BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
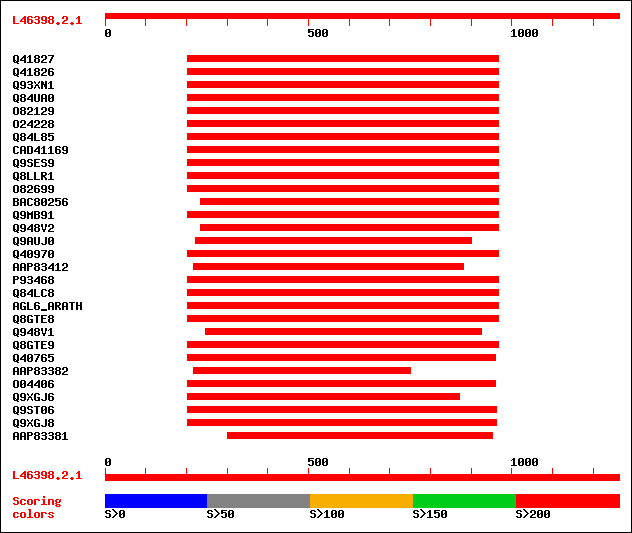
Query= L46398.2.1
(1263 letters)
Database: /db/trembl-ebi/tmp/swall
1,242,019 sequences; 397,940,306 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q41827|Q41827 MADS box protein. 523 e-147
sptr|Q41826|Q41826 MADS box protein. 459 e-128
sptr|Q93XN1|Q93XN1 Mads1. 399 e-110
sptr|Q84UA0|Q84UA0 MADS4. 395 e-109
sptr|O82129|O82129 MADS box transcription factor. 393 e-108
sptr|O24228|O24228 MADS box protein. 392 e-108
sptr|Q84L85|Q84L85 MADS-box transcription factor SEP1. 330 2e-89
sptrnew|CAD41169|CAD41169 OSJNBa0064M23.14 protein. 324 1e-87
sptr|Q9SES9|Q9SES9 MADS box transcription factor MADS17. 324 1e-87
sptr|Q8LLR1|Q8LLR1 MADS-box protein 3. 311 1e-83
sptr|O82699|O82699 MADS-box protein. 304 2e-81
sptrnew|BAC80256|BAC80256 MADS-box transcription factor (Fragment). 300 3e-80
sptr|Q9MB91|Q9MB91 PMADS4 protein. 296 4e-79
sptr|Q948V2|Q948V2 Putative MADS-domain transcription factor MpM... 295 9e-79
sptr|Q9AUJ0|Q9AUJ0 MADS box protein nmads3 (Fragment). 289 5e-77
sptr|Q40970|Q40970 Putative MADS-box family transcription factor. 283 3e-75
sptrnew|AAP83412|AAP83412 AGL6-like MADS-box (Fragment). 282 6e-75
sptr|P93468|P93468 MADS-box family transcription factor. 281 1e-74
sptr|Q84LC8|Q84LC8 MADS-box transcription factor CDM104. 280 2e-74
sw|P29386|AGL6_ARATH Agamous-like MADS box protein AGL6. 278 1e-73
sptr|Q8GTE8|Q8GTE8 MADS-box protein AGL6-a. 275 1e-72
sptr|Q948V1|Q948V1 Putative MADS-domain transcription factor MpM... 273 5e-72
sptr|Q8GTE9|Q8GTE9 MADS-box protein AGL6-a. 268 2e-70
sptr|Q40765|Q40765 Dal1 protein. 267 2e-70
sptrnew|AAP83382|AAP83382 AGL6-like MADS-box (Fragment). 265 8e-70
sptr|O04406|O04406 MADS-box protein. 262 7e-69
sptr|Q9XGJ6|Q9XGJ6 Putative MADS domain transcription factor GGM11. 262 7e-69
sptr|Q9ST06|Q9ST06 GpMADS3 protein. 251 2e-65
sptr|Q9XGJ8|Q9XGJ8 Putative MADS domain transcription factor GGM9. 248 1e-64
sptrnew|AAP83381|AAP83381 AGL6-like MADS-box (Fragment). 243 3e-63
>sptr|Q41827|Q41827 MADS box protein.
Length = 255
Score = 523 bits (1347), Expect = e-147
Identities = 255/255 (100%), Positives = 255/255 (100%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 120
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT
Sbjct: 121 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 180
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPR 921
TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPR
Sbjct: 181 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPR 240
Query: 922 SAATGENNFMLGWVL 966
SAATGENNFMLGWVL
Sbjct: 241 SAATGENNFMLGWVL 255
>sptr|Q41826|Q41826 MADS box protein.
Length = 255
Score = 459 bits (1182), Expect = e-128
Identities = 234/257 (91%), Positives = 239/257 (92%), Gaps = 2/257 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+TKTLERYQHCCYNAQDSN ALSE+QSWYQEMSKLRAKFEALQRTQRHLLGE+LGPLS
Sbjct: 61 GITKTLERYQHCCYNAQDSNG-ALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQQLEKQLECALSQARQRKTQ+MMEQVEELRR ERHLGEMNRQLKHKLEAEGCSNY
Sbjct: 120 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 179
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHP-PAHSVAMDCEPTLQIGY-PHHQFPPPEAVNNI 915
TLQHAA WPAPG T+VEHDGATY VHP A SVAMDCEPTLQIGY PHHQF P EA NNI
Sbjct: 180 TLQHAA-WPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNI 238
Query: 916 PRSAATGENNFMLGWVL 966
PRS GENNFMLGWVL
Sbjct: 239 PRSPPGGENNFMLGWVL 255
>sptr|Q93XN1|Q93XN1 Mads1.
