLLHRTCYEEMLKE-GYRCPLCMHSALDMTRYWRQLDDEVAQTPMPSEYQNMTVD 219
Query: 220 ILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
ILCNDC S VQFH++ KC+ SYNT
Sbjct: 220 ILCNDCNGRSTVQFHILGMKCKICESYNT 248
>sptrnew|EAA34290|EAA34290 Hypothetical protein.
Length = 888
Score = 56.6 bits (135), Expect = 7e-08
Identities = 31/99 (31%), Positives = 42/99 (42%), Gaps = 2/99 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--L 189
YLF+S V+++ HTIH C E + CP+C+ M L +
Sbjct: 490 YLFNSPRPVAIMKCGHTIHKHCFSAHRER-SYRCPVCNKSCVNMEIQFRNLDIAIATQPM 548
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
D I CNDC A S ++H + KC SYNT
Sbjct: 549 PDEYQDARAVISCNDCSAKSQTRYHWLGLKCSVCHSYNT 587
>sptr|Q873B9|Q873B9 Hypothetical protein B24N11.050.
Length = 888
Score = 56.6 bits (135), Expect = 7e-08
Identities = 31/99 (31%), Positives = 42/99 (42%), Gaps = 2/99 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--L 189
YLF+S V+++ HTIH C E + CP+C+ M L +
Sbjct: 490 YLFNSPRPVAIMKCGHTIHKHCFSAHRER-SYRCPVCNKSCVNMEIQFRNLDIAIATQPM 548
Query: 190 SDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
D I CNDC A S ++H + KC SYNT
Sbjct: 549 PDEYQDARAVISCNDCSAKSQTRYHWLGLKCSVCHSYNT 587
>sptr|Q8LPQ5|Q8LPQ5 AT3g18290/MIE15_8.
Length = 1254
Score = 56.2 bits (134), Expect = 9e-08
Identities = 33/103 (32%), Positives = 45/103 (43%), Gaps = 3/103 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
+LF S+ V LP H +H C + C + CP+C M L
Sbjct: 1141 FLFTSSEAVRALPCGHYMHSACFQAYT--CSHYTCPICGKSLGDMAVYFGMLDALLAAEE 1198
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
L + ++ ILCNDC +FH + HKC + SYNTR I
Sbjct: 1199 LPEEYKNRCQDILCNDCERKGTTRFHWLYHKCGSCGSYNTRVI 1241
>sptr|Q9LJQ1|Q9LJQ1 Gb|AAD55299.1.
Length = 1232
Score = 56.2 bits (134), Expect = 9e-08
Identities = 33/103 (32%), Positives = 45/103 (43%), Gaps = 3/103 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
+LF S+ V LP H +H C + C + CP+C M L
Sbjct: 1119 FLFTSSEAVRALPCGHYMHSACFQAYT--CSHYTCPICGKSLGDMAVYFGMLDALLAAEE 1176
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
L + ++ ILCNDC +FH + HKC + SYNTR I
Sbjct: 1177 LPEEYKNRCQDILCNDCERKGTTRFHWLYHKCGSCGSYNTRVI 1219
>sptr|Q8LJR1|Q8LJR1 Putative zinc finger protein.
Length = 400
Score = 54.3 bits (129), Expect = 4e-07
Identities = 33/103 (32%), Positives = 45/103 (43%), Gaps = 3/103 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
+LF S+ V LP H++H C + C + CP+C M L
Sbjct: 289 FLFTSSAAVRALPCGHSMHSACFQAYT--CSHYTCPICCKSLGDMAVYFGMLDALLAAEE 346
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
L + + ILCNDC +FH + HKC + SYNTR I
Sbjct: 347 LPEEYRDRCQDILCNDCERKGRCRFHWLYHKCGSCGSYNTRVI 389
>sptr|Q8S9V3|Q8S9V3 Putative zinc finger protein.
Length = 1063
Score = 52.8 bits (125), Expect = 1e-06
Identities = 33/103 (32%), Positives = 44/103 (42%), Gaps = 3/103 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQ-FACPLCSXXXXXMXXXXERLXXXXXX-- 186
+LF S+ V LP H +H C + C + CP+C M L
Sbjct: 952 FLFTSSAAVRALPCGHFMHSACFQAYT--CSHYTCPICCKSLGDMAVYFGMLDALLAAEE 1009
Query: 187 LSDSCDHKMVRILCNDCGAISDVQFHLIAHKCQNXXSYNTRXI 315
L + + ILCNDC +FH + HKC + SYNTR I
Sbjct: 1010 LPEEYRDRCQDILCNDCERKGRSRFHWLYHKCGSCGSYNTRVI 1052
>sptrnew|EAA15075|EAA15075 ENSANGP00000012255 (Fragment).
