BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
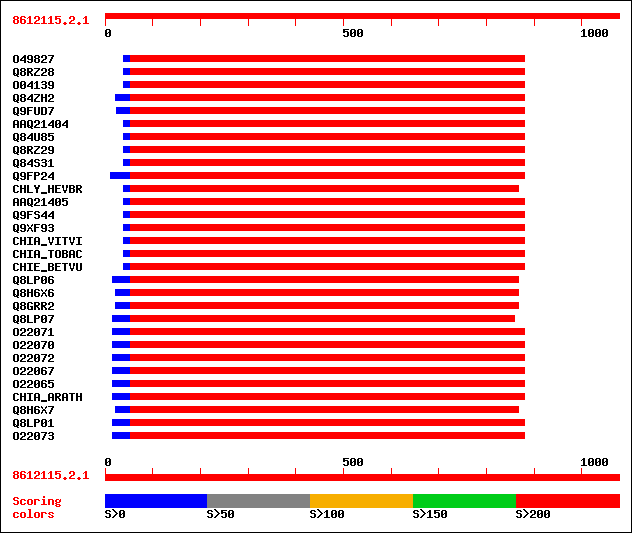
Query= 8612115.2.1
(1078 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O49827|O49827 Chitinase (EC 3.2.1.14). 352 8e-99
sptr|Q8RZ28|Q8RZ28 Chitinase. 352 8e-99
sptr|O04139|O04139 Basic class III chitinase OsChib3b precursor. 351 2e-98
sptr|Q84ZH2|Q84ZH2 Putative class III acidic chitinase. 295 2e-80
sptr|Q9FUD7|Q9FUD7 Class III acidic chitinase. 290 2e-80
sptrnew|AAQ21404|AAQ21404 Class III chitinase. 290 1e-79
sptr|Q84U85|Q84U85 Acidic class III chitinase (EC 3.2.1.14). 285 4e-78
sptr|Q8RZ29|Q8RZ29 Putative chitinase. 288 3e-77
sptr|Q84S31|Q84S31 Chitinase III. 280 3e-77
sptr|Q9FP24|Q9FP24 Putative class III chitinase. 277 3e-76
sw|P23472|CHLY_HEVBR Hevamine A precursor [Includes: Chitinase (... 275 1e-75
sptrnew|AAQ21405|AAQ21405 Putative class III chitinase. 281 2e-75
sptr|Q9FS44|Q9FS44 Chitinase precursor (EC 3.2.1.17). 273 5e-75
sptr|Q9XF93|Q9XF93 Chitinase (EC 3.2.1.14). 273 7e-75
sw|P51614|CHIA_VITVI Acidic endochitinase precursor (EC 3.2.1.14). 270 4e-74
sw|P29060|CHIA_TOBAC Acidic endochitinase precursor (EC 3.2.1.14). 271 5e-74
sw|P36910|CHIE_BETVU Acidic endochitinase SE2 precursor (EC 3.2.... 268 2e-72
sptr|Q8LP06|Q8LP06 Acidic endochitinase. 256 7e-72
sptr|Q8H6X6|Q8H6X6 Class III chitinase (EC 3.2.1.14). 260 2e-71
sptr|Q8GRR2|Q8GRR2 Class III chitinase (EC 3.2.1.14). 260 3e-71
sptr|Q8LP07|Q8LP07 Acidic endochitinase (Fragment). 254 3e-71
sptr|O22071|O22071 Acidic endochitinase. 254 4e-71
sptr|O22070|O22070 Acidic endochitinase. 253 5e-71
sptr|O22072|O22072 Acidic endochitinase. 253 7e-71
sptr|O22067|O22067 Acidic endochitinase. 253 9e-71
sptr|O22065|O22065 Acidic endochitinase. 253 9e-71
sw|P19172|CHIA_ARATH Acidic endochitinase precursor (EC 3.2.1.14). 253 9e-71
sptr|Q8H6X7|Q8H6X7 Class III chitinase (EC 3.2.1.14). 258 9e-71
sptr|Q8LP01|Q8LP01 Acidic endochitinase. 252 1e-70
sptr|O22073|O22073 Acidic endochitinase. 252 1e-70
>sptr|O49827|O49827 Chitinase (EC 3.2.1.14).
Length = 305
Score = 352 bits (904), Expect(2) = 8e-99
Identities = 177/276 (64%), Positives = 194/276 (70%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTCATGNYRFV VAFLP FGKGQTP LNLAGHCDPAS GCTGVGAD+K+C
Sbjct: 37 GQNGNEGTLAQTCATGNYRFVIVAFLPVFGKGQTPVLNLAGHCDPASNGCTGVGADIKSC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q +G+KV+ SIGGGVG+YGLSSR DA+ VAAYLW+NYL
Sbjct: 97 QSLGIKVMFSIGGGVGNYGLSSRDDAKQVAAYLWNNYLGGTSPSRPLGDAVMDGIDFDIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
G+ P +AAPQCPFPDASLG AL+TGLFDY
Sbjct: 157 SGGGMYWDDLARYLKA-------YSRQGSSKKPVYLTAAPQCPFPDASLGVALSTGLFDY 209
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNPPCQYS+S GVG+LA AW QWTSI AGRVFLGLPAA +AAGSGFV SDLV+
Sbjct: 210 VWVQFYNNPPCQYSSSNGVGNLASAWKQWTSIPAGRVFLGLPAAAEAAGSGFVETSDLVS 269
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+VLPVVK S KYGGIMLWSRYYDGLTGYSD VKS V
Sbjct: 270 KVLPVVKKSPKYGGIMLWSRYYDGLTGYSDKVKSSV 305
Score = 67.0 bits (162), Expect = 5e-10
Identities = 28/32 (87%), Positives = 32/32 (100%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAV+DG+DFDIESGGGMYWDDLAR+LK+YS
Sbjct: 143 LGDAVMDGIDFDIESGGGMYWDDLARYLKAYS 174
Score = 31.6 bits (70), Expect(2) = 8e-99
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 32 IAIYWGQNGNEG 43
>sptr|Q8RZ28|Q8RZ28 Chitinase.