Length = 259
Score = 399 bits (1024), Expect = e-110
Identities = 211/263 (80%), Positives = 223/263 (84%), Gaps = 8/263 (3%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G TKTLERYQHCCYNAQDSN SALSE+QSWYQEMSKL+AKFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GTTKTLERYQHCCYNAQDSN-SALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCS--N 735
VKELQQLEKQLEC+LSQARQRKTQ+M+EQVEELRR ER LGE+NRQLKHKL+AEG S N
Sbjct: 120 VKELQQLEKQLECSLSQARQRKTQLMVEQVEELRRKERQLGEINRQLKHKLDAEGSSSNN 179
Query: 736 YTTLQHAACWPAPGGTIVEHDGATY-----QVHPPAHSVAMDCEPTLQIGYPHHQFPPPE 900
Y +Q W A GT+V+ A Y Q P HS AMDCEPTLQIGYPH P +
Sbjct: 180 YRAMQQLT-WAA--GTVVDEGAAAYHMQHQQQQQPNHSAAMDCEPTLQIGYPHQFAAPEQ 236
Query: 901 AVNNIPRSAAT-GENNFMLGWVL 966
A NNIPRS+ GENNFMLGWVL
Sbjct: 237 AANNIPRSSGQGGENNFMLGWVL 259
>sptr|Q84UA0|Q84UA0 MADS4.
Length = 260
Score = 395 bits (1015), Expect = e-109
Identities = 208/260 (80%), Positives = 220/260 (84%), Gaps = 5/260 (1%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G TKTLERYQHCCYNAQDS+ SALSE+QSWYQEMSKL+AKFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GTTKTLERYQHCCYNAQDSS-SALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEG-CSNY 738
VKELQQLEKQLEC+LSQARQRKTQ+MMEQVEELRR ER LGE+NRQLKHKL+AEG SN
Sbjct: 120 VKELQQLEKQLECSLSQARQRKTQLMMEQVEELRRKERQLGEINRQLKHKLDAEGSSSNS 179
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVH---PPAHSVAMDCEPTLQIGYPHHQFPPPEAVN 909
W A GT+V+ Y +H P HS AMDCEPTLQIGYPH +A N
Sbjct: 180 FRAMQQITWAA--GTVVDEGAGAYHMHHQQQPNHSAAMDCEPTLQIGYPHQFAAAEQAAN 237
Query: 910 NIPRSAAT-GENNFMLGWVL 966
NIPRS+A GENNFMLGWVL
Sbjct: 238 NIPRSSAPGGENNFMLGWVL 257
>sptr|O82129|O82129 MADS box transcription factor.
Length = 258
Score = 393 bits (1009), Expect = e-108
Identities = 211/263 (80%), Positives = 225/263 (85%), Gaps = 8/263 (3%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G TKTLERYQHCCYNAQDSN ALSE+QSWYQEMSKL+AKFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GTTKTLERYQHCCYNAQDSNG-ALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEG--CSN 735
VKELQQLEKQLEC+LS ARQRKTQ+MMEQVEELRR ER LG++NRQLKHKL+AEG +N
Sbjct: 120 VKELQQLEKQLECSLSLARQRKTQLMMEQVEELRRKERQLGDINRQLKHKLDAEGSNSNN 179
Query: 736 YTTLQHAACWPAPGGTIVEHDGATY----QVHPPAHSVAMDCEPTLQIGYPHHQF-PPPE 900
Y +Q + W A GT+V+ A Y Q P HS AMDCEPTLQIGY HHQF P +
Sbjct: 180 YRAMQQIS-WAA--GTVVDEGAAAYHMQQQQQHPNHSAAMDCEPTLQIGY-HHQFAAPDQ 235
Query: 901 AVNNIPRSAAT-GENNFMLGWVL 966
A NNIPRS+A GENNFMLGWVL
Sbjct: 236 AANNIPRSSAPGGENNFMLGWVL 258
>sptr|O24228|O24228 MADS box protein.
Length = 250
Score = 392 bits (1006), Expect = e-108
Identities = 214/258 (82%), Positives = 223/258 (86%), Gaps = 3/258 (1%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+TKTLERYQHCCYNAQDSNN ALSE+QSWY EMSKL+AKFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GITKTLERYQHCCYNAQDSNN-ALSETQSWYHEMSKLKAKFEALQRTQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEG-CSNY 738
VKELQQLEKQLECALSQARQRKTQ+MMEQVEELRR ER LGE+NRQLKHKLE EG SNY
Sbjct: 120 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERQLGEINRQLKHKLEVEGSTSNY 179
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
+Q A+ W G +VE +GA Y V PP HS AMD EPTLQIGYP HQF P EA N I
Sbjct: 180 RAMQQAS-WAQ--GAVVE-NGAAY-VQPPPHSAAMDSEPTLQIGYP-HQFVPAEA-NTIQ 232
Query: 919 RSAAT--GENNFMLGWVL 966
RS A ENNFMLGWVL
Sbjct: 233 RSTAPAGAENNFMLGWVL 250
>sptr|Q84L85|Q84L85 MADS-box transcription factor SEP1.
