BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
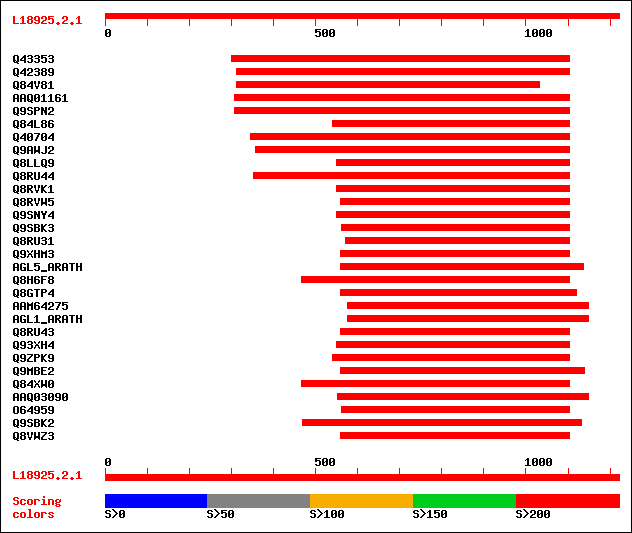
Query= L18925.2.1
(1221 letters)
Database: /db/trembl-ebi/tmp/swall
1,242,019 sequences; 397,940,306 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q43353|Q43353 ZAG2 gene. 517 e-145
sptr|Q42389|Q42389 MADS box protein. 474 e-132
sptr|Q84V81|Q84V81 Putative MADS-domain transcription factor (Fr... 428 e-119
sptrnew|AAQ01161|AAQ01161 MADS protein. 347 1e-94
sptr|Q9SPN2|Q9SPN2 Transcription factor. 347 3e-94
sptr|Q84L86|Q84L86 MADS-box transcription factor AG. 261 1e-68
sptr|Q40704|Q40704 MADS box protein. 254 2e-66
sptr|Q9AWJ2|Q9AWJ2 Putative MADS-box protein. 252 9e-66
sptr|Q8LLQ9|Q8LLQ9 MADS-box protein 5. 249 4e-65
sptr|Q8RU44|Q8RU44 Agamous-like protein 1 HvAG1. 249 4e-65
sptr|Q8RVK1|Q8RVK1 AP3-like protein. 249 6e-65
sptr|Q8RVW5|Q8RVW5 MADS-box transcription factor. 248 9e-65
sptr|Q9SNY4|Q9SNY4 Transcription factor MADS1. 247 2e-64
sptr|Q9SBK3|Q9SBK3 Agamous-like putative transcription factor. 247 2e-64
sptr|Q8RU31|Q8RU31 Putative transcription factor agamous. 247 3e-64
sptr|Q9XHM3|Q9XHM3 Agamous homolog (Fragment). 247 3e-64
sw|P29385|AGL5_ARATH Agamous-like MADS box protein AGL5. 245 8e-64
sptr|Q8H6F8|Q8H6F8 MADS box protein GHMADS-2. 245 8e-64
sptr|Q8GTP4|Q8GTP4 MADS box transcription factor. 244 1e-63
sptrnew|AAM64275|AAM64275 Shatterproof 1 (SHP1)/ agamous-like 1 ... 244 1e-63
sw|P29381|AGL1_ARATH Agamous-like MADS box protein AGL1 (Protein... 244 1e-63
sptr|Q8RU43|Q8RU43 Agamous-like protein 2 HvAG2. 244 1e-63
sptr|Q93XH4|Q93XH4 MAD-box transcripion factor. 244 2e-63
sptr|Q9ZPK9|Q9ZPK9 Agamous homolog transcription factor. 243 4e-63
sptr|Q9MBE2|Q9MBE2 MADS-box protein. 242 7e-63
sptr|Q84XW0|Q84XW0 Mads-box transcription factor. 242 7e-63
sptrnew|AAQ03090|AAQ03090 AGAMOUS-like protein. 241 1e-62
sptr|O64959|O64959 CUM10. 241 1e-62
sptr|Q9SBK2|Q9SBK2 Agamous-like putative transcription factor. 241 2e-62
sptr|Q8VWZ3|Q8VWZ3 C-type MADS box protein. 240 3e-62
>sptr|Q43353|Q43353 ZAG2 gene.
Length = 268
Score = 517 bits (1331), Expect = e-145
Identities = 263/268 (98%), Positives = 263/268 (98%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN
Sbjct: 1 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN
Sbjct: 61 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 120
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL
Sbjct: 121 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 180
Query: 564 QQVTVARSVXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQRQ 385
QQVTVARSV ATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQRQ
Sbjct: 181 QQVTVARSVAAAAAATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQRQ 240
Query: 384 MLATELNLGYQLAPPGSDAANNNPHHQF 301
MLATELNLGYQLAPPGSDAANNNPHHQF
Sbjct: 241 MLATELNLGYQLAPPGSDAANNNPHHQF 268
>sptr|Q42389|Q42389 MADS box protein.
