BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
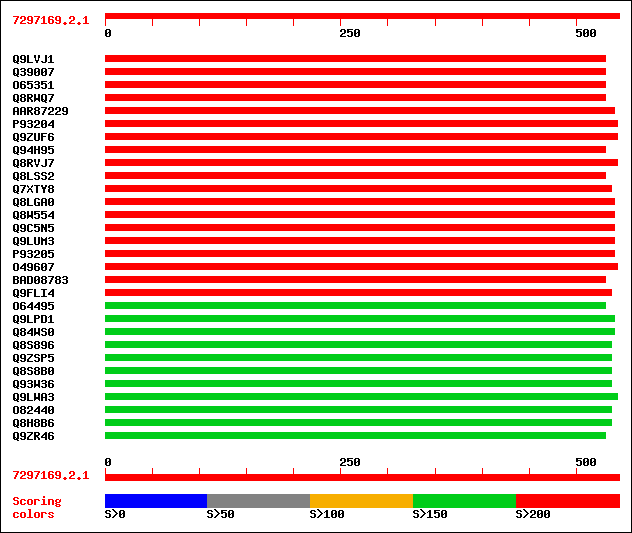
Query= 7297169.2.1
(545 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9LVJ1|Q9LVJ1 Cucumisin-like serine protease, subtilisin-li... 276 2e-73
sptr|Q39007|Q39007 Subtilisin-like protease (Fragment). 249 1e-65
sptr|O65351|O65351 CUCUMISIN-like serine protease (Putative seri... 249 1e-65
sptr|Q8RWQ7|Q8RWQ7 AT5g67360/K8K14_8. 244 4e-64
sptrnew|AAR87229|AAR87229 Putaive subtilisin-like proteinase. 236 1e-61
sptr|P93204|P93204 Serine protease, SBT1. 235 3e-61
sptr|Q9ZUF6|Q9ZUF6 Putative serine protease (Putative subtilisin... 233 9e-61
sptr|Q94H95|Q94H95 Putative serine protease. 231 3e-60
sptr|Q8RVJ7|Q8RVJ7 Putative serine protease. 227 6e-59
sptr|Q8LSS2|Q8LSS2 Putative cucumisin-like serine protease. 223 9e-58
sptr|Q7XTY8|Q7XTY8 OSJNBa0019K04.9 protein. 222 2e-57
sptr|Q8LGA0|Q8LGA0 Subtilisin-like serine protease. 219 2e-56
sptr|Q8W554|Q8W554 AT3g14240/MLN21_2. 218 4e-56
sptr|Q9C5N5|Q9C5N5 Hypothetical protein. 218 4e-56
sptr|Q9LUM3|Q9LUM3 Subtilisin proteinase-like protein. 218 4e-56
sptr|P93205|P93205 Serine protease, SBT2. 218 5e-56
sptr|O49607|O49607 Subtilisin proteinase - like (Putative subtil... 216 1e-55
sptrnew|BAD08783|BAD08783 Putative subtilisin-like proteinase. 216 1e-55
sptr|Q9FLI4|Q9FLI4 Serine protease-like protein (Putative subtil... 213 9e-55
sptr|O64495|O64495 F20D22.12 protein. 191 5e-48
sptr|Q9LPD1|Q9LPD1 F22M8.3 protein. 183 1e-45
sptr|Q84WS0|Q84WS0 Putative subtilisin-like serine protease. 183 1e-45
sptr|Q8S896|Q8S896 Subtilisin-like serine protease AIR3 (Fragment). 181 5e-45
sptr|Q9ZSP5|Q9ZSP5 Subtilisin-like protease. 181 5e-45
sptr|Q8S8B0|Q8S8B0 Subtilisin-like serine protease AIR3 (Fragment). 181 5e-45
sptr|Q93W36|Q93W36 At2g04160/T16B23.1. 181 5e-45
sptr|Q9LWA3|Q9LWA3 Subtilisin-like protease. 180 9e-45
sptr|O82440|O82440 Subtilisin-like protease (Fragment). 180 1e-44
sptr|Q8H8B6|Q8H8B6 Putatvie subtilisin-like serine protease. 179 3e-44
sptr|Q9ZR46|Q9ZR46 P69C protein. 177 6e-44
>sptr|Q9LVJ1|Q9LVJ1 Cucumisin-like serine protease, subtilisin-like
protease.