Length = 296
Score = 352 bits (904), Expect(2) = 8e-99
Identities = 177/276 (64%), Positives = 194/276 (70%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTCATGNYRFV VAFLP FGKGQTP LNLAGHCDPAS GCTGVGAD+K+C
Sbjct: 28 GQNGNEGTLAQTCATGNYRFVIVAFLPVFGKGQTPVLNLAGHCDPASNGCTGVGADIKSC 87
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q +G+KV+ SIGGGVG+YGLSSR DA+ VAAYLW+NYL
Sbjct: 88 QSLGIKVMFSIGGGVGNYGLSSRDDAKQVAAYLWNNYLGGTSPSRPLGDAVMDGIDFDIE 147
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
G+ P +AAPQCPFPDASLG AL+TGLFDY
Sbjct: 148 SGGGMYWDDLARYLKA-------YSRQGSSKKPVYLTAAPQCPFPDASLGVALSTGLFDY 200
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNPPCQYS+S GVG+LA AW QWTSI AGRVFLGLPAA +AAGSGFV SDLV+
Sbjct: 201 VWVQFYNNPPCQYSSSNGVGNLASAWKQWTSIPAGRVFLGLPAAAEAAGSGFVETSDLVS 260
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+VLPVVK S KYGGIMLWSRYYDGLTGYSD VKS V
Sbjct: 261 KVLPVVKKSPKYGGIMLWSRYYDGLTGYSDKVKSSV 296
Score = 67.0 bits (162), Expect = 5e-10
Identities = 28/32 (87%), Positives = 32/32 (100%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAV+DG+DFDIESGGGMYWDDLAR+LK+YS
Sbjct: 134 LGDAVMDGIDFDIESGGGMYWDDLARYLKAYS 165
Score = 31.6 bits (70), Expect(2) = 8e-99
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 23 IAIYWGQNGNEG 34
>sptr|O04139|O04139 Basic class III chitinase OsChib3b precursor.
Length = 305
Score = 351 bits (900), Expect(2) = 2e-98
Identities = 176/276 (63%), Positives = 193/276 (69%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTCATGNYRFV VAFLP FGKGQTP LNLAGHCDPAS GCTGVGAD+K+C
Sbjct: 37 GQNGNEGTLAQTCATGNYRFVIVAFLPVFGKGQTPVLNLAGHCDPASNGCTGVGADIKSC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q +G+KV+ SIGGGVG+YGLSSR DA+ VAAYLW+NYL
Sbjct: 97 QSLGIKVMFSIGGGVGNYGLSSRDDAKQVAAYLWNNYLGGTSPSRPLGDAVMDGIDFDIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
G+ P +AAPQCPFPDASLG AL+TGLFDY
Sbjct: 157 SGGGMYWDDLARYLKA-------YSRQGSSKKPVYLTAAPQCPFPDASLGVALSTGLFDY 209
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNPPCQYS+S GVG+LA AW QWTSI AGRVFLGLP A +AAGSGFV SDLV+
Sbjct: 210 VWVQFYNNPPCQYSSSNGVGNLASAWKQWTSIPAGRVFLGLPVAAEAAGSGFVETSDLVS 269
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+VLPVVK S KYGGIMLWSRYYDGLTGYSD VKS V
Sbjct: 270 KVLPVVKKSPKYGGIMLWSRYYDGLTGYSDKVKSSV 305
Score = 67.0 bits (162), Expect = 5e-10
Identities = 28/32 (87%), Positives = 32/32 (100%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAV+DG+DFDIESGGGMYWDDLAR+LK+YS
Sbjct: 143 LGDAVMDGIDFDIESGGGMYWDDLARYLKAYS 174
Score = 31.6 bits (70), Expect(2) = 2e-98
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 32 IAIYWGQNGNEG 43
>sptr|Q84ZH2|Q84ZH2 Putative class III acidic chitinase.
Length = 297
Score = 295 bits (755), Expect(2) = 2e-80
Identities = 147/276 (53%), Positives = 173/276 (62%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATGNY+FVN+AFL FG GQ P NLAGHCDP +GGC +D+K+C
Sbjct: 33 GQNNGEGTLADTCATGNYKFVNIAFLAAFGNGQPPVFNLAGHCDPTNGGCASQSSDIKSC 92
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q GVK++LSIGGG GSY LSS DA++VA YLW+N+L
Sbjct: 93 QSRGVKIMLSIGGGAGSYYLSSSEDAKNVATYLWNNFLGGQSSSRPLGDAVLDGIDFDIE 152
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G G +AAPQCPFPDA +G AL TGLFDY
Sbjct: 153 GGTNQHWDDLARY----------LKGYSNSGRRVYLTAAPQCPFPDACIGDALNTGLFDY 202
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNPPCQYS S +LA AW QW S+ A ++FLGLPA+PQAAGSGF+PA DL +
Sbjct: 203 VWVQFYNNPPCQYS-SGSTSNLADAWKQWLSVPAKQIFLGLPASPQAAGSGFIPADDLKS 261
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
QVLPV+K+S KYGGIMLWS+YYD YS +VKS V
Sbjct: 262 QVLPVIKSSGKYGGIMLWSKYYDDQDDYSSSVKSDV 297
Score = 56.6 bits (135), Expect = 6e-07
Identities = 24/32 (75%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DFDIE G +WDDLAR+LK YS
Sbjct: 139 LGDAVLDGIDFDIEGGTNQHWDDLARYLKGYS 170
Score = 27.7 bits (60), Expect(2) = 2e-80
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +3
Query: 21 SASRRXIAIYWGQNGNEG 74
S+ IAIYWGQN EG
Sbjct: 22 SSQAGSIAIYWGQNNGEG 39
>sptr|Q9FUD7|Q9FUD7 Class III acidic chitinase.
Length = 299
Score = 290 bits (743), Expect(2) = 2e-80
Identities = 147/277 (53%), Positives = 177/277 (63%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + +TCA+GNY+FVNVAFL TFG GQTPA+NLAGHCDP + CT + ++K+C
Sbjct: 34 GQNGNEGTLAETCASGNYQFVNVAFLTTFGNGQTPAINLAGHCDPTTEECTKLSPEIKSC 93
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LSIGG GSY L+S DAR VA YLW+N+L
Sbjct: 94 QAKGIKVILSIGGASGSYSLTSADDARQVATYLWNNFLGGQSSSRPLGAAVLDGIDFDIE 153
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G RG +AAPQCPFPDA +G AL TGLFD
Sbjct: 154 GGTDQHWDDLARY----------LSGYSKRGKKVYLTAAPQCPFPDAYVGNALKTGLFDN 203
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNPPCQY AS V +L AW QWTS I A ++FLGLPAAPQAAGSGF+PA+DL
Sbjct: 204 VWVQFYNNPPCQY-ASGDVTNLEDAWKQWTSAIPADKIFLGLPAAPQAAGSGFIPATDLS 262
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+QVLP +K+S KYGG+MLWS+YYD GYS ++K+ V
Sbjct: 263 SQVLPAIKSSAKYGGVMLWSKYYDDPDGYSSSIKNDV 299
Score = 51.2 bits (121), Expect = 3e-05
Identities = 22/32 (68%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG AVLDG+DFDIE G +WDDLAR+L YS
Sbjct: 140 LGAAVLDGIDFDIEGGTDQHWDDLARYLSGYS 171
Score = 32.0 bits (71), Expect(2) = 2e-80
Identities = 13/17 (76%), Positives = 14/17 (82%)
Frame = +3
Query: 24 ASRRXIAIYWGQNGNEG 74
A+ IAIYWGQNGNEG
Sbjct: 24 ANAGGIAIYWGQNGNEG 40
>sptrnew|AAQ21404|AAQ21404 Class III chitinase.