Length = 265
Score = 474 bits (1220), Expect = e-132
Identities = 244/264 (92%), Positives = 251/264 (95%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVAL+VFSSRGRLYEYANN
Sbjct: 1 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALVVFSSRGRLYEYANN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVKAT+ERYKKAH VGSSSGPPLLEHNAQQFYQQES KLRNQIQMLQNTNRHLVGDSVGN
Sbjct: 61 SVKATIERYKKAHAVGSSSGPPLLEHNAQQFYQQESVKLRNQIQMLQNTNRHLVGDSVGN 120
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
LSLKELKQLESRLEKGISKIRARKSELLAAEI+YMAKRETELQNDHM LRTKIEEGEQQL
Sbjct: 121 LSLKELKQLESRLEKGISKIRARKSELLAAEINYMAKRETELQNDHMNLRTKIEEGEQQL 180
Query: 564 QQVTVARSVXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQRQ 385
QQVTVA+SV AT++ELNPFLEMDTKCFF GGPFATLDMKCF PGSL QMLEAQQRQ
Sbjct: 181 QQVTVAQSV-AAAAATDVELNPFLEMDTKCFFPGGPFATLDMKCFFPGSL-QMLEAQQRQ 238
Query: 384 MLATELNLGYQLAPPGSDAANNNP 313
MLATELNLGYQLAPP +D ANNNP
Sbjct: 239 MLATELNLGYQLAPPDTDVANNNP 262
>sptr|Q84V81|Q84V81 Putative MADS-domain transcription factor
(Fragment).
Length = 241
Score = 428 bits (1100), Expect = e-119
Identities = 221/240 (92%), Positives = 228/240 (95%)
Frame = -1
Query: 1032 RNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLL 853
RNGLLKKAYELSVLCDAEVAL+VFSSRGRLYEYANNSVKAT+ERYKKAH VGSSSGPPLL
Sbjct: 1 RNGLLKKAYELSVLCDAEVALVVFSSRGRLYEYANNSVKATIERYKKAHAVGSSSGPPLL 60
Query: 852 EHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARK 673
EHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARK
Sbjct: 61 EHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARK 120
Query: 672 SELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQLQQVTVARSVXXXXXATNLELNPFL 493
SELLAAEI+YMAKRETELQNDHM LRTKIEEGEQQLQQVTVA+SV AT++ELNPFL
Sbjct: 121 SELLAAEINYMAKRETELQNDHMNLRTKIEEGEQQLQQVTVAQSV-AAAAATDVELNPFL 179
Query: 492 EMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQRQMLATELNLGYQLAPPGSDAANNNP 313
EMDTKCFF GGPFATLDMKCF PGSL QMLEAQQRQMLATELNLGYQLAPP +D ANNNP
Sbjct: 180 EMDTKCFFPGGPFATLDMKCFFPGSL-QMLEAQQRQMLATELNLGYQLAPPDTDVANNNP 238
>sptrnew|AAQ01161|AAQ01161 MADS protein.
Length = 270
Score = 347 bits (891), Expect = 1e-94
Identities = 196/279 (70%), Positives = 220/279 (78%), Gaps = 13/279 (4%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIEN TSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGRIEIKRIENTTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 S-VKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVG 748
+ VKAT++RYKKAH GS+SG PL+E NAQQ+YQQESAKLR+QIQMLQNTN+HLVGD+V
Sbjct: 61 NNVKATIDRYKKAHACGSTSGAPLIEVNAQQYYQQESAKLRHQIQMLQNTNKHLVGDNVS 120
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
NLSLKELKQLESRLEKGISKIRARK+ELLA+EI+YMAKRE ELQND+M LRTKI E EQQ
Sbjct: 121 NLSLKELKQLESRLEKGISKIRARKNELLASEINYMAKREIELQNDNMDLRTKIAEEEQQ 180
Query: 567 LQQVTVARS----VXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSL----- 415
LQQVTVARS + A + NPF A LDMKCF P +L
Sbjct: 181 LQQVTVARSAAMELQAAAAAQQQQQNPF----------AVAAAQLDMKCFFPLNLFEAAA 230
Query: 414 -QQMLEAQQRQMLATELNLGY--QLAPPGSDAANNNPHH 307
Q + AQ++Q++ TELNLGY LA PG+ AA+ P H
Sbjct: 231 QVQAVAAQRQQIIPTELNLGYHHHLAIPGAAAADAPPPH 269
>sptr|Q9SPN2|Q9SPN2 Transcription factor.