Length = 777
Score = 276 bits (705), Expect = 2e-73
Identities = 135/177 (76%), Positives = 151/177 (85%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HLVPATMVG K GD+IR Y++T SPTA I F GT+IG SP +PRVAAFSSRGPN+ P
Sbjct: 445 HLVPATMVGAKAGDQIRDYIKTSDSPTAKISFLGTLIGPSPPSPRVAAFSSRGPNHLTPV 504
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPDVIAPGVNILA WTG PTDLDID RRV+FNIISGTSMSCPHVSGLAALLR+AHP
Sbjct: 505 ILKPDVIAPGVNILAGWTGMVGPTDLDIDPRRVQFNIISGTSMSCPHVSGLAALLRKAHP 564
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
+WSPAAIKSAL+TTAY+++NSGE I+DLATG S F+ GAGHVDPN AL+P LVYD
Sbjct: 565 DWSPAAIKSALVTTAYDVENSGEPIEDLATGKSSNSFIHGAGHVDPNKALNPGLVYD 621
>sptr|Q39007|Q39007 Subtilisin-like protease (Fragment).
Length = 746
Score = 249 bits (637), Expect = 1e-65
Identities = 126/177 (71%), Positives = 140/177 (79%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PAT VG+K GD IR+YV TDP+PTA+I GTV+G PS P VAAFSSRGPN P
Sbjct: 431 HLLPATTVGEKAGDIIRHYVTTDPNPTASISILGTVVGVKPS-PVVAAFSSRGPNSITPN 489
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAWTGAA PT L DSRRVEFNIISGTSMSCPHVSGLAALL+ HP
Sbjct: 490 ILKPDLIAPGVNILAAWTGAAGPTGLASDSRRVEFNIISGTSMSCPHVSGLAALLKSVHP 549
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
EWSPAAI+SALMTTAY G+ + D+ATG STPF GAGHV P A +P L+YD
Sbjct: 550 EWSPAAIRSALMTTAYKTYKDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYD 606
>sptr|O65351|O65351 CUCUMISIN-like serine protease (Putative serine
protease) (Putative subtilisin serine protease ARA12).
Length = 757
Score = 249 bits (637), Expect = 1e-65
Identities = 126/177 (71%), Positives = 140/177 (79%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PAT VG+K GD IR+YV TDP+PTA+I GTV+G PS P VAAFSSRGPN P
Sbjct: 442 HLLPATTVGEKAGDIIRHYVTTDPNPTASISILGTVVGVKPS-PVVAAFSSRGPNSITPN 500
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAWTGAA PT L DSRRVEFNIISGTSMSCPHVSGLAALL+ HP
Sbjct: 501 ILKPDLIAPGVNILAAWTGAAGPTGLASDSRRVEFNIISGTSMSCPHVSGLAALLKSVHP 560
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
EWSPAAI+SALMTTAY G+ + D+ATG STPF GAGHV P A +P L+YD
Sbjct: 561 EWSPAAIRSALMTTAYKTYKDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYD 617
>sptr|Q8RWQ7|Q8RWQ7 AT5g67360/K8K14_8.
Length = 757
Score = 244 bits (624), Expect = 4e-64
Identities = 125/177 (70%), Positives = 139/177 (78%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PAT VG+K GD IR+YV TDP+PTA+I GTV+G PS P VAAFSSRGPN P
Sbjct: 442 HLLPATTVGEKAGDIIRHYVTTDPNPTASISILGTVVGVKPS-PVVAAFSSRGPNSITPN 500
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAWTGAA PT L DSRRVEFNIISGTSMSCPHVSGLAALL+ HP
Sbjct: 501 ILKPDLIAPGVNILAAWTGAAGPTGLASDSRRVEFNIISGTSMSCPHVSGLAALLKSVHP 560
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
E SPAAI+SALMTTAY G+ + D+ATG STPF GAGHV P A +P L+YD
Sbjct: 561 ECSPAAIRSALMTTAYKTYKDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYD 617
>sptrnew|AAR87229|AAR87229 Putaive subtilisin-like proteinase.