Length = 298
Score = 290 bits (741), Expect(2) = 1e-79
Identities = 143/277 (51%), Positives = 179/277 (64%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + + CATGNY +V +AFLPTFG GQTP +NLAGHCDP S CTG+ +D+K+C
Sbjct: 35 GQNGNEGTLAEACATGNYEYVIIAFLPTFGDGQTPMINLAGHCDPYSNECTGLSSDIKSC 94
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KVLLS+GGG GSY ++S DA+SVA YLW+N+L
Sbjct: 95 QAKGIKVLLSLGGGAGSYSIASTQDAKSVATYLWNNFLGGQSSSRPLGPAVLDGIDFDIE 154
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
G + + L G G +AAPQCPFPDA +G AL TGLFDY
Sbjct: 155 GGSNQHW---------GDLAKYL---KGYNGKKVYITAAPQCPFPDAWIGNALTTGLFDY 202
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNPPCQY+ + +L AW QWTS I A ++FLGLPA+P+AAGSGF+PA+DL
Sbjct: 203 VWVQFYNNPPCQYNPGE-ISNLEDAWKQWTSGIPANKIFLGLPASPEAAGSGFIPATDLT 261
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+ VLP +K S KYGG+MLWSRY D +GYS ++KS+V
Sbjct: 262 STVLPAIKGSAKYGGVMLWSRYDDVQSGYSSSIKSHV 298
Score = 49.7 bits (117), Expect = 8e-05
Identities = 20/32 (62%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG AVLDG+DFDIE G +W DLA++LK Y+
Sbjct: 141 LGPAVLDGIDFDIEGGSNQHWGDLAKYLKGYN 172
Score = 30.4 bits (67), Expect(2) = 1e-79
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
I+IYWGQNGNEG
Sbjct: 30 ISIYWGQNGNEG 41
>sptr|Q84U85|Q84U85 Acidic class III chitinase (EC 3.2.1.14).
Length = 294
Score = 285 bits (728), Expect(2) = 4e-78
Identities = 143/277 (51%), Positives = 173/277 (62%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPA-SGGCTGVGADVKA 229
G G + +TCATGNY +VN+AFLPTFG GQTP +NLAGHCDP + GCT + + +K+
Sbjct: 32 GQNGNEGTLAETCATGNYHYVNIAFLPTFGNGQTPMINLAGHCDPTITNGCTHLSSQIKS 91
Query: 230 CQRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXX 409
CQ G+KV+LSIGGG GSY LSS DA+ VA YL++N+L
Sbjct: 92 CQAKGIKVMLSIGGGAGSYYLSSSQDAKQVATYLFNNFLSGKSSPRPLGDAILDGIDLDI 151
Query: 410 XXXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFD 589
L G RG +AAPQCPFPD +G AL TGLFD
Sbjct: 152 EGGTDLYWDDLARY----------LSNYGKRGRKVYLTAAPQCPFPDYYIGNALQTGLFD 201
Query: 590 YVWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
YVWVQFYNNPPCQYS+ G+ S KAW W SI AG +FLGLPA+ QAAG+GFVPA DL
Sbjct: 202 YVWVQFYNNPPCQYSS--GMDSFEKAWKDWNSIPAGEIFLGLPASAQAAGTGFVPAGDLT 259
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+QVLP +K S KYGG+MLW +Y+D TGYS ++K V
Sbjct: 260 SQVLPAIKGSAKYGGVMLWDKYHD--TGYSSSIKKDV 294
Score = 53.1 bits (126), Expect = 7e-06
Identities = 21/31 (67%), Positives = 26/31 (83%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSY 482
LGDA+LDG+D DIE G +YWDDLAR+L +Y
Sbjct: 139 LGDAILDGIDLDIEGGTDLYWDDLARYLSNY 169
Score = 30.4 bits (67), Expect(2) = 4e-78
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
I+IYWGQNGNEG
Sbjct: 27 ISIYWGQNGNEG 38
>sptr|Q8RZ29|Q8RZ29 Putative chitinase.
Length = 297
Score = 288 bits (737), Expect(2) = 3e-77
Identities = 147/277 (53%), Positives = 176/277 (63%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G + +TCA+GNY FV +AFLP FGKGQTP ++LA HCDPASGGCTG D++AC
Sbjct: 33 GQNDGEASLAETCASGNYEFVIIAFLPKFGKGQTPRVDLASHCDPASGGCTGQSKDIRAC 92
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
QR GVKVLLSIGGG GSYGLSS DAR VA YLW+N+L
Sbjct: 93 QRRGVKVLLSIGGGDGSYGLSSPGDARQVAMYLWNNFLGGSSSSRPLGDAVLDGIDFDIE 152
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGP-CTCSAAPQCPFPDASLGTALATGLFD 589
L G GG SAAPQCPFPD G A++TGLFD
Sbjct: 153 LGGAKFWDDLARD----------LKSLGRSGGRRVVLSAAPQCPFPDEWDGGAISTGLFD 202
Query: 590 YVWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNP CQ+SA G G+ AW +W S+ AGR+FLGLPA+ AAG+GFVPA +L
Sbjct: 203 AVWVQFYNNPECQFSA--GRGAFMDAWRKWESVPAGRLFLGLPASKDAAGTGFVPAGELN 260
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
++VLP+++ S KYGG+MLWS+YYD TGYS A+KS+V
Sbjct: 261 SRVLPLIRGSPKYGGVMLWSKYYDDQTGYSSAIKSHV 297
Score = 53.9 bits (128), Expect = 4e-06
Identities = 24/30 (80%), Positives = 26/30 (86%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKS 479
LGDAVLDG+DFDIE GG +WDDLAR LKS
Sbjct: 139 LGDAVLDGIDFDIELGGAKFWDDLARDLKS 168
Score = 23.9 bits (50), Expect(2) = 3e-77
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = +3
Query: 39 IAIYWGQNGNE 71
IA+YWGQN E
Sbjct: 28 IAVYWGQNDGE 38
>sptr|Q84S31|Q84S31 Chitinase III.