Length = 270
Score = 347 bits (889), Expect = 3e-94
Identities = 195/279 (69%), Positives = 220/279 (78%), Gaps = 13/279 (4%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIEN TSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGRIEIKRIENTTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 S-VKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVG 748
+ VKAT++RYKKAH GS+SG PL+E NAQQ+YQQESAKLR+QIQMLQNTN+HLVGD+V
Sbjct: 61 NNVKATIDRYKKAHACGSTSGAPLIEVNAQQYYQQESAKLRHQIQMLQNTNKHLVGDNVS 120
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
NLSLKELKQLESRLEKGI+KIRARK+ELLA+EI+YMAKRE ELQND+M LRTKI E EQQ
Sbjct: 121 NLSLKELKQLESRLEKGIAKIRARKNELLASEINYMAKREIELQNDNMDLRTKIAEEEQQ 180
Query: 567 LQQVTVARS----VXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSL----- 415
LQQVTVARS + A + NPF A LDMKCF P +L
Sbjct: 181 LQQVTVARSAAMELQAAAAAQQQQQNPF----------AVAAAQLDMKCFFPLNLFEAAA 230
Query: 414 -QQMLEAQQRQMLATELNLGY--QLAPPGSDAANNNPHH 307
Q + AQ++Q++ TELNLGY LA PG+ AA+ P H
Sbjct: 231 QVQAVAAQRQQIIPTELNLGYHHHLAIPGATAADAPPPH 269
>sptr|Q84L86|Q84L86 MADS-box transcription factor AG.
Length = 235
Score = 261 bits (667), Expect = 1e-68
Identities = 134/188 (71%), Positives = 163/188 (86%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS+RGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
S+K+T+ERYKKA SS+ ++E N QQ+YQQE+AKLR+QIQ LQN+NRHL+GDS+ +
Sbjct: 61 SIKSTIERYKKA-CADSSNSTAVVEVNTQQYYQQEAAKLRHQIQSLQNSNRHLMGDSLSS 119
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
LS+KELKQLE+RLE+GI++IR++K ELL AEI YM KRE ELQND+M LR KI + E +
Sbjct: 120 LSIKELKQLENRLERGITRIRSKKHELLFAEIEYMQKREAELQNDNMYLRAKITDNE-RA 178
Query: 564 QQVTVARS 541
QV+V +S
Sbjct: 179 HQVSVVQS 186
>sptr|Q40704|Q40704 MADS box protein.
Length = 236
Score = 254 bits (648), Expect = 2e-66
Identities = 147/254 (57%), Positives = 179/254 (70%), Gaps = 1/254 (0%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTN-RHLVGDSVG 748
SVK+TVERYKKA++ S+SG + E NAQ YQQES+KLR QI LQN N R +VGDS+
Sbjct: 61 SVKSTVERYKKANSDTSNSG-TVAEVNAQH-YQQESSKLRQQISSLQNANSRTIVGDSIN 118
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
+SL++LKQ+E+RLEKGI+KIRARK+ELL AE+ YM KRE ELQND+M LR+K+ E E+
Sbjct: 119 TMSLRDLKQVENRLEKGIAKIRARKNELLYAEVEYMQKREVELQNDNMYLRSKVVENERG 178
Query: 567 LQQVTVARSVXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQR 388
Q + + ++ NP+ D + FL ++ Q +
Sbjct: 179 QQPLNM-MGAASTSEYDHMVNNPY-----------------DSRNFLQVNIMQQPQHYAH 220
Query: 387 QMLATELNLGYQLA 346
Q+ T L LG Q A
Sbjct: 221 QLQPTTLQLGQQPA 234
>sptr|Q9AWJ2|Q9AWJ2 Putative MADS-box protein.
Length = 247
Score = 252 bits (643), Expect = 9e-66
Identities = 145/250 (58%), Positives = 177/250 (70%), Gaps = 1/250 (0%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTN-RHLVGDSVG 748
SVK+TVERYKKA++ S+SG + E NAQ YQQES+KLR QI LQN N R +VGDS+
Sbjct: 61 SVKSTVERYKKANSDTSNSG-TVAEVNAQH-YQQESSKLRQQISSLQNANSRTIVGDSIN 118
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
+SL++LKQ+E+RLEKGI+KIRARK+ELL AE+ YM KRE ELQND+M LR+K+ E E+
Sbjct: 119 TMSLRDLKQVENRLEKGIAKIRARKNELLYAEVEYMQKREVELQNDNMYLRSKVVENERG 178
Query: 567 LQQVTVARSVXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQR 388
Q + + ++ NP+ D + FL ++ Q +
Sbjct: 179 QQPLNM-MGAASTSEYDHMVNNPY-----------------DSRNFLQVNIMQQPQHYAH 220
Query: 387 QMLATELNLG 358
Q+ T L LG
Sbjct: 221 QLQPTTLQLG 230
>sptr|Q8LLQ9|Q8LLQ9 MADS-box protein 5.