Length = 243
Score = 330 bits (847), Expect = 2e-89
Identities = 176/255 (69%), Positives = 205/255 (80%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G +KTLERYQ CCY +QD+ A E Q+WYQE+++L+AKFE+LQ QRHLLGEDLGPLS
Sbjct: 61 GTSKTLERYQRCCYTSQDA-TIADREKQNWYQEVARLKAKFESLQSAQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQQLE+QLE +LSQARQRKTQ+M +Q+EELR+ E HLGE+N+QLK KLEAEG N
Sbjct: 120 VKELQQLERQLEASLSQARQRKTQIMFDQMEELRKKEHHLGEINKQLKTKLEAEG-ENLR 178
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPR 921
+Q W + + G + +H P+HS AM+CEPTLQIGY HQ PE ++PR
Sbjct: 179 AIQ--GSWESDATNV--GGGNVFSMH-PSHSSAMECEPTLQIGY--HQLVQPE--GSLPR 229
Query: 922 SAATGENNFMLGWVL 966
++ GENNFMLGWVL
Sbjct: 230 NSG-GENNFMLGWVL 243
>sptrnew|CAD41169|CAD41169 OSJNBa0064M23.14 protein.
Length = 249
Score = 324 bits (831), Expect = 1e-87
Identities = 178/259 (68%), Positives = 200/259 (77%), Gaps = 4/259 (1%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNS-ALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPL 558
G+ KTLE+Y CCYNAQ SN++ A E QSWYQEMS+L+ K E LQR+QRH+LGEDLGPL
Sbjct: 61 GINKTLEKYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPL 120
Query: 559 SVKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNY 738
S+KELQQLEKQLE +LSQARQRKTQ+MMEQV++LRR ER LGE+N+QLK+KLEAE S+
Sbjct: 121 SIKELQQLEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSN 180
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
W GT+V G PP +DCEPTLQIGY +QF PEA N P
Sbjct: 181 CRSAIQDSW--VHGTVVS-GGRVLNAQPPPD---IDCEPTLQIGY--YQFVRPEAAN--P 230
Query: 919 RSAATG---ENNFMLGWVL 966
RS G NNF++GW L
Sbjct: 231 RSNGGGGDQNNNFVMGWPL 249
>sptr|Q9SES9|Q9SES9 MADS box transcription factor MADS17.
Length = 249
Score = 324 bits (831), Expect = 1e-87
Identities = 178/259 (68%), Positives = 200/259 (77%), Gaps = 4/259 (1%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNS-ALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPL 558
G+ KTLE+Y CCYNAQ SN++ A E QSWYQEMS+L+ K E LQR+QRH+LGEDLGPL
Sbjct: 61 GINKTLEKYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPL 120
Query: 559 SVKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNY 738
S+KELQQLEKQLE +LSQARQRKTQ+MMEQV++LRR ER LGE+N+QLK+KLEAE S+
Sbjct: 121 SIKELQQLEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSN 180
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
W GT+V G PP +DCEPTLQIGY +QF PEA N P
Sbjct: 181 CRSAIQDSW--VHGTVVS-GGRVLNAQPPPD---IDCEPTLQIGY--YQFVRPEAAN--P 230
Query: 919 RSAATG---ENNFMLGWVL 966
RS G NNF++GW L
Sbjct: 231 RSNGGGGDQNNNFVMGWPL 249
>sptr|Q8LLR1|Q8LLR1 MADS-box protein 3.
Length = 244
Score = 311 bits (797), Expect = 1e-83
Identities = 175/257 (68%), Positives = 201/257 (78%), Gaps = 2/257 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G TKTLERYQ CY QD+N E+QSWYQE+SKL+AK+E+LQRTQRHLLGEDLGPLS
Sbjct: 61 GTTKTLERYQRVCYTPQDNNMEC--ETQSWYQEVSKLKAKYESLQRTQRHLLGEDLGPLS 118
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQ LEKQLE AL+QARQRKTQ+M+EQ+E+LRR ER LG++N+QLK KLEAEG +
Sbjct: 119 VKELQNLEKQLEGALAQARQRKTQMMIEQMEDLRRKERQLGDLNKQLKLKLEAEG-QSLK 177
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC--EPTLQIGYPHHQFPPPEAVNNI 915
+Q W T +++ VH P+ S MDC EP LQIGY H + P E ++
Sbjct: 178 AIQ--GSWNPSTATA---GNSSFPVH-PSQSNPMDCEPEPILQIGY--HHYVPAEG-PSV 228
Query: 916 PRSAATGENNFMLGWVL 966
+S A GE+NF+ GWVL
Sbjct: 229 SKSMA-GESNFIQGWVL 244
>sptr|O82699|O82699 MADS-box protein.