Length = 297
Score = 280 bits (717), Expect(2) = 3e-77
Identities = 139/276 (50%), Positives = 170/276 (61%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTC TG Y +VN+AFL FG GQTP +NLAGHC+PAS GCT V ++ C
Sbjct: 33 GQNGNEGTLTQTCNTGKYSYVNIAFLNKFGNGQTPEINLAGHCNPASNGCTSVSTGIRNC 92
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LSIGGGVGSY LSS DA++VA YLW+N+L
Sbjct: 93 QNRGIKVMLSIGGGVGSYSLSSSNDAQNVANYLWNNFLGGQSSSRPLGDAVLDGIDFDIE 152
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G RG +AAPQCPFPD LGTAL TGLFDY
Sbjct: 153 LGSTLHWDDLARA----------LSGFSKRGRKVYLTAAPQCPFPDKFLGTALNTGLFDY 202
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNP CQYS S +L +W +WTS ++F+GLPA+ AAGSGF+PA+ L +
Sbjct: 203 VWVQFYNNPQCQYS-SGNTNNLLNSWNRWTSSINSQIFMGLPASSAAAGSGFIPANVLTS 261
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
Q+LPV+K S KYGG+MLWS+YYD +GYS ++KS V
Sbjct: 262 QILPVIKRSPKYGGVMLWSKYYDDQSGYSSSIKSSV 297
Score = 51.2 bits (121), Expect = 3e-05
Identities = 22/32 (68%), Positives = 26/32 (81%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DFDIE G ++WDDLAR L +S
Sbjct: 139 LGDAVLDGIDFDIELGSTLHWDDLARALSGFS 170
Score = 31.6 bits (70), Expect(2) = 3e-77
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 28 IAIYWGQNGNEG 39
>sptr|Q9FP24|Q9FP24 Putative class III chitinase.
Length = 317
Score = 277 bits (708), Expect(2) = 3e-76
Identities = 145/281 (51%), Positives = 171/281 (60%), Gaps = 5/281 (1%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATGNY FVN+AFL +FG GQ P LNLAGHCD SG C + AD+ C
Sbjct: 40 GQNGNEGTLADTCATGNYAFVNLAFLCSFGSGQAPQLNLAGHCDAYSGACANLTADIARC 99
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q MGVKVLLSIGGG G Y L+S+ D +A YLW+++L
Sbjct: 100 QSMGVKVLLSIGGGAGGYSLASKQDVSHLARYLWESFLGGRPSAPGGRRPLGDAVLDGVD 159
Query: 413 XXXXXXXXXXXXXXXPGPVPQVL--LPGAGARGGPCTCSAAPQCPFPDASLGTALATGLF 586
G + L G GA G SAAPQCPFPD +G AL TGLF
Sbjct: 160 FDIEGGGGDPRYY---GDLAAYLKAYSGKGAAGKEVLLSAAPQCPFPDQWVGKALDTGLF 216
Query: 587 DYVWVQFYNNPPCQYSASAGVGS--LAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPA 757
DYVWVQFYNNPPCQY+A +G G+ L AW QWTS + A +FLGLPA+P AAGSGF+P
Sbjct: 217 DYVWVQFYNNPPCQYAAGSGGGAANLLDAWRQWTSGVEARYIFLGLPASPGAAGSGFIPV 276
Query: 758 SDLVAQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
L +QVLP +K S+KYGG+MLWSRYYD GYS A+K+ V
Sbjct: 277 GSLESQVLPALKASSKYGGVMLWSRYYDDQDGYSSAIKNAV 317
Score = 50.8 bits (120), Expect = 3e-05
Identities = 25/34 (73%), Positives = 28/34 (82%), Gaps = 2/34 (5%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGG--MYWDDLARFLKSYS 485
LGDAVLDGVDFDIE GGG Y+ DLA +LK+YS
Sbjct: 150 LGDAVLDGVDFDIEGGGGDPRYYGDLAAYLKAYS 183
Score = 32.0 bits (71), Expect(2) = 3e-76
Identities = 14/21 (66%), Positives = 14/21 (66%)
Frame = +3
Query: 12 SPHSASRRXIAIYWGQNGNEG 74
S A IAIYWGQNGNEG
Sbjct: 26 SSRGAHGGRIAIYWGQNGNEG 46
>sw|P23472|CHLY_HEVBR Hevamine A precursor [Includes: Chitinase (EC
3.2.1.14); Lysozyme (EC 3.2.1.17)].
Length = 311
Score = 275 bits (704), Expect(2) = 1e-75
Identities = 133/273 (48%), Positives = 174/273 (63%), Gaps = 1/273 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTC+T Y +VN+AFL FG GQTP +NLAGHC+PA+GGCT V +++C
Sbjct: 34 GQNGNEGTLTQTCSTRKYSYVNIAFLNKFGNGQTPQINLAGHCNPAAGGCTIVSNGIRSC 93
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+GSY L+S+ADA++VA YLW+N+L
Sbjct: 94 QIQGIKVMLSLGGGIGSYTLASQADAKNVADYLWNNFLGGKSSSRPLGDAVLDGIDFDIE 153
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L +G +AAPQCPFPD LGTAL TGLFDY
Sbjct: 154 HGSTLYWDDLARY----------LSAYSKQGKKVYLTAAPQCPFPDRYLGTALNTGLFDY 203
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNPPCQYS S + ++ +W +WT SI AG++FLGLPAAP+AAGSG+VP L+
Sbjct: 204 VWVQFYNNPPCQYS-SGNINNIINSWNRWTTSINAGKIFLGLPAAPEAAGSGYVPPDVLI 262
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAV 868
+++LP +K S KYGG+MLWS++YD GYS ++
Sbjct: 263 SRILPEIKKSPKYGGVMLWSKFYDDKNGYSSSI 295
Score = 57.4 bits (137), Expect = 4e-07
Identities = 24/32 (75%), Positives = 28/32 (87%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DFDIE G +YWDDLAR+L +YS
Sbjct: 140 LGDAVLDGIDFDIEHGSTLYWDDLARYLSAYS 171
Score = 31.6 bits (70), Expect(2) = 1e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 29 IAIYWGQNGNEG 40
>sptrnew|AAQ21405|AAQ21405 Putative class III chitinase.