Length = 223
Score = 249 bits (637), Expect = 4e-65
Identities = 130/185 (70%), Positives = 162/185 (87%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGR+YEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRVYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
++K+T++RYKKA + S++G +E NA Q+YQQESAKLR QIQMLQN+NRHL+GDS+ +
Sbjct: 61 NIKSTIDRYKKASS-DSTNGGFTMEINA-QYYQQESAKLRQQIQMLQNSNRHLMGDSLAS 118
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L++KELKQLE+RLE+GI++IR++K ELL AEI Y+ KRE EL+N+ + LRTKI E E +L
Sbjct: 119 LTVKELKQLENRLERGITRIRSKKHELLLAEIEYLQKREIELENESVYLRTKIAEVE-RL 177
Query: 564 QQVTV 550
QQ +
Sbjct: 178 QQANM 182
>sptr|Q8RU44|Q8RU44 Agamous-like protein 1 HvAG1.
Length = 234
Score = 249 bits (637), Expect = 4e-65
Identities = 145/252 (57%), Positives = 177/252 (70%), Gaps = 1/252 (0%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL+VFSSRGRLYEY+NN
Sbjct: 1 MGRGRIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTN-RHLVGDSVG 748
SVKAT+ERYKKA++ S+SG + E NA Q+YQQES+KLR QI LQN+N R LV DSV
Sbjct: 61 SVKATIERYKKANSDTSNSG-TVAEVNA-QYYQQESSKLRQQISSLQNSNSRSLVRDSVS 118
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
++L++LKQLE RLEKGI+KIRARK+EL+ AE+ YM KRE EL ND++ LR+K+ E E+
Sbjct: 119 TMTLRDLKQLEGRLEKGIAKIRARKNELMYAEVEYMQKREMELHNDNIYLRSKVSENERG 178
Query: 567 LQQVTVARSVXXXXXATNLELNPFLEMDTKCFFTGGPFATLDMKCFLPGSLQQMLEAQQR 388
Q + + S ++ A D + FL ++QQ Q
Sbjct: 179 QQPMNMMASGSTSSEYDHM------------------VAPYDSRNFLQVNMQQQQHYSQ- 219
Query: 387 QMLATELNLGYQ 352
Q+ T L LG Q
Sbjct: 220 QLQPTALQLGQQ 231
>sptr|Q8RVK1|Q8RVK1 AP3-like protein.
Length = 244
Score = 249 bits (636), Expect = 6e-65
Identities = 130/185 (70%), Positives = 154/185 (83%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL+ FSSRGRLYEYANN
Sbjct: 16 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVAFSSRGRLYEYANN 75
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVKAT+ERYKKA SS+ + E NA QFYQQE+ KLRNQI+ LQN NRH++G+S+G
Sbjct: 76 SVKATIERYKKAS--DSSNTGSVAEVNA-QFYQQEADKLRNQIRNLQNANRHMLGESIGG 132
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L +KELK LESRLEKGIS+IR++K+ELL AEI YM KRE +L N++ LR KI E E++
Sbjct: 133 LPMKELKSLESRLEKGISRIRSKKNELLFAEIEYMQKREIDLHNNNQLLRAKIAENERKQ 192
Query: 564 QQVTV 550
Q + +
Sbjct: 193 QSMNL 197
>sptr|Q8RVW5|Q8RVW5 MADS-box transcription factor.
Length = 239
Score = 248 bits (634), Expect = 9e-65
Identities = 129/182 (70%), Positives = 155/182 (85%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALI+FS+RGRLYEYANN
Sbjct: 12 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIIFSTRGRLYEYANN 71
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVK T+ERYKKA T S++G + E N+ Q+YQQE+ KLR QI LQN+NR+L+GD++
Sbjct: 72 SVKGTIERYKKASTDNSNTG-SISEANS-QYYQQEATKLRQQITNLQNSNRNLLGDALTT 129
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
+SL++LKQLE+RLEKGI+KIRA+K+ELL AEI YM KRE ELQ D+M LR KI + E+
Sbjct: 130 MSLRDLKQLETRLEKGINKIRAKKNELLHAEIDYMQKREMELQTDNMFLRNKISDNERAQ 189
Query: 564 QQ 559
QQ
Sbjct: 190 QQ 191
>sptr|Q9SNY4|Q9SNY4 Transcription factor MADS1.
Length = 234
Score = 247 bits (631), Expect = 2e-64
Identities = 126/185 (68%), Positives = 155/185 (83%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS+RGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
S+K+T+ER KKA SSS ++E N Q++YQQE++KLR QIQ+LQN NRHL+G+S+
Sbjct: 61 SIKSTIERDKKA-CADSSSSSAVIEVNTQRYYQQEASKLRQQIQILQNANRHLMGESLDP 119
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L++KELKQLE+RLE+GI+++R++K ELL AE+ YM KRE ELQ D+M LR KI E E+
Sbjct: 120 LNVKELKQLETRLERGITRVRSKKHELLFAELEYMQKREVELQTDNMYLRAKIGENERAH 179
Query: 564 QQVTV 550
Q V
Sbjct: 180 QASVV 184
>sptr|Q9SBK3|Q9SBK3 Agamous-like putative transcription factor.