Length = 765
Score = 236 bits (602), Expect = 1e-61
Identities = 123/183 (67%), Positives = 138/183 (75%), Gaps = 3/183 (1%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PA VG K G I+ YV +DPSPTATIV GT + PS P VAAFSSRGPN PE
Sbjct: 436 HLLPAAGVGAKEGAAIKAYVASDPSPTATIVVAGTQVDVRPS-PVVAAFSSRGPNMLTPE 494
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAWTG A PT + D+RRV FNIISGTSMSCPHVSGLAALLR AHP
Sbjct: 495 ILKPDIIAPGVNILAAWTGKAGPTGIAADTRRVAFNIISGTSMSCPHVSGLAALLRSAHP 554
Query: 361 EWSPAAIKSALMTTAYNL---DNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
EWSPAA++SALMTTAY+ + D ATG +TPF GAGHVDP +A+DP LVYD
Sbjct: 555 EWSPAAVRSALMTTAYSTYAGAGDANPLLDAATGAPATPFDYGAGHVDPASAVDPGLVYD 614
Query: 532 AGS 540
G+
Sbjct: 615 LGT 617
>sptr|P93204|P93204 Serine protease, SBT1.
Length = 766
Score = 235 bits (599), Expect = 3e-61
Identities = 117/181 (64%), Positives = 140/181 (77%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+P VGQ G+ I+ Y+ ++ +PTATI F GT +G PS P VAAFSSRGPN P+
Sbjct: 442 HLIPTAAVGQTAGNLIKQYIASNSNPTATIAFGGTKLGVQPS-PVVAAFSSRGPNPITPD 500
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
+LKPD+IAPGVNILA WTG PT L D+R V FNIISGTSMSCPHVSGLAALL+ AHP
Sbjct: 501 VLKPDLIAPGVNILAGWTGKVGPTGLQEDTRNVGFNIISGTSMSCPHVSGLAALLKAAHP 560
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAGS 540
EWSPAAI+SALMTT+Y+ +G+TI+D+ATG+ STPF GAGHV+P AA+ P LVYD
Sbjct: 561 EWSPAAIRSALMTTSYSTYKNGKTIEDVATGMSSTPFDYGAGHVNPTAAVSPGLVYDLTV 620
Query: 541 D 543
D
Sbjct: 621 D 621
>sptr|Q9ZUF6|Q9ZUF6 Putative serine protease (Putative subtilisin
serine protease).
Length = 754
Score = 233 bits (595), Expect = 9e-61
Identities = 116/181 (64%), Positives = 136/181 (75%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PA VG+K GD +R YV++D PTA +VF+GTV+ PS P VAAFSSRGPN PE
Sbjct: 436 HLLPAIAVGKKTGDLLREYVKSDSKPTALLVFKGTVLDVKPS-PVVAAFSSRGPNTVTPE 494
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPDVI PGVNILA W+ A PT LD DSRR +FNI+SGTSMSCPH+SGLA LL+ AHP
Sbjct: 495 ILKPDVIGPGVNILAGWSDAIGPTGLDKDSRRTQFNIMSGTSMSCPHISGLAGLLKAAHP 554
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAGS 540
EWSP+AIKSALMTTAY LDN+ + D A S P+ G+GHVDP AL P LVYD +
Sbjct: 555 EWSPSAIKSALMTTAYVLDNTNAPLHDAADNSLSNPYAHGSGHVDPQKALSPGLVYDIST 614
Query: 541 D 543
+
Sbjct: 615 E 615
>sptr|Q94H95|Q94H95 Putative serine protease.