Length = 243
Score = 304 bits (778), Expect = 2e-81
Identities = 167/257 (64%), Positives = 201/257 (78%), Gaps = 2/257 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFS RGKLYEF SA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRGKLYEFASA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G++KTLERYQ C + + NS E+QSWYQE++KL+AK+E+LQRTQRHLLGEDLGPLS
Sbjct: 61 GMSKTLERYQRCSFTPPE--NSIERETQSWYQEVTKLKAKYESLQRTQRHLLGEDLGPLS 118
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQ LEKQLE AL+Q RQRKTQ+M+EQ+E+LR+ ERHLG++N+QL+ KLEAEG N
Sbjct: 119 VKELQNLEKQLEGALAQTRQRKTQLMIEQMEDLRKKERHLGDLNKQLRVKLEAEG-QNLN 177
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC--EPTLQIGYPHHQFPPPEAVNNI 915
+Q+ A G+ + + +H + + MDC EP +Q+GY HQ+ P E ++I
Sbjct: 178 VIQNMWSSDAAAGS------SNFSLH-SSQTNPMDCTPEPVIQMGY--HQYHPAEG-SSI 227
Query: 916 PRSAATGENNFMLGWVL 966
PRS TGE NF+ GWVL
Sbjct: 228 PRS-LTGETNFIQGWVL 243
>sptrnew|BAC80256|BAC80256 MADS-box transcription factor (Fragment).
Length = 227
Score = 300 bits (768), Expect = 3e-80
Identities = 160/244 (65%), Positives = 192/244 (78%)
Frame = +1
Query: 235 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKTLERYQH 414
ENKINRQVTFSKRRNGLLKKAYELS+LCDAE+ALIIFS RGKLYEFGS+G+TKTLERYQ
Sbjct: 1 ENKINRQVTFSKRRNGLLKKAYELSILCDAEIALIIFSSRGKLYEFGSSGLTKTLERYQR 60
Query: 415 CCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQQLEKQL 594
C Y Q+ NN A E+Q W+QE+SKL+AK+E L R+QRHLLGEDLGPLSVKELQQLE+QL
Sbjct: 61 CSYVPQE-NNPADREAQVWHQEISKLKAKYELLLRSQRHLLGEDLGPLSVKELQQLERQL 119
Query: 595 ECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQHAACWPAP 774
E ALSQARQRKTQ+MMEQ+EELR+ ER LG++N+QLK KLEAEG ++++Q A W +
Sbjct: 120 EVALSQARQRKTQIMMEQMEELRKKERCLGDINKQLKGKLEAEGIGAFSSIQGA--WESA 177
Query: 775 GGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPRSAATGENNFML 954
+V P+ S +DCEP+LQIGY HQF P EA +P +A GE+NF+
Sbjct: 178 APVVVH----------PSQSADVDCEPSLQIGY--HQFVPQEAA--MPCRSAGGESNFIQ 223
Query: 955 GWVL 966
GW+L
Sbjct: 224 GWML 227
>sptr|Q9MB91|Q9MB91 PMADS4 protein.
Length = 253
Score = 296 bits (758), Expect = 4e-79
Identities = 169/262 (64%), Positives = 190/262 (72%), Gaps = 7/262 (2%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFG+A
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGNA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+TKTLERYQ CC N QD N E+QSWYQE+SKL+ KFEALQRTQRHLLGEDLGPLS
Sbjct: 61 GITKTLERYQRCCLNPQD--NCGERETQSWYQEVSKLKGKFEALQRTQRHLLGEDLGPLS 118
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHK-------LEA 720
VKELQ LEKQLE AL+QARQRKTQ+MMEQ+EELRR ERHLG++N+QLK K LEA
Sbjct: 119 VKELQNLEKQLEGALAQARQRKTQIMMEQMEELRRKERHLGDVNKQLKVKVSLELSSLEA 178
Query: 721 EGCSNYTTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPE 900
EG + W + +E + + VH + S MDC P + Y H + +
Sbjct: 179 EGQAGLNRAL-PFLWTS---NALEAGNSNFPVH-HSQSNPMDCGPDPVLQYRDHHYMAAD 233
Query: 901 AVNNIPRSAATGENNFMLGWVL 966
+ RS A E N M GW L
Sbjct: 234 GPSG-SRSMAV-ETNIMQGWGL 253
>sptr|Q948V2|Q948V2 Putative MADS-domain transcription factor
MpMADS3 (Fragment).
Length = 231
Score = 295 bits (755), Expect = 9e-79
Identities = 157/244 (64%), Positives = 193/244 (79%)
Frame = +1
Query: 235 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKTLERYQH 414
ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FS RGKLYEFGSAG+ KTLERYQ
Sbjct: 1 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSAGINKTLERYQR 60
Query: 415 CCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQQLEKQL 594
CCY D+N + ++Q WYQE+SKL AK ++LQR+QRHLLGEDLGPLSVKELQ+LE+QL
Sbjct: 61 CCYTFHDANITD-RDTQGWYQEVSKLNAKCDSLQRSQRHLLGEDLGPLSVKELQKLERQL 119
Query: 595 ECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQHAACWPAP 774
E AL+Q RQRKTQ+M+E +E LR ER LG++N++LK+KLEA+G + +Q A W +
Sbjct: 120 ESALTQTRQRKTQIMLEHMEALREKERQLGDINKELKNKLEAKGQGAFRAMQ--ASWES- 176
Query: 775 GGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPRSAATGENNFML 954
G +V ++G + +H P+ S A++CEPTLQIGY H F PEA NIPR+ E+NFM
Sbjct: 177 -GPLVGNNG--FPMH-PSQSAAIECEPTLQIGY--HSFAAPEA--NIPRT-VVAESNFMH 227
Query: 955 GWVL 966
GW+L
Sbjct: 228 GWIL 231
>sptr|Q9AUJ0|Q9AUJ0 MADS box protein nmads3 (Fragment).