Length = 225
Score = 247 bits (631), Expect = 2e-64
Identities = 128/181 (70%), Positives = 156/181 (86%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
S+K T+ERYKKA + SS+ + E N Q+YQQESAKLR QIQMLQN+NRHL+GDS+
Sbjct: 61 SIKTTIERYKKACS-DSSATSSVTELNT-QYYQQESAKLRQQIQMLQNSNRHLMGDSLSA 118
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L++KELKQLE+RLE+GI++IR++K E+L AEI Y+ KRE EL+N+++ +RTKI E E+
Sbjct: 119 LTVKELKQLENRLERGITRIRSKKHEMLLAEIEYLQKREIELENENVCIRTKIAEVERVQ 178
Query: 564 Q 562
Q
Sbjct: 179 Q 179
>sptr|Q8RU31|Q8RU31 Putative transcription factor agamous.
Length = 265
Score = 247 bits (630), Expect = 3e-64
Identities = 120/179 (67%), Positives = 155/179 (86%), Gaps = 1/179 (0%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYAN- 928
MGRG+IEIKRIEN TSRQVTFCKRRNGLLKKAYEL++LCDAE+ALIVFSSRGRLYE++N
Sbjct: 1 MGRGKIEIKRIENKTSRQVTFCKRRNGLLKKAYELAILCDAEIALIVFSSRGRLYEFSNV 60
Query: 927 NSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVG 748
NS ++T+ERYKKA + +S P+++ N+ Q++QQE+AK+R+QIQ LQN NRHL+G+S+G
Sbjct: 61 NSTRSTIERYKKA-SASTSGSAPVIDVNSHQYFQQEAAKMRHQIQTLQNANRHLIGESIG 119
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQ 571
N++ KELK LE+RLEKGIS+IR++K ELL +EI YM KRE +LQN++M LR K+ E E+
Sbjct: 120 NMTAKELKSLENRLEKGISRIRSKKHELLFSEIEYMQKREADLQNENMFLRAKVAEAER 178
>sptr|Q9XHM3|Q9XHM3 Agamous homolog (Fragment).
Length = 244
Score = 247 bits (630), Expect = 3e-64
Identities = 126/182 (69%), Positives = 150/182 (82%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAE+ALIVFSSRGRLYEYANN
Sbjct: 20 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEIALIVFSSRGRLYEYANN 79
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVK+T+ERYKKA +S P + QFYQQES+KLR QI+ +QN NRH++G+++ +
Sbjct: 80 SVKSTIERYKKA---SDTSNPGSVSETNAQFYQQESSKLRRQIRDIQNLNRHIMGEALSS 136
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L+ +ELK LE RLEKGIS+IR++K+ELL AEI YM KRE ELQN +M LR KI E E+
Sbjct: 137 LTFRELKNLEGRLEKGISRIRSKKNELLFAEIEYMQKREIELQNANMYLRAKIAENERNQ 196
Query: 564 QQ 559
QQ
Sbjct: 197 QQ 198
>sw|P29385|AGL5_ARATH Agamous-like MADS box protein AGL5.
Length = 246
Score = 245 bits (626), Expect = 8e-64
Identities = 125/193 (64%), Positives = 153/193 (79%)
Frame = -1
Query: 1137 AREEQAGIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS 958
A E A +GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL++FS
Sbjct: 5 ASNEVAESSKKIGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFS 64
Query: 957 SRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNT 778
+RGRLYEYANNSV+ T+ERYKKA + PP + Q+YQQE++KLR QI+ +QN
Sbjct: 65 TRGRLYEYANNSVRGTIERYKKA--CSDAVNPPTITEANTQYYQQEASKLRRQIRDIQNL 122
Query: 777 NRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTL 598
NRH++G+S+G+L+ KELK LESRLEKGIS++R++K E+L AEI YM KRE ELQND+M L
Sbjct: 123 NRHILGESLGSLNFKELKNLESRLEKGISRVRSKKHEMLVAEIEYMQKREIELQNDNMYL 182
Query: 597 RTKIEEGEQQLQQ 559
R+KI E QQ
Sbjct: 183 RSKITERTGLQQQ 195
>sptr|Q8H6F8|Q8H6F8 MADS box protein GHMADS-2.
Length = 223
Score = 245 bits (626), Expect = 8e-64
Identities = 132/213 (61%), Positives = 171/213 (80%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
++++T++RYKKA + +S+ + E NA Q+YQQESAKLR QIQMLQN+NRHL+GDS+ +
Sbjct: 61 NIRSTIDRYKKACS-DTSNTNTVTEINA-QYYQQESAKLRQQIQMLQNSNRHLMGDSLSS 118
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L++KELKQ+E+RLE+GI++IR++K E+L AEI ++ KRE EL+N+ + LRTKI E E +L
Sbjct: 119 LTVKELKQVENRLERGITRIRSKKHEMLLAEIEFLQKREIELENESVCLRTKIAEIE-RL 177
Query: 564 QQVTVARSVXXXXXATNLELNPFLEMDTKCFFT 466
QQ + T ELN + ++ FF+
Sbjct: 178 QQANMV---------TGPELNAIQALASRNFFS 201
>sptr|Q8GTP4|Q8GTP4 MADS box transcription factor.