Length = 297
Score = 281 bits (718), Expect(2) = 2e-75
Identities = 142/277 (51%), Positives = 174/277 (62%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + + CAT NY +V +AFLPTFG GQTP +NLAGHCDP S GCTG+ +D+K+C
Sbjct: 35 GQNGNEGTLAEACATENYEYVIIAFLPTFGDGQTPMINLAGHCDPYSNGCTGLSSDIKSC 94
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KVLLSIGGG GSY ++S DA SVA YLW+N+L
Sbjct: 95 QAKGIKVLLSIGGGAGSYSIASTKDANSVATYLWNNFLGGKSSSRPLGPAVLDGIDFDIE 154
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G + +AAPQCPFPDA +G AL TGLFDY
Sbjct: 155 GGSSQHWGDLARY----------LKGFNKK---VYITAAPQCPFPDAWIGNALTTGLFDY 201
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNPPCQY+ A + + AW QWTS I A ++FLGLPA+P AAGSGF+ A DL
Sbjct: 202 VWVQFYNNPPCQYNPDAFM-NFEDAWKQWTSGIPANKIFLGLPASPTAAGSGFISADDLT 260
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+ VLPV+K S+KYGG+MLWSRY D +GYS ++KS+V
Sbjct: 261 STVLPVIKGSSKYGGVMLWSRYDDVQSGYSSSIKSHV 297
Score = 49.3 bits (116), Expect = 1e-04
Identities = 20/32 (62%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG AVLDG+DFDIE G +W DLAR+LK ++
Sbjct: 141 LGPAVLDGIDFDIEGGSSQHWGDLARYLKGFN 172
Score = 25.0 bits (53), Expect(2) = 2e-75
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IA+Y GQNGNEG
Sbjct: 30 IAMYRGQNGNEG 41
>sptr|Q9FS44|Q9FS44 Chitinase precursor (EC 3.2.1.17).
Length = 297
Score = 273 bits (698), Expect(2) = 5e-75
Identities = 137/276 (49%), Positives = 164/276 (59%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTC TG Y +VN+AFL FG GQTP +NLAGHCD AS GCT V D+ C
Sbjct: 33 GQNGNEGTLTQTCNTGKYSYVNIAFLNKFGNGQTPEINLAGHCDSASNGCTSVSTDISNC 92
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q GVKV+LSIGG +GSY LSS DA++VA YLW+N+L
Sbjct: 93 QSQGVKVMLSIGGAIGSYSLSSSDDAQNVANYLWNNFLGGRSSSRPLGDAVLDGIDFVIL 152
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G RG +AAPQCPFPD +GTAL TG FD
Sbjct: 153 LGSTQYWDDLARA----------LSGFSQRGRKVYLTAAPQCPFPDKFMGTALNTGRFDN 202
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VW QFYNNPPCQY+ S +L +W +WTS R+FLGLPAA AAGSGF+P + L +
Sbjct: 203 VWAQFYNNPPCQYT-SGNTTNLLNSWNRWTSSINSRIFLGLPAASAAAGSGFIPPNVLTS 261
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
Q+LPV+K S KYGG+MLWS+YYD +GYS +KS V
Sbjct: 262 QILPVIKTSPKYGGVMLWSKYYDDQSGYSSTIKSSV 297
Score = 45.8 bits (107), Expect = 0.001
Identities = 21/32 (65%), Positives = 23/32 (71%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF I G YWDDLAR L +S
Sbjct: 139 LGDAVLDGIDFVILLGSTQYWDDLARALSGFS 170
Score = 31.6 bits (70), Expect(2) = 5e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 28 IAIYWGQNGNEG 39
>sptr|Q9XF93|Q9XF93 Chitinase (EC 3.2.1.14).
Length = 299
Score = 273 bits (697), Expect(2) = 7e-75
Identities = 140/278 (50%), Positives = 175/278 (62%), Gaps = 2/278 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTCA+GNY+FVN+AF +FG G+TP LNLAGHCDP+S CT + +K+C
Sbjct: 32 GQNGNEGTLAQTCASGNYQFVNIAFHSSFGNGRTPTLNLAGHCDPSSNTCTKFSSQIKSC 91
Query: 233 QRMGVKVLLSIGGGVGS-YGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXX 409
Q G+KV+LS+GGG G Y L+S DAR V AYLW+N+L
Sbjct: 92 QAEGIKVILSVGGGWGGQYSLASSEDARQVGAYLWNNFLGGHSTSRPLGDAVLDGVDFDI 151
Query: 410 XXXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFD 589
+ + L + G +AAPQCPFPDA++G AL TGLFD
Sbjct: 152 EGGNDQYWDD---------LARYLSAHSKKGGKKVYLTAAPQCPFPDANIGNALKTGLFD 202
Query: 590 YVWVQFYNNPPCQYSASAGVGSLAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPASDL 766
VWVQFYNNPPCQY+ S V +L AW QWTS I A +VFLGLPAAP+AAGSGF+PA+ L
Sbjct: 203 NVWVQFYNNPPCQYT-SGNVTNLEDAWKQWTSAIPAQQVFLGLPAAPEAAGSGFIPAAAL 261
Query: 767 VAQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
VLP +K S KYGG+MLWS+YYD L GYS ++K++V
Sbjct: 262 TTTVLPGIKTSDKYGGVMLWSKYYDDLYGYSSSIKNHV 299
Score = 55.8 bits (133), Expect = 1e-06
Identities = 24/32 (75%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDGVDFDIE G YWDDLAR+L ++S
Sbjct: 139 LGDAVLDGVDFDIEGGNDQYWDDLARYLSAHS 170
Score = 31.6 bits (70), Expect(2) = 7e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 27 IAIYWGQNGNEG 38
>sw|P51614|CHIA_VITVI Acidic endochitinase precursor (EC 3.2.1.14).
Length = 301
Score = 270 bits (690), Expect(2) = 4e-74
Identities = 137/277 (49%), Positives = 167/277 (60%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + QTC TG Y +VN+AFL FG GQTP +NLAGHC+PAS GCT V ++ C
Sbjct: 33 GQNGNEGTLTQTCNTGKYSYVNIAFLNKFGNGQTPEINLAGHCNPASNGCTSVSTGIRNC 92
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LSIGGG GSY LSS DA++VA YLW+N+L
Sbjct: 93 QNRGIKVMLSIGGGAGSYSLSSSNDAQNVANYLWNNFLGGQSSSRPLGDAVLDGIDFDIE 152
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
Q RG +AAPQCPFPD GTAL TGLFDY
Sbjct: 153 LGSTLHWDDLARALSRIEFQQ-------ERGRKVYLTAAPQCPFPDKVPGTALNTGLFDY 205
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTS-IRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VWVQFYNNPPCQYS S +L +W +WTS I + F+GLPA+ AAG GF+PA+ L
Sbjct: 206 VWVQFYNNPPCQYS-SGNTNNLLNSWNRWTSSINSTGSFMGLPASSAAAGRGFIPANVLT 264
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LPV+K S KYGG+MLWS+YYD +GYS ++KS V
Sbjct: 265 SQILPVIKRSPKYGGVMLWSKYYDDQSGYSSSIKSSV 301
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/28 (75%), Positives = 24/28 (85%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFL 473
LGDAVLDG+DFDIE G ++WDDLAR L
Sbjct: 139 LGDAVLDGIDFDIELGSTLHWDDLARAL 166
Score = 31.6 bits (70), Expect(2) = 4e-74
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
IAIYWGQNGNEG
Sbjct: 28 IAIYWGQNGNEG 39
>sw|P29060|CHIA_TOBAC Acidic endochitinase precursor (EC 3.2.1.14).