Length = 236
Score = 289 bits (740), Expect = 5e-77
Identities = 156/227 (68%), Positives = 177/227 (77%), Gaps = 1/227 (0%)
Frame = +1
Query: 223 LKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKTLE 402
+K IENKI RQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSAG+ KTLE
Sbjct: 1 IKSIENKITRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGINKTLE 60
Query: 403 RYQHCCYNAQDSNNS-ALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQQ 579
+Y CCYNAQ SN++ A E QSWYQEMS+L+ K E LQR+QRH+LGEDLGPLS+KELQQ
Sbjct: 61 KYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPLSIKELQQ 120
Query: 580 LEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQHAA 759
LEKQLE +LSQARQRKTQ+MMEQV++LRR ER LGE+N+QLK+KLEAE S+
Sbjct: 121 LEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSNCRSAIQD 180
Query: 760 CWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPE 900
W GT+V G PP +DCEPTLQIGY +QF PE
Sbjct: 181 SW--VHGTVVS-GGRVLNAQPPPD---IDCEPTLQIGY--YQFVRPE 219
>sptr|Q40970|Q40970 Putative MADS-box family transcription factor.
Length = 242
Score = 283 bits (725), Expect = 3e-75
Identities = 160/256 (62%), Positives = 190/256 (74%), Gaps = 1/256 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+ KTLERYQ C Y QD+ S E+Q+W+QE+ KL+A+ E LQR+QRHLLGEDLGPLS
Sbjct: 61 GMLKTLERYQKCSYVLQDATVSD-REAQNWHQEVGKLKARVELLQRSQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
+KELQQLE+QLE AL+ R RKTQVM+E ++ELRR ER L E+N+ L+ KL EAEG +
Sbjct: 120 IKELQQLERQLEVALTHVRSRKTQVMLEMMDELRRKERILQEVNKSLRKKLQEAEGQAFN 179
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
W + + A HP S A+DCEPTLQIGY Q+ PPE +++P
Sbjct: 180 AMQPPPHAWDSHAVA----NNAYAMQHP---SNAVDCEPTLQIGY---QYAPPE--SSMP 227
Query: 919 RSAATGENNFMLGWVL 966
R +NN+M GW++
Sbjct: 228 RH-EQAQNNYMQGWMV 242
>sptrnew|AAP83412|AAP83412 AGL6-like MADS-box (Fragment).
Length = 242
Score = 282 bits (722), Expect = 6e-75
Identities = 142/222 (63%), Positives = 173/222 (77%)
Frame = +1
Query: 217 VELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKT 396
V+L+R+ENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFS RGKLYEFGSAG+T T
Sbjct: 2 VQLRRMENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRGKLYEFGSAGMTAT 61
Query: 397 LERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQ 576
LERYQ CC+N Q++ A E+QSWYQE+SKL+AK+E+LQRTQRHLLGEDLGPL+VKEL+
Sbjct: 62 LERYQRCCFNPQNAGAGAERETQSWYQEVSKLKAKYESLQRTQRHLLGEDLGPLNVKELE 121
Query: 577 QLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQHA 756
LEKQLE +LSQARQRKT++MMEQ+E+LRR ER LGEMN+QLK ++ E S+ T Q
Sbjct: 122 NLEKQLEGSLSQARQRKTKIMMEQMEDLRRKERQLGEMNKQLKIRVSLELSSHETEGQGL 181
Query: 757 ACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHH 882
+P + + P + + + EP LQ+GY H+
Sbjct: 182 RGFPCQXNAAASAGTSNFMAQPGTNPMEFEPEPVLQMGYHHY 223
>sptr|P93468|P93468 MADS-box family transcription factor.
Length = 242
Score = 281 bits (720), Expect = 1e-74
Identities = 159/256 (62%), Positives = 189/256 (73%), Gaps = 1/256 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+ KTLERYQ C Y QD+ S E+Q+W+QE+ KL+A+ E LQR+QRHLLGEDLGPLS
Sbjct: 61 GMLKTLERYQKCSYVLQDATVSD-REAQNWHQEVGKLKARVELLQRSQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
+KELQQLE+QLE AL+ R RKTQVM+E ++ELRR ER L E+N+ L+ KL EAEG +
Sbjct: 120 IKELQQLERQLEVALTHVRSRKTQVMLEMMDELRRKERILQEVNKSLRKKLQEAEGQAFN 179
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
W + + A HP S A+DCEPTLQ GY Q+ PPE +++P
Sbjct: 180 AMQPPPHAWDSHAVA----NNAYAMQHP---SNAVDCEPTLQTGY---QYAPPE--SSMP 227
Query: 919 RSAATGENNFMLGWVL 966
R +NN+M GW++
Sbjct: 228 RH-EQAQNNYMQGWMV 242
>sptr|Q84LC8|Q84LC8 MADS-box transcription factor CDM104.