Length = 764
Score = 231 bits (590), Expect = 3e-60
Identities = 119/177 (67%), Positives = 139/177 (78%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++P VGQK GD +R Y +DP+PTA+IVF GT +G PS P VAAFSSRGPN P
Sbjct: 446 HVLPGAGVGQKAGDTMRAYALSDPNPTASIVFAGTQVGIQPS-PVVAAFSSRGPNTVTPG 504
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAW+G+ P+ L DSRRV FNIISGTSMSCPHVSGLAALLR AH
Sbjct: 505 ILKPDLIAPGVNILAAWSGSVGPSGLAGDSRRVGFNIISGTSMSCPHVSGLAALLRAAHQ 564
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
+WSPAAI+SALMTT+YN +G I D+ATG+ +TP GAGHVDP+ A+DP LVYD
Sbjct: 565 DWSPAAIRSALMTTSYNGYPNGNGILDVATGLPATPLDVGAGHVDPSKAVDPGLVYD 621
>sptr|Q8RVJ7|Q8RVJ7 Putative serine protease.
Length = 566
Score = 227 bits (579), Expect = 6e-59
Identities = 118/181 (65%), Positives = 132/181 (72%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+P VG + + I+ Y DP P TI GT +G PS P VAAFSSRGPN PE
Sbjct: 242 HLLPTAAVGLRTANAIKNYAFLDPKPMGTIASGGTKLGVEPS-PVVAAFSSRGPNLVTPE 300
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
+LKPD+IAPGVNILA WTG A PT L D R VEFNIISGTSMSCPHVSGLAAL++ AH
Sbjct: 301 VLKPDLIAPGVNILAGWTGGAGPTGLTNDKRHVEFNIISGTSMSCPHVSGLAALIKAAHQ 360
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAGS 540
+WSPAAIKSALMTTAY +GE + D+ATG STPF GAGHV+P AALDP LVYDA
Sbjct: 361 DWSPAAIKSALMTTAYATYKNGEDLLDVATGQPSTPFDYGAGHVNPVAALDPGLVYDATV 420
Query: 541 D 543
D
Sbjct: 421 D 421
>sptr|Q8LSS2|Q8LSS2 Putative cucumisin-like serine protease.
Length = 773
Score = 223 bits (569), Expect = 9e-58
Identities = 115/182 (63%), Positives = 133/182 (73%), Gaps = 5/182 (2%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPS-----PTATIVFRGTVIGKSPSAPRVAAFSSRGPN 165
HL+PA VG+ GDKIR Y + P A + F GTV+G PS P VAAFSSRGPN
Sbjct: 451 HLLPAVAVGKLAGDKIREYASRRAAGGAGAPMAILSFGGTVLGVRPS-PVVAAFSSRGPN 509
Query: 166 YRAPEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALL 345
PEILKPD+I PGVNILA W+G A PT L D RR FNIISGTSMSCPH+SG+AALL
Sbjct: 510 TVVPEILKPDMIGPGVNILAGWSGVAGPTGLVKDGRRTHFNIISGTSMSCPHISGVAALL 569
Query: 346 RQAHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLV 525
+ AHPEWSPAAIKSALMTTAY +DN+ +++D A G+ +TPF GAGHVDP AL P L+
Sbjct: 570 KAAHPEWSPAAIKSALMTTAYTVDNTNSSLRDAAGGLLATPFAFGAGHVDPQKALSPGLL 629
Query: 526 YD 531
YD
Sbjct: 630 YD 631
>sptr|Q7XTY8|Q7XTY8 OSJNBa0019K04.9 protein.
Length = 776
Score = 222 bits (566), Expect = 2e-57
Identities = 114/179 (63%), Positives = 134/179 (74%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PA VG+ G + Y ++ P PTAT+ F GT +G PS P VAAFSSRGPN E
Sbjct: 458 HLLPAVAVGEAEGIAAKSYSKSAPKPTATLSFGGTKLGIRPS-PVVAAFSSRGPNILTLE 516
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPDV+APGVNILAAW+G ASP+ L DSRRV FNI+SGTSMSCPHV+G+AAL++ +HP
Sbjct: 517 ILKPDVVAPGVNILAAWSGDASPSSLSSDSRRVGFNILSGTSMSCPHVAGVAALIKASHP 576
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
+WSPA IKSALMTTAY DN+ +KD ATG STPF GAGH+ P AL P LVYD G
Sbjct: 577 DWSPAQIKSALMTTAYVHDNTYRPMKDAATGKASTPFEHGAGHIHPVRALTPGLVYDIG 635
>sptr|Q8LGA0|Q8LGA0 Subtilisin-like serine protease.