Length = 291
Score = 271 bits (693), Expect(2) = 5e-74
Identities = 136/276 (49%), Positives = 167/276 (60%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCAT NY VN+AFL FG GQ P LNLAGHCDP +G CTG+ D++AC
Sbjct: 30 GQNGNEGSLADTCATNNYAIVNIAFLVVFGNGQNPVLNLAGHCDPNAGACTGLSNDIRAC 89
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG GSY LSS DAR+VA YLW+NYL
Sbjct: 90 QNQGIKVMLSLGGGAGSYFLSSADDARNVANYLWNNYLGGQSNTRPLGDAVLDGIDFDIE 149
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
+ + +AAPQCPFPD L AL+TGLFDY
Sbjct: 150 GGTTQHWDELAKTLSQFSQQRKVY-----------LTAAPQCPFPDTWLNGALSTGLFDY 198
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDLVA 772
VWVQFYNNPPCQYS + +L W QW +I+AG++FLGLPAA AAGSGF+P+ LV+
Sbjct: 199 VWVQFYNNPPCQYSGGSA-DNLKNYWNQWNAIQAGKIFLGLPAAQGAAGSGFIPSDVLVS 257
Query: 773 QVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
QVLP++ S KYGG+MLWS++YD GYS A+K+ V
Sbjct: 258 QVLPLINGSPKYGGVMLWSKFYD--NGYSSAIKANV 291
Score = 47.8 bits (112), Expect = 3e-04
Identities = 20/32 (62%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DFDIE G +WD+LA+ L +S
Sbjct: 136 LGDAVLDGIDFDIEGGTTQHWDELAKTLSQFS 167
Score = 30.0 bits (66), Expect(2) = 5e-74
Identities = 11/12 (91%), Positives = 11/12 (91%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
I IYWGQNGNEG
Sbjct: 25 IVIYWGQNGNEG 36
>sw|P36910|CHIE_BETVU Acidic endochitinase SE2 precursor (EC
3.2.1.14).
Length = 293
Score = 268 bits (684), Expect(2) = 2e-72
Identities = 140/278 (50%), Positives = 168/278 (60%), Gaps = 2/278 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TC +GNY V +AF+ TFG GQTPALNLAGHCDPA+ C + +D+K C
Sbjct: 33 GQNGDEGSLADTCNSGNYGTVILAFVATFGNGQTPALNLAGHCDPATN-CNSLSSDIKTC 91
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q+ G+KVLLSIGGG G Y LSS DA + A YLW+ YL
Sbjct: 92 QQAGIKVLLSIGGGAGGYSLSSTDDANTFADYLWNTYLGGQSSTRPLGDAVLDGIDFDIE 151
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTC--SAAPQCPFPDASLGTALATGLF 586
AG G T SAAPQCP PDASL TA+ATGLF
Sbjct: 152 SGDGRFWDDLARAL------------AGHNNGQKTVYLSAAPQCPLPDASLSTAIATGLF 199
Query: 587 DYVWVQFYNNPPCQYSASAGVGSLAKAWAQWTSIRAGRVFLGLPAAPQAAGSGFVPASDL 766
DYVWVQFYNNPPCQY SA +L +W QWT+++A ++FLGLPA+ AAGSGF+PA L
Sbjct: 200 DYVWVQFYNNPPCQYDTSAD--NLLSSWNQWTTVQANQIFLGLPASTDAAGSGFIPADAL 257
Query: 767 VAQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+QVLP +K S KYGG+MLWS+ YD +GYS A+KS V
Sbjct: 258 TSQVLPTIKGSAKYGGVMLWSKAYD--SGYSSAIKSSV 293
Score = 53.5 bits (127), Expect = 5e-06
Identities = 23/32 (71%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DFDIESG G +WDDLAR L ++
Sbjct: 138 LGDAVLDGIDFDIESGDGRFWDDLARALAGHN 169
Score = 28.1 bits (61), Expect(2) = 2e-72
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = +3
Query: 39 IAIYWGQNGNEG 74
I IYWGQNG+EG
Sbjct: 28 IVIYWGQNGDEG 39
>sptr|Q8LP06|Q8LP06 Acidic endochitinase.
Length = 302
Score = 256 bits (655), Expect(2) = 7e-72
Identities = 127/273 (46%), Positives = 165/273 (60%), Gaps = 1/273 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTRFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ VA YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVVADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L +G + APQCPFPD +G+AL T LFDY
Sbjct: 157 LGSPQHWDDLARS----------LSKLSYKGRKVYLTGAPQCPFPDRLMGSALNTRLFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC Y++S +L +W +WT SI A ++FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYTSS-NTQNLFDSWNKWTTSITAQKIFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAV 868
+Q+LPV+K S KYGG+MLWS+++D GYS ++
Sbjct: 266 SQILPVLKKSRKYGGVMLWSKFWDDKNGYSSSI 298
Score = 47.0 bits (110), Expect = 5e-04
Identities = 21/32 (65%), Positives = 24/32 (75%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARSLSKLS 174
Score = 37.7 bits (86), Expect(2) = 7e-72
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|Q8H6X6|Q8H6X6 Class III chitinase (EC 3.2.1.14).