Length = 250
Score = 280 bits (717), Expect = 2e-74
Identities = 163/265 (61%), Positives = 186/265 (70%), Gaps = 10/265 (3%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFS R KLYEFGS
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRDKLYEFGSV 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
GV KTLERYQ CC+N QD+NN E+QSWYQE+SKL+AKFE+LQRTQRHLLGEDLGPLS
Sbjct: 61 GVMKTLERYQRCCFNPQDNNNE--RETQSWYQEVSKLKAKFESLQRTQRHLLGEDLGPLS 118
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-------EA 720
VKEL LEKQLE AL+QARQRKTQ+++EQ+EELR ER LG+MN+ LK K+ E
Sbjct: 119 VKELHNLEKQLEGALTQARQRKTQILVEQMEELRCKERELGDMNKHLKIKVSHELSTFET 178
Query: 721 EGCSNYTTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC---EPTLQIGYPHHQFP 891
EG Y T Q W + T+ +H P+ S M C +P LQIG +QF
Sbjct: 179 EG-QGYRTHQLPCPWNSSNNN-------TFLMH-PSQSNPMGCQQEQPILQIG--DNQFM 227
Query: 892 PPEAVNNIPRSAATGENNFMLGWVL 966
E + + E N WV+
Sbjct: 228 QGEGSSG--QRNMVDETNIHGNWVI 250
>sw|P29386|AGL6_ARATH Agamous-like MADS box protein AGL6.
Length = 252
Score = 278 bits (711), Expect = 1e-73
Identities = 149/257 (57%), Positives = 184/257 (71%), Gaps = 2/257 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSV 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G+ T+ERY C YN SNN +QSW QE++KL++K+E+L RT R+LLGEDLG +
Sbjct: 61 GIESTIERYNRC-YNCSLSNNKPEETTQSWCQEVTKLKSKYESLVRTNRNLLGEDLGEMG 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VKELQ LE+QLE AL+ RQRKTQVMME++E+LR+ ER LG++N+QLK K E EG + +
Sbjct: 120 VKELQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFETEGHA-FK 178
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC--EPTLQIGYPHHQFPPPEAVNNI 915
T Q W ++ + P+H +DC EP LQIG+ H + E +++
Sbjct: 179 TFQD--LWANSAASVAGDPNNSEFPVEPSHPNVLDCNTEPFLQIGFQQHYYVQGEG-SSV 235
Query: 916 PRSAATGENNFMLGWVL 966
+S GE NF+ GWVL
Sbjct: 236 SKSNVAGETNFVQGWVL 252
>sptr|Q8GTE8|Q8GTE8 MADS-box protein AGL6-a.
Length = 252
Score = 275 bits (702), Expect = 1e-72
Identities = 145/257 (56%), Positives = 182/257 (70%), Gaps = 2/257 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FS RGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSV 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
GV +T+ERY H CYN +NN Q+W QE++KL+AK+E+L RT RHLLGED+G +
Sbjct: 61 GVERTIERY-HRCYNCSVTNNRPEESKQNWCQEVAKLKAKYESLVRTNRHLLGEDIGEMG 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VK+LQ LE+QLE AL+ RQRKTQVMME++E+LR+ ER LG++N+QLK K EA G +
Sbjct: 120 VKQLQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFEAGG---HA 176
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC--EPTLQIGYPHHQFPPPEAVNNI 915
WP ++ + P+H ++DC EP LQIG+ H + E +++
Sbjct: 177 FKSFQDFWPNSAASMAGDPNNSKFPVQPSHPDSVDCNTEPFLQIGFQQHYYVQGEG-SSV 235
Query: 916 PRSAATGENNFMLGWVL 966
P+S E NF+ W L
Sbjct: 236 PKSNVACETNFVQDWFL 252
>sptr|Q948V1|Q948V1 Putative MADS-domain transcription factor
MpMADS4 (Fragment).
Length = 248
Score = 273 bits (697), Expect = 5e-72
Identities = 150/226 (66%), Positives = 176/226 (77%)
Frame = +1
Query: 247 NRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKTLERYQHCCYN 426
NRQVTF KRRNGLLKKAYELSVLCDAEVALIIFS RGKLY FGSAG KT ERYQ CCY
Sbjct: 1 NRQVTFCKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYGFGSAGTNKTPERYQRCCYT 60
Query: 427 AQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQQLEKQLECAL 606
QD S E+Q WYQE+S+L+AK+E+LQR+QRHLLGEDLGPLSVKELQ LE+QLE AL
Sbjct: 61 PQDVVVSD-RETQGWYQEVSRLKAKYESLQRSQRHLLGEDLGPLSVKELQHLERQLEVAL 119
Query: 607 SQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQHAACWPAPGGTI 786
SQARQRKTQ+M+EQ+EELR+ ER LG++N+QLK KLEAEG + +Q W + G +
Sbjct: 120 SQARQRKTQIMIEQMEELRKKERQLGDINKQLKIKLEAEGQGPFRCIQ--GSWES--GAM 175
Query: 787 VEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIPRS 924
V ++ + ++ P + M+CEPTLQIGY H F PPEA NIPRS
Sbjct: 176 VGNN--NFSMNAP-QAAPMECEPTLQIGY--HHFVPPEA--NIPRS 214
>sptr|Q8GTE9|Q8GTE9 MADS-box protein AGL6-a.