Length = 254
Score = 244 bits (624), Expect = 1e-63
Identities = 130/187 (69%), Positives = 151/187 (80%)
Frame = -1
Query: 1119 GIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLY 940
G MGRGRIEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS RGRLY
Sbjct: 17 GAAEKMGRGRIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSGRGRLY 76
Query: 939 EYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVG 760
EY+NNSVKAT+ERYKKA T +SS + E NAQ YQQESAKL+ QI LQN+NR L+G
Sbjct: 77 EYSNNSVKATIERYKKA-TSDTSSAGTVAEINAQH-YQQESAKLKQQITTLQNSNRTLIG 134
Query: 759 DSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEE 580
D++ +S ++LKQLE RL+KG+ KIRARK+ELL AEI YM +RE ELQN++ LR K+ E
Sbjct: 135 DTMATMSHRDLKQLEGRLDKGLGKIRARKNELLCAEIEYMQRREMELQNNNFFLREKVAE 194
Query: 579 GEQQLQQ 559
E+ QQ
Sbjct: 195 TERGQQQ 201
>sptrnew|AAM64275|AAM64275 Shatterproof 1 (SHP1)/ agamous-like 1
(AGL1).
Length = 248
Score = 244 bits (624), Expect = 1e-63
Identities = 122/191 (63%), Positives = 153/191 (80%)
Frame = -1
Query: 1149 LQEPAREEQAGIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVAL 970
++E A +GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL
Sbjct: 1 MEEGGSSHDAESSKKLGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVAL 60
Query: 969 IVFSSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQM 790
++FS+RGRLYEYANNSV+ T+ERYKKA + PP + Q+YQQE++KLR QI+
Sbjct: 61 VIFSTRGRLYEYANNSVRGTIERYKKA--CSDAVNPPSVTEANTQYYQQEASKLRRQIRD 118
Query: 789 LQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQND 610
+QN+NRH+VG+S+G+L+ KELK LE RLEKGIS++R++K+ELL AEI YM KRE ELQ++
Sbjct: 119 IQNSNRHIVGESLGSLNFKELKNLEGRLEKGISRVRSKKNELLVAEIEYMQKREMELQHN 178
Query: 609 HMTLRTKIEEG 577
+M LR KI EG
Sbjct: 179 NMYLRAKIAEG 189
>sw|P29381|AGL1_ARATH Agamous-like MADS box protein AGL1 (Protein
Shatterproof 1).
Length = 248
Score = 244 bits (624), Expect = 1e-63
Identities = 122/191 (63%), Positives = 153/191 (80%)
Frame = -1
Query: 1149 LQEPAREEQAGIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVAL 970
++E A +GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL
Sbjct: 1 MEEGGSSHDAESSKKLGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVAL 60
Query: 969 IVFSSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQM 790
++FS+RGRLYEYANNSV+ T+ERYKKA + PP + Q+YQQE++KLR QI+
Sbjct: 61 VIFSTRGRLYEYANNSVRGTIERYKKA--CSDAVNPPSVTEANTQYYQQEASKLRRQIRD 118
Query: 789 LQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQND 610
+QN+NRH+VG+S+G+L+ KELK LE RLEKGIS++R++K+ELL AEI YM KRE ELQ++
Sbjct: 119 IQNSNRHIVGESLGSLNFKELKNLEGRLEKGISRVRSKKNELLVAEIEYMQKREMELQHN 178
Query: 609 HMTLRTKIEEG 577
+M LR KI EG
Sbjct: 179 NMYLRAKIAEG 189
>sptr|Q8RU43|Q8RU43 Agamous-like protein 2 HvAG2.
Length = 232
Score = 244 bits (624), Expect = 1e-63
Identities = 129/182 (70%), Positives = 152/182 (83%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRGRIEIKRIEN T+RQ+TFCKRRNGLLKKAYELSVLCDAEVAL+VFSSRGRLYEY+NN
Sbjct: 1 MGRGRIEIKRIENTTNRQLTFCKRRNGLLKKAYELSVLCDAEVALVVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SVKAT+ERYKKA T +SS + E NAQ YQQESAKLR QI LQN+NR L+GD++
Sbjct: 61 SVKATIERYKKA-TSDTSSAGTVAEINAQH-YQQESAKLRQQITTLQNSNRTLIGDTMAT 118
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
+S ++LKQLE RL+KG+ KIRARK+ELL+AEI YM +RE ELQN++ LR K+ E E+
Sbjct: 119 MSHRDLKQLEGRLDKGLGKIRARKNELLSAEIEYMQRREMELQNNNFYLREKVAETERGQ 178
Query: 564 QQ 559
QQ
Sbjct: 179 QQ 180
>sptr|Q93XH4|Q93XH4 MAD-box transcripion factor.