Length = 295
Score = 260 bits (665), Expect(2) = 2e-71
Identities = 130/273 (47%), Positives = 168/273 (61%), Gaps = 1/273 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G + TCA+GN+ +VN++FL FGKGQTP +NLAGHC+PA GCT +G +K C
Sbjct: 30 GQNGNEATLNDTCASGNFAYVNLSFLNKFGKGQTPEINLAGHCNPAVNGCTILGPQIKFC 89
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q++GVKV+LS+GGGVG+Y L+S+ DA+ VA YL++N+L
Sbjct: 90 QKLGVKVMLSMGGGVGNYSLASKKDAKDVARYLYNNFLGGRSSFRPLGNARLDGIDFDIE 149
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G +AAPQCPFPD LGTAL TGLFD
Sbjct: 150 LGSSLYYEDLAKY----------LKRYSKLGRKMYLTAAPQCPFPDRLLGTALNTGLFDN 199
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNP CQY+ + V L +W +WT S+ A R+FLGLPAAPQAAGSGF+PA L
Sbjct: 200 VWIQFYNNPSCQYTTN-NVDDLKNSWTRWTTSVNARRIFLGLPAAPQAAGSGFIPAEVLT 258
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAV 868
+LPV+K S KYGG+MLWS+++D TGYS ++
Sbjct: 259 GGILPVIKKSRKYGGVMLWSKFWDEQTGYSASI 291
Score = 48.1 bits (113), Expect = 2e-04
Identities = 20/32 (62%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG+A LDG+DFDIE G +Y++DLA++LK YS
Sbjct: 136 LGNARLDGIDFDIELGSSLYYEDLAKYLKRYS 167
Score = 32.0 bits (71), Expect(2) = 2e-71
Identities = 13/17 (76%), Positives = 14/17 (82%)
Frame = +3
Query: 21 SASRRXIAIYWGQNGNE 71
S +R IAIYWGQNGNE
Sbjct: 19 SIARSGIAIYWGQNGNE 35
>sptr|Q8GRR2|Q8GRR2 Class III chitinase (EC 3.2.1.14).
Length = 295
Score = 260 bits (664), Expect(2) = 3e-71
Identities = 130/273 (47%), Positives = 167/273 (61%), Gaps = 1/273 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G + TCA+GNY +VN++FL FG GQTP +NLAGHC+PA GCT +G +K C
Sbjct: 30 GQNGNEATLNDTCASGNYAYVNLSFLNKFGNGQTPEINLAGHCNPAVNGCTILGPQIKFC 89
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q++GVKV+LS+GGGVG+Y L+S+ DA+ VA YL++N+L
Sbjct: 90 QKLGVKVMLSMGGGVGNYSLASKKDAKDVARYLYNNFLGGRSSFRPLGNARLDGIDFDIE 149
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G +AAPQCPFPD LGTAL TGLFD
Sbjct: 150 LGSSLYYEDLAQY----------LKRYSKLGRKMYLTAAPQCPFPDRLLGTALNTGLFDN 199
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNP CQY+ + V L +W +WT S+ A R+FLGLPAAPQAAGSGF+PA L
Sbjct: 200 VWIQFYNNPSCQYTTN-NVDDLKNSWTRWTTSVNARRIFLGLPAAPQAAGSGFIPAEVLT 258
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAV 868
+LPV+K S KYGG+MLWS+++D TGYS ++
Sbjct: 259 GGILPVIKKSRKYGGVMLWSKFWDEQTGYSASI 291
Score = 47.8 bits (112), Expect = 3e-04
Identities = 20/32 (62%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG+A LDG+DFDIE G +Y++DLA++LK YS
Sbjct: 136 LGNARLDGIDFDIELGSSLYYEDLAQYLKRYS 167
Score = 32.0 bits (71), Expect(2) = 3e-71
Identities = 13/17 (76%), Positives = 14/17 (82%)
Frame = +3
Query: 21 SASRRXIAIYWGQNGNE 71
S +R IAIYWGQNGNE
Sbjct: 19 SIARSGIAIYWGQNGNE 35
>sptr|Q8LP07|Q8LP07 Acidic endochitinase (Fragment).
Length = 295
Score = 254 bits (649), Expect(2) = 3e-71
Identities = 126/270 (46%), Positives = 162/270 (60%), Gaps = 1/270 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VN+AFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNLAFLVKFGNGQTPELNLAGHCNPAANTCTRFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ VA YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVVADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T LFDY
Sbjct: 157 LGSPQHWDDLART----------LSKLSHRGRKVYLTGAPQCPFPDRLMGSALNTRLFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC Y+ S +L +W +WT SI A ++FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYT-SGNTQNLFDSWNKWTTSITAQKIFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYS 859
+Q+LP++K S KYGG+MLWS+++D GYS
Sbjct: 266 SQILPILKKSRKYGGVMLWSKFWDDKNGYS 295
Score = 47.0 bits (110), Expect = 5e-04
Identities = 21/32 (65%), Positives = 24/32 (75%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARTLSKLS 174
Score = 37.7 bits (86), Expect(2) = 3e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22071|O22071 Acidic endochitinase.
Length = 302
Score = 254 bits (648), Expect(2) = 4e-71
Identities = 128/277 (46%), Positives = 164/277 (59%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG G+TP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGRTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ VA YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSRKDAKVVADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLARS----------LSKYSHRGRKIYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC YS S +L +W +WT SI A + FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYS-SGNTQNLFDSWNKWTTSIAAQKFFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +KNS KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKNSRKYGGVMLWSKFWDDKNGYSSSILASV 302
Score = 50.1 bits (118), Expect = 6e-05
Identities = 22/32 (68%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L YS
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARSLSKYS 174
Score = 37.7 bits (86), Expect(2) = 4e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22070|O22070 Acidic endochitinase.
Length = 302
Score = 253 bits (647), Expect(2) = 5e-71
Identities = 127/277 (45%), Positives = 163/277 (58%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLART----------LSKFSHRGRKIYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC YS S +L +W +WT SI A + FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYS-SGNTQNLFDSWNKWTTSIAAQKFFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSISASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARTLSKFS 174
Score = 37.7 bits (86), Expect(2) = 5e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22072|O22072 Acidic endochitinase.
Length = 302
Score = 253 bits (646), Expect(2) = 7e-71
Identities = 126/277 (45%), Positives = 164/277 (59%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLARS----------LSKFSHRGRKVYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC Y+ S +L +W +WT SI A ++FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYT-SGNTQNLFDSWNKWTTSIAAQKIFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSILASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARSLSKFS 174
Score = 37.7 bits (86), Expect(2) = 7e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22067|O22067 Acidic endochitinase.
Length = 302
Score = 253 bits (645), Expect(2) = 9e-71
Identities = 127/277 (45%), Positives = 163/277 (58%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLART----------LSKFSHRGRKIYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC YS S +L +W +WT SI A + FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYS-SGNTQNLFDSWNKWTTSITAQKFFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSILASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARTLSKFS 174
Score = 37.7 bits (86), Expect(2) = 9e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22065|O22065 Acidic endochitinase.
Length = 302
Score = 253 bits (645), Expect(2) = 9e-71
Identities = 127/277 (45%), Positives = 163/277 (58%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLART----------LSKFSHRGRKIYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC YS S +L +W +WT SI A + FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYS-SGNTQNLFDSWNKWTTSIAAQKFFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSIVASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARTLSKFS 174
Score = 37.7 bits (86), Expect(2) = 9e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sw|P19172|CHIA_ARATH Acidic endochitinase precursor (EC 3.2.1.14).