Length = 259
Score = 268 bits (684), Expect = 2e-70
Identities = 145/264 (54%), Positives = 182/264 (68%), Gaps = 9/264 (3%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FS RGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSV 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
GV +T+ERY H CYN +NN Q+W QE++KL+AK+E+L RT RHLLGED+G +
Sbjct: 61 GVERTIERY-HRCYNCSVTNNRPEESKQNWCQEVAKLKAKYESLVRTNRHLLGEDIGEMG 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
VK+LQ LE+QLE AL+ RQRKTQVMME++E+LR+ ER LG++N+QLK K EA G +
Sbjct: 120 VKQLQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFEAGG---HA 176
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDC--EPTLQIG-------YPHHQFPP 894
WP ++ + P+H ++DC EP LQIG + H +
Sbjct: 177 FKSFQDFWPNSAASMAGDPNNSKFPVQPSHPDSVDCNTEPFLQIGLVLVYIRFQQHYYVQ 236
Query: 895 PEAVNNIPRSAATGENNFMLGWVL 966
E +++P+S E NF+ W L
Sbjct: 237 GEG-SSVPKSNVACETNFVQDWFL 259
>sptr|Q40765|Q40765 Dal1 protein.
Length = 261
Score = 267 bits (683), Expect = 2e-70
Identities = 153/265 (57%), Positives = 189/265 (71%), Gaps = 12/265 (4%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
+ KTLERY+ C Y QD+ + E+Q+W+QE++KL+ K E LQR+QRHLLGEDLGPL+
Sbjct: 61 SMNKTLERYEKCSYAMQDTTGVSDREAQNWHQEVTKLKGKVELLQRSQRHLLGEDLGPLN 120
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
VKELQQLE+QLE AL+ R RKTQVM++Q+EELR+ ER L E+N+ L+ KL E EG
Sbjct: 121 VKELQQLERQLEVALAHLRSRKTQVMLDQIEELRQRERLLHEVNKSLQKKLSETEGRDVI 180
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQV-----HPPAHSVA----MDCEPTLQIGYPHHQFP 891
T ++ + GT D + HP +S A +DCEPTLQIGY Q
Sbjct: 181 TGIEQTS--NTNTGTNGPWDSSITNTAYALSHPQQNSNASLHHVDCEPTLQIGY---QPV 235
Query: 892 PPEAVN--NIPRSAATGENNFMLGW 960
PPE++ + P+ T +N +M GW
Sbjct: 236 PPESIGPPHQPQHNQT-QNQYMQGW 259
>sptrnew|AAP83382|AAP83382 AGL6-like MADS-box (Fragment).
Length = 206
Score = 265 bits (678), Expect = 8e-70
Identities = 136/178 (76%), Positives = 154/178 (86%)
Frame = +1
Query: 217 VELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSAGVTKT 396
V+LKR+ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSAG KT
Sbjct: 1 VQLKRMENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGTNKT 60
Query: 397 LERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLSVKELQ 576
LERYQ CCY QD S E+Q WYQE+SKL+AK+E+LQR+QRHLL EDLGPLSVKELQ
Sbjct: 61 LERYQRCCYTPQDVVVSD-RETQGWYQEVSKLKAKYESLQRSQRHLLXEDLGPLSVKELQ 119
Query: 577 QLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYTTLQ 750
LE+QLE ALSQARQRKTQ+M+EQ+EELR+ ER LG++N+QLK KLEAEG + +Q
Sbjct: 120 HLERQLEVALSQARQRKTQIMIEQMEELRKKERQLGDINKQLKIKLEAEGQGPFRCIQ 177
>sptr|O04406|O04406 MADS-box protein.
Length = 261
Score = 262 bits (670), Expect = 7e-69
Identities = 150/267 (56%), Positives = 183/267 (68%), Gaps = 14/267 (5%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
+ KTLERY+ C Y QD+ + E+Q+W+QE++KL+ K E LQR+QRHLLGEDLGPL+
Sbjct: 61 SMNKTLERYEKCSYAMQDTTGVSDREAQNWHQEVTKLKGKVELLQRSQRHLLGEDLGPLN 120
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
VKELQQLE+QLE AL+ R RKTQVM++Q+EELR+ ER L E+N+ L+ KL E EG
Sbjct: 121 VKELQQLERQLEVALTHLRSRKTQVMLDQIEELRQRERLLHEVNKSLQKKLSETEGRDVI 180
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQV-----HPPAHSVA----MDCEPTLQIGY----PH 879
T ++ + GT D + HP S + +DCEPTLQIGY P
Sbjct: 181 TGIEQTS--NTNTGTNGPWDSSITNTAYALSHPQQDSNSSLHHVDCEPTLQIGYQPVAPE 238
Query: 880 HQFPPPEAVNNIPRSAATGENNFMLGW 960
PP + +N N +M GW
Sbjct: 239 SIVPPHQPPHN------QTPNQYMQGW 259
>sptr|Q9XGJ6|Q9XGJ6 Putative MADS domain transcription factor GGM11.
Length = 254
Score = 262 bits (670), Expect = 7e-69
Identities = 140/223 (62%), Positives = 168/223 (75%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
G KTLERYQ C Y Q+SNNS ++Q+W+ E+SKL+ K E LQR+QRHLLGEDLGPLS
Sbjct: 61 GTLKTLERYQKCSYALQESNNSD-RDAQTWHHEVSKLKTKVEILQRSQRHLLGEDLGPLS 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 741
++ELQ LE+Q+E AL+Q R RKTQVMM+ +++L++ ER L E+N+ L+ KL+ Y+
Sbjct: 120 IRELQTLERQIEVALTQVRARKTQVMMDMMDDLKKKERLLQEVNKSLRKKLDETEGQVYS 179
Query: 742 TLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIG 870
Q A P P Y + PP A+DCEPT ++G
Sbjct: 180 NAQLQAA-PPPEWDSNAIANPVYAL-PPTPQNAVDCEPTCKLG 220
>sptr|Q9ST06|Q9ST06 GpMADS3 protein.