Length = 225
Score = 244 bits (622), Expect = 2e-63
Identities = 126/185 (68%), Positives = 156/185 (84%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SV+ T+ERYKK + S++G + E NA QFYQQE++KLR QI+ +QN NRH++G+++ +
Sbjct: 61 SVRTTIERYKKVCSDSSNTG-SVSEANA-QFYQQEASKLRRQIRDIQNLNRHILGEALSS 118
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L+ KELK LE+RLEKGIS+IR++K+ELL AEI YM KRE ELQN ++ LR +I E E+
Sbjct: 119 LNFKELKNLETRLEKGISRIRSKKNELLFAEIEYMQKREIELQNSNLFLRAQIAENERAQ 178
Query: 564 QQVTV 550
QQ+ +
Sbjct: 179 QQMNL 183
>sptr|Q9ZPK9|Q9ZPK9 Agamous homolog transcription factor.
Length = 228
Score = 243 bits (620), Expect = 4e-63
Identities = 130/189 (68%), Positives = 156/189 (82%), Gaps = 1/189 (0%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYAN- 928
MGRG+IEIKRIEN TSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS+RGRLYEY+N
Sbjct: 1 MGRGKIEIKRIENTTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYSNS 60
Query: 927 NSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVG 748
NSVK T+ERYKKA T +++G + E N+Q +YQQE+ KLR QI LQNTNR L+G+S+
Sbjct: 61 NSVKTTIERYKKACTDTTNTGT-VSEANSQ-YYQQEATKLRQQITNLQNTNRTLMGESLS 118
Query: 747 NLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQ 568
+SL+ELKQLE RLE+GI+KIR +K+ELL+AEI YM KRE E+ ND+M LR KI E E+
Sbjct: 119 TMSLRELKQLEGRLERGINKIRTKKNELLSAEIEYMQKREAEMHNDNMYLRNKIAENERA 178
Query: 567 LQQVTVARS 541
QQ+ + S
Sbjct: 179 QQQMNMLPS 187
>sptr|Q9MBE2|Q9MBE2 MADS-box protein.
Length = 249
Score = 242 bits (618), Expect = 7e-63
Identities = 126/194 (64%), Positives = 157/194 (80%)
Frame = -1
Query: 1140 PAREEQAGIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVF 961
PA + ++ +GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVF
Sbjct: 9 PADDPESSSQKKLGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVF 68
Query: 960 SSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQN 781
S+RGRLYEYANNSV+AT+ERYKKA SS+ + E N QFYQQE++KLR QI+ +QN
Sbjct: 69 STRGRLYEYANNSVRATIERYKKA--CDSSNTGSVTETNV-QFYQQEASKLRRQIREIQN 125
Query: 780 TNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMT 601
+NRH++G+++ L++KELK LE RLEKGIS+IR++K+E+L AEI YM KRE ELQN +
Sbjct: 126 SNRHILGEALSTLNVKELKNLEGRLEKGISRIRSKKNEMLFAEIEYMQKREIELQNHNNF 185
Query: 600 LRTKIEEGEQQLQQ 559
LR KI E ++ QQ
Sbjct: 186 LRAKIAENDRAQQQ 199
>sptr|Q84XW0|Q84XW0 Mads-box transcription factor.
Length = 227
Score = 242 bits (618), Expect = 7e-63
Identities = 135/217 (62%), Positives = 168/217 (77%), Gaps = 4/217 (1%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTN----RHLVGD 757
S+K T+ RYKKA + SS+ + E N Q+YQQESAKLR QIQMLQN+N RHL+GD
Sbjct: 61 SIKTTIGRYKKACS-DSSATSSVTELNT-QYYQQESAKLRQQIQMLQNSNSNLVRHLMGD 118
Query: 756 SVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEG 577
S+ L++KELKQLE+RLE+GI++IR++K E+L AEI Y+ KRE EL+N+++ +RTKI E
Sbjct: 119 SLSALTVKELKQLENRLERGITRIRSKKHEMLLAEIEYLQKREIELENENVCIRTKIAEV 178
Query: 576 EQQLQQVTVARSVXXXXXATNLELNPFLEMDTKCFFT 466
E +LQQ + + ELN + ++ FFT
Sbjct: 179 E-RLQQANMV---------SGQELNAIQALASRNFFT 205
>sptrnew|AAQ03090|AAQ03090 AGAMOUS-like protein.
Length = 242
Score = 241 bits (616), Expect = 1e-62
Identities = 124/199 (62%), Positives = 163/199 (81%)
Frame = -1
Query: 1149 LQEPAREEQAGIVASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVAL 970
++ P + ++ +GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVAL
Sbjct: 1 MEFPNQAPESSSQKKLGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVAL 60
Query: 969 IVFSSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQM 790
IVFS+RGRLYEYANNSV+AT++RYKKA+ ++SG + E N QFYQQE++KLR QI+
Sbjct: 61 IVFSNRGRLYEYANNSVRATIDRYKKAYADPTNSG-SVSEANT-QFYQQEASKLRRQIRE 118
Query: 789 LQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQND 610
+QN+NRH++G+++ +L+ KELK LE RLEKGIS+IR++K+E+L +EI +M KRETELQ+
Sbjct: 119 IQNSNRHILGEALSSLNAKELKNLEGRLEKGISRIRSKKNEMLFSEIEFMQKRETELQHH 178
Query: 609 HMTLRTKIEEGEQQLQQVT 553
+ LR KI E E++ QQ T
Sbjct: 179 NNFLRAKIAENEREEQQHT 197
>sptr|O64959|O64959 CUM10.