Length = 578
Score = 50.1 bits (118), Expect = 7e-06
Identities = 26/84 (30%), Positives = 37/84 (44%), Gaps = 2/84 (2%)
Frame = +1
Query: 61 HTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXX--LSDSCDHKMVRILCND 234
H +H C +E+ +ACP C M E L + + V ILC D
Sbjct: 242 HLLHRTCFEELLSSGHYACPTCQTSMMDMNQLWEYLDAEVAATPMPKEYANYFVDILCKD 301
Query: 235 CGAISDVQFHLIAHKCQNXXSYNT 306
C S V+FH++ KC + +YNT
Sbjct: 302 CHKESTVKFHVVGLKCTHCGAYNT 325
>sptr|O14099|O14099 Zinc finger protein.
Length = 425
Score = 47.8 bits (112), Expect = 3e-05
Identities = 27/99 (27%), Positives = 41/99 (41%), Gaps = 2/99 (2%)
Frame = +1
Query: 16 YLFDSTNDVSVLPXXHTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSD 195
Y+F+S V L H +H +C +E + CP C + L
Sbjct: 277 YMFNSRERVIFLSCSHPLHQRCHEEYIR-TNYRCPTCYKTIINVNSLFRILDMEIERQPM 335
Query: 196 SCDHK--MVRILCNDCGAISDVQFHLIAHKCQNXXSYNT 306
+ + I CNDC + D ++H + HKC + SYNT
Sbjct: 336 PYPYNTWISTIRCNDCNSRCDTKYHFLGHKCNSCHSYNT 374
>sptr|Q96BL1|Q96BL1 Hypothetical protein (Fragment).
Length = 117
Score = 47.8 bits (112), Expect = 3e-05
Identities = 25/71 (35%), Positives = 32/71 (45%), Gaps = 2/71 (2%)
Frame = +1
Query: 100 HCQFACPLCSXXXXXMXXXXERLXXXXXX--LSDSCDHKMVRILCNDCGAISDVQFHLIA 273
H + CPLC M +L + + V ILCNDC S VQFH++
Sbjct: 34 HRGYRCPLCMHSALDMTRYWRQLDDEVAQTPMPSEYQNMTVDILCNDCNGRSTVQFHILG 93
Query: 274 HKCQNXXSYNT 306
KC+ SYNT
Sbjct: 94 MKCKICESYNT 104
>sptr|Q9VMD4|Q9VMD4 CG16947 protein.
Length = 433
Score = 45.8 bits (107), Expect = 1e-04
Identities = 23/84 (27%), Positives = 34/84 (40%), Gaps = 2/84 (2%)
Frame = +1
Query: 61 HTIHVKCLKEMEEHCQFACPLCSXXXXXMXXXXERLXXXXXXLSDSC--DHKMVRILCND 234
H +H C ++ + CP C M L + +++ V I CND
Sbjct: 332 HLLHKMCFDQLLASGHYTCPTCQTSLIDMTALWVYLDDQAERMPVPLKYENQRVHIFCND 391
Query: 235 CGAISDVQFHLIAHKCQNXXSYNT 306
C S +FH I KC + +YNT
Sbjct: 392 CHKTSKTKFHFIGLKCVHCGAYNT 415
>sptrnew|EAA41056|EAA41056 GLP_447_25082_24141.
Length = 313
Score = 33.5 bits (75), Expect = 0.64
Identities = 15/31 (48%), Positives = 17/31 (54%)
Frame = +1
Query: 34 NDVSVLPXXHTIHVKCLKEMEEHCQFACPLC 126
NDV VLP HT H CL + +CPLC
Sbjct: 102 NDVVVLPCTHTFHRICLIRWLHYDNTSCPLC 132
>sptr|Q8BP31|Q8BP31 Unknown EST.
Length = 155
Score = 31.6 bits (70), Expect = 2.4
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +1
Query: 22 FDSTNDVSVLPXXHTIHVKCL-KEMEEHCQFACPLCS 129
F +++ VLP H H KCL K +E C CP+C+
Sbjct: 100 FKGKDELGVLPCQHAFHRKCLVKWLEVRC--VCPMCN 134
>sptr|Q9H9V4|Q9H9V4 Hypothetical protein FLJ12526.
Length = 155
Score = 31.6 bits (70), Expect = 2.4
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +1
Query: 22 FDSTNDVSVLPXXHTIHVKCL-KEMEEHCQFACPLCS 129
F +++ VLP H H KCL K +E C CP+C+
Sbjct: 100 FKGKDELGVLPCQHAFHRKCLVKWLEVRC--VCPMCN 134
>sptr|Q80VA7|Q80VA7 Hypothetical protein (Fragment).
Length = 151
Score = 31.6 bits (70), Expect = 2.4
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +1
Query: 22 FDSTNDVSVLPXXHTIHVKCL-KEMEEHCQFACPLCS 129
F +++ VLP H H KCL K +E C CP+C+
Sbjct: 96 FKGKDELGVLPCQHAFHRKCLVKWLEVRC--VCPMCN 130
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 200,691,973
Number of Sequences: 1272877
Number of extensions: 2703884
Number of successful extensions: 12486
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 11982
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 12473
length of database: 406,886,216
effective HSP length: 90
effective length of database: 292,327,286
effective search space used: 7015854864
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