Length = 252
Score = 251 bits (640), Expect = 2e-65
Identities = 137/255 (53%), Positives = 178/255 (69%), Gaps = 1/255 (0%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
++KTLERY+ C Y+ Q+ N S ++Q+W+ E++KL+AK E+L + QR+L+GEDLGPL+
Sbjct: 61 SMSKTLERYEKCSYSMQE-NASTDRDAQNWHHEVTKLKAKLESLHKAQRNLMGEDLGPLN 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
+KELQ LE+QLE AL R RKTQ++++ ++ELR ER L E+N+ L+ KL E EG +
Sbjct: 120 IKELQSLEQQLEVALGHVRNRKTQLLIQTIDELRDKERTLQEVNKSLQKKLSETEGRNVI 179
Query: 739 TTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAVNNIP 918
L HA + Q P+ +DCEPTL IG P+HQ EA+ P
Sbjct: 180 PALPHATTGSGEWESSTLTTTYALQTQQPSGIHHVDCEPTLHIG-PYHQAVHHEAIPGPP 238
Query: 919 RSAATGENNFMLGWV 963
+ + N + + WV
Sbjct: 239 ATHSEPHNQY-IWWV 252
>sptr|Q9XGJ8|Q9XGJ8 Putative MADS domain transcription factor GGM9.
Length = 253
Score = 248 bits (634), Expect = 1e-64
Identities = 140/259 (54%), Positives = 180/259 (69%), Gaps = 5/259 (1%)
Frame = +1
Query: 202 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 381
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 382 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 561
++KTLERY+ C Y+ Q+ N S ++Q+W+ E++KL+AK E+L + QR L+GEDLGPL+
Sbjct: 61 SMSKTLERYEKCSYSMQE-NASTDRDAQNWHHEVTKLKAKLESLHKAQRSLMGEDLGPLN 119
Query: 562 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKL-EAEGCSNY 738
+KELQ LE+QLE AL R RKTQ++++ ++ELR ER L E+N+ L+ KL E EG +
Sbjct: 120 IKELQSLEQQLEVALGHVRNRKTQLLIQTIDELRDKERTLQEVNKSLQKKLSETEGRNVI 179
Query: 739 TTLQHA----ACWPAPGGTIVEHDGATYQVHPPAHSVAMDCEPTLQIGYPHHQFPPPEAV 906
L HA W + T + T Q H +DCEPTL IG P+HQ EA+
Sbjct: 180 PALPHAPNGSGEWESSTLTTTTYALQTQQTSNIHH---VDCEPTLHIG-PYHQAVHHEAI 235
Query: 907 NNIPRSAATGENNFMLGWV 963
P + + N + + WV
Sbjct: 236 TAPPATHSEPHNQY-IWWV 253
>sptrnew|AAP83381|AAP83381 AGL6-like MADS-box (Fragment).
Length = 214
Score = 243 bits (621), Expect = 3e-63
Identities = 131/217 (60%), Positives = 169/217 (77%)
Frame = +1
Query: 301 ELSVLCDAEVALIIFSGRGKLYEFGSAGVTKTLERYQHCCYNAQDSNNSALSESQSWYQE 480
E+S+LCDAEVALI+FS RGKLYEFGSAGV KTLERYQ CCY QD+N + ++Q WYQE
Sbjct: 5 EMSMLCDAEVALIVFSSRGKLYEFGSAGVNKTLERYQRCCYTFQDANITD-RDTQGWYQE 63
Query: 481 MSKLRAKFEALQRTQRHLLGEDLGPLSVKELQQLEKQLECALSQARQRKTQVMMEQVEEL 660
+SKL+AK ++LQR+QRHLLGEDLGPLSVKELQ+LE+QLE AL+Q RQRKTQ+M+E +E L
Sbjct: 64 VSKLKAKCDSLQRSQRHLLGEDLGPLSVKELQKLERQLESALTQTRQRKTQIMLEHMEAL 123
Query: 661 RRTERHLGEMNRQLKHKLEAEGCSNYTTLQHAACWPAPGGTIVEHDGATYQVHPPAHSVA 840
R ER LG++N++LK+KLEA+G + +Q A W + G +V ++G + +H P+ S A
Sbjct: 124 REKERQLGDINKELKNKLEAKGQGAFRAMQ--ASWES--GPLVGNNG--FPMH-PSQSAA 176
Query: 841 MDCEPTLQIGYPHHQFPPPEAVNNIPRSAATGENNFM 951
++CEPTLQIGY H F PEA +IPR+ E+NFM
Sbjct: 177 IECEPTLQIGY--HSFAAPEA--SIPRT-VVAESNFM 208
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 12, 2003 8:28 PM
Number of letters in database: 397,940,306
Number of sequences in database: 1,242,019
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 884,406,961
Number of Sequences: 1242019
Number of extensions: 19439753
Number of successful extensions: 64892
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 59146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 63818
length of database: 397,940,306
effective HSP length: 125
effective length of database: 242,687,931
effective search space used: 71592939645
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