Length = 302
Score = 253 bits (645), Expect(2) = 9e-71
Identities = 127/277 (45%), Positives = 163/277 (58%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLART----------LSKFSHRGRKIYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC YS S +L +W +WT SI A + FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYS-SGNTQNLFDSWNKWTTSIAAQKFFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSILASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARTLSKFS 174
Score = 37.7 bits (86), Expect(2) = 9e-71
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|Q8H6X7|Q8H6X7 Class III chitinase (EC 3.2.1.14).
Length = 295
Score = 258 bits (660), Expect(2) = 9e-71
Identities = 129/273 (47%), Positives = 167/273 (61%), Gaps = 1/273 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G + TCA+GN+ +VN++FL FG GQTP +NLAGHC+PA GCT +G +K C
Sbjct: 30 GQNGNEATLNDTCASGNFAYVNLSFLNKFGNGQTPEINLAGHCNPAVNGCTILGPQIKFC 89
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q++GVKV+LS+GGGVG+Y L+S+ DA+ VA YL++N+L
Sbjct: 90 QKLGVKVMLSMGGGVGNYSLASKKDAKDVARYLYNNFLGGRSSFRPLGNARLDGIDFDIE 149
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L G +AAPQCPFPD LGTAL TGLFD
Sbjct: 150 LGSSLYYEDLAKY----------LKRYSKLGRKMYLTAAPQCPFPDRLLGTALNTGLFDN 199
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNP CQY+ + V L +W +WT S+ A R+FLGLPAAPQAAGSGF+PA L
Sbjct: 200 VWIQFYNNPSCQYTTN-NVDDLKNSWTRWTTSVNARRIFLGLPAAPQAAGSGFIPAEVLT 258
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAV 868
+LPV+K S KYGG+MLWS+++D TGYS ++
Sbjct: 259 GGILPVIKKSRKYGGVMLWSKFWDEQTGYSASI 291
Score = 48.1 bits (113), Expect = 2e-04
Identities = 20/32 (62%), Positives = 27/32 (84%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LG+A LDG+DFDIE G +Y++DLA++LK YS
Sbjct: 136 LGNARLDGIDFDIELGSSLYYEDLAKYLKRYS 167
Score = 32.0 bits (71), Expect(2) = 9e-71
Identities = 13/17 (76%), Positives = 14/17 (82%)
Frame = +3
Query: 21 SASRRXIAIYWGQNGNE 71
S +R IAIYWGQNGNE
Sbjct: 19 SIARSGIAIYWGQNGNE 35
>sptr|Q8LP01|Q8LP01 Acidic endochitinase.
Length = 302
Score = 252 bits (644), Expect(2) = 1e-70
Identities = 126/277 (45%), Positives = 163/277 (58%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKYC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + S+ DA+ VA YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSKEDAKVVADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G AL T FDY
Sbjct: 157 LGSPQHWDDLARS----------LSKLSHRGRKVYLTGAPQCPFPDRLMGNALNTRRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC Y+ S +L +W +WT SI A R+FLGLPAAP+A+GSG++P L
Sbjct: 207 VWIQFYNNPPCSYT-SGNTQNLFDSWNKWTTSITAQRIFLGLPAAPEASGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSILATV 302
Score = 47.0 bits (110), Expect = 5e-04
Identities = 21/32 (65%), Positives = 24/32 (75%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARSLSKLS 174
Score = 37.7 bits (86), Expect(2) = 1e-70
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
>sptr|O22073|O22073 Acidic endochitinase.
Length = 302
Score = 252 bits (644), Expect(2) = 1e-70
Identities = 126/277 (45%), Positives = 164/277 (59%), Gaps = 1/277 (0%)
Frame = +2
Query: 53 GPERQRGXVXQTCATGNYRFVNVAFLPTFGKGQTPALNLAGHCDPASGGCTGVGADVKAC 232
G G + TCATG Y +VNVAFL FG GQTP LNLAGHC+PA+ CT G+ VK C
Sbjct: 37 GQNGNEGNLSATCATGRYAYVNVAFLVKFGNGQTPELNLAGHCNPAANTCTHFGSQVKDC 96
Query: 233 QRMGVKVLLSIGGGVGSYGLSSRADARSVAAYLWDNYLXXXXXXXXXXXXXXXXXXXXXX 412
Q G+KV+LS+GGG+G+Y + SR DA+ +A YLW+N+L
Sbjct: 97 QSRGIKVMLSLGGGIGNYSIGSREDAKVIADYLWNNFLGGKSSSRPLGDAVLDGIDFNIE 156
Query: 413 XXXXXXXXXXXXXXXPGPVPQVLLPGAGARGGPCTCSAAPQCPFPDASLGTALATGLFDY 592
L RG + APQCPFPD +G+AL T FDY
Sbjct: 157 LGSPQHWDDLARS----------LSKFSHRGRKVYLTGAPQCPFPDRLMGSALNTKRFDY 206
Query: 593 VWVQFYNNPPCQYSASAGVGSLAKAWAQWT-SIRAGRVFLGLPAAPQAAGSGFVPASDLV 769
VW+QFYNNPPC Y+ S +L +W +WT SI A ++FLGLPAAP+AAGSG++P L
Sbjct: 207 VWIQFYNNPPCSYT-SGNTQNLFDSWNKWTTSIAAQKLFLGLPAAPEAAGSGYIPPDVLT 265
Query: 770 AQVLPVVKNSTKYGGIMLWSRYYDGLTGYSDAVKSYV 880
+Q+LP +K S KYGG+MLWS+++D GYS ++ + V
Sbjct: 266 SQILPTLKKSRKYGGVMLWSKFWDDKNGYSSSILASV 302
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/32 (65%), Positives = 25/32 (78%)
Frame = +3
Query: 390 LGDAVLDGVDFDIESGGGMYWDDLARFLKSYS 485
LGDAVLDG+DF+IE G +WDDLAR L +S
Sbjct: 143 LGDAVLDGIDFNIELGSPQHWDDLARSLSKFS 174
Score = 37.7 bits (86), Expect(2) = 1e-70
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 15 PHSASRRXIAIYWGQNGNEG 74
P ASR IAIYWGQNGNEG
Sbjct: 24 PSDASRGGIAIYWGQNGNEG 43
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 680,293,232
Number of Sequences: 1395590
Number of extensions: 14008679
Number of successful extensions: 60887
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 53138
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 60175
length of database: 442,889,342
effective HSP length: 124
effective length of database: 269,836,182
effective search space used: 63141666588
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