Length = 229
Score = 241 bits (616), Expect = 1e-62
Identities = 128/185 (69%), Positives = 156/185 (84%), Gaps = 4/185 (2%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
MGRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEY+NN
Sbjct: 1 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 60
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTN----RHLVGD 757
S+K T+ERYKKA + SS+ + E N Q+YQQESAKLR QIQMLQN+N RHL+GD
Sbjct: 61 SIKTTIERYKKACS-DSSATSSVTELNT-QYYQQESAKLRQQIQMLQNSNSNLVRHLMGD 118
Query: 756 SVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEG 577
S+ L++KELKQLE+RLE+GI++IR++K E+L AEI Y+ KRE EL+N+++ +RTKI E
Sbjct: 119 SLSALTVKELKQLENRLERGITRIRSKKHEMLLAEIEYLQKREIELENENVCIRTKIAEV 178
Query: 576 EQQLQ 562
E+ Q
Sbjct: 179 ERVQQ 183
>sptr|Q9SBK2|Q9SBK2 Agamous-like putative transcription factor.
Length = 254
Score = 241 bits (614), Expect = 2e-62
Identities = 136/236 (57%), Positives = 168/236 (71%), Gaps = 15/236 (6%)
Frame = -1
Query: 1131 EEQAGIV--------ASMGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEV 976
+E++G+V + GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEV
Sbjct: 8 DEESGVVGLRRSSSSSRTGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEV 67
Query: 975 ALIVFSSRGRLYEYANNSVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQI 796
ALIVFSSRGRLYEYANNSV+AT+ RYKKA++ S+ + E N QFYQQESAKLR QI
Sbjct: 68 ALIVFSSRGRLYEYANNSVRATISRYKKAYS-DPSTAMTVSEANT-QFYQQESAKLRAQI 125
Query: 795 QMLQNTNRHLVGDSVGNLSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQ 616
LQN NRHL+G+S+ +LS+K+LK LE +LEKGIS+IR+RK+ELL +EI YM KRE EL
Sbjct: 126 GNLQNLNRHLLGESISSLSVKDLKSLEVKLEKGISRIRSRKNELLFSEIEYMQKREIELH 185
Query: 615 NDHMTLRTKIEEGEQQLQQVTVA-------RSVXXXXXATNLELNPFLEMDTKCFF 469
++ +R KI E E+ Q + R TNLE N + D+ +F
Sbjct: 186 TNNQLIRAKIAETERSXQNTNASNNNGIATRRGEEGSMGTNLEDNNHHQYDSTNYF 241
>sptr|Q8VWZ3|Q8VWZ3 C-type MADS box protein.
Length = 242
Score = 240 bits (613), Expect = 3e-62
Identities = 123/182 (67%), Positives = 153/182 (84%)
Frame = -1
Query: 1104 MGRGRIEIKRIENNTSRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANN 925
+GRG+IEIKRIEN T+RQVTFCKRRNGLLKKAYELSVLCDAEVALIVFS+RGRLYEYANN
Sbjct: 16 LGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYANN 75
Query: 924 SVKATVERYKKAHTVGSSSGPPLLEHNAQQFYQQESAKLRNQIQMLQNTNRHLVGDSVGN 745
SV+AT++RYKKA S+ G + E N QFYQQE++KLR QI+ +QN+NRH++G+S+
Sbjct: 76 SVRATIDRYKKA-CADSTDGGSVSEANT-QFYQQEASKLRRQIREIQNSNRHILGESLST 133
Query: 744 LSLKELKQLESRLEKGISKIRARKSELLAAEISYMAKRETELQNDHMTLRTKIEEGEQQL 565
L +KELK LE RLEKGIS+IR++K+E+L +EI +M KRETELQ+ + LR KI E E++
Sbjct: 134 LKVKELKNLEGRLEKGISRIRSKKNEILFSEIEFMQKRETELQHHNNFLRAKIAESEREQ 193
Query: 564 QQ 559
QQ
Sbjct: 194 QQ 195
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 12, 2003 8:28 PM
Number of letters in database: 397,940,306
Number of sequences in database: 1,242,019
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 813,374,984
Number of Sequences: 1242019
Number of extensions: 16757207
Number of successful extensions: 47801
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 44028
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 47142
length of database: 397,940,306
effective HSP length: 125
effective length of database: 242,687,931
effective search space used: 68195308611
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