Length = 775
Score = 219 bits (557), Expect = 2e-56
Identities = 113/186 (60%), Positives = 138/186 (74%), Gaps = 6/186 (3%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYV------QTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGP 162
H++PAT VG GD+IR Y+ ++ PTATIVF+GT +G P AP VA+FS+RGP
Sbjct: 443 HVLPATSVGASGGDEIRRYISESSKSRSSKHPTATIVFKGTRLGIRP-APVVASFSARGP 501
Query: 163 NYRAPEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAAL 342
N PEILKPDVIAPG+NILAAW P+ + D+RR EFNI+SGTSM+CPHVSGLAAL
Sbjct: 502 NPETPEILKPDVIAPGLNILAAWPDRIGPSGVTSDNRRTEFNILSGTSMACPHVSGLAAL 561
Query: 343 LRQAHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRL 522
L+ AHP+WSPAAI+SALMTTAY +DNSGE + D +TG S+ G+GHV P A+DP L
Sbjct: 562 LKAAHPDWSPAAIRSALMTTAYTVDNSGEPMMDESTGNTSSVTDYGSGHVHPTRAMDPGL 621
Query: 523 VYDAGS 540
VYD S
Sbjct: 622 VYDITS 627
>sptr|Q8W554|Q8W554 AT3g14240/MLN21_2.
Length = 581
Score = 218 bits (555), Expect = 4e-56
Identities = 112/186 (60%), Positives = 138/186 (74%), Gaps = 6/186 (3%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYV------QTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGP 162
H++PAT VG GD+IR Y+ ++ PTATIVF+GT +G P AP VA+FS+RGP
Sbjct: 249 HVLPATSVGASGGDEIRRYISESSKSRSSKHPTATIVFKGTRLGIRP-APVVASFSARGP 307
Query: 163 NYRAPEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAAL 342
N PEILKPDVIAPG+NILAAW P+ + D+RR EFNI+SGTSM+CPHVSGLAAL
Sbjct: 308 NPETPEILKPDVIAPGLNILAAWPDRIGPSGVTSDNRRTEFNILSGTSMACPHVSGLAAL 367
Query: 343 LRQAHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRL 522
L+ AHP+WSPAAI+SAL+TTAY +DNSGE + D +TG S+ G+GHV P A+DP L
Sbjct: 368 LKAAHPDWSPAAIRSALITTAYTVDNSGEPMMDESTGNTSSVMDYGSGHVHPTKAMDPGL 427
Query: 523 VYDAGS 540
VYD S
Sbjct: 428 VYDITS 433
>sptr|Q9C5N5|Q9C5N5 Hypothetical protein.
Length = 775
Score = 218 bits (555), Expect = 4e-56
Identities = 112/186 (60%), Positives = 138/186 (74%), Gaps = 6/186 (3%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYV------QTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGP 162
H++PAT VG GD+IR Y+ ++ PTATIVF+GT +G P AP VA+FS+RGP
Sbjct: 443 HVLPATSVGASGGDEIRRYISESSKSRSSKHPTATIVFKGTRLGIRP-APVVASFSARGP 501
Query: 163 NYRAPEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAAL 342
N PEILKPDVIAPG+NILAAW P+ + D+RR EFNI+SGTSM+CPHVSGLAAL
Sbjct: 502 NPETPEILKPDVIAPGLNILAAWPDRIGPSGVTSDNRRTEFNILSGTSMACPHVSGLAAL 561
Query: 343 LRQAHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRL 522
L+ AHP+WSPAAI+SAL+TTAY +DNSGE + D +TG S+ G+GHV P A+DP L
Sbjct: 562 LKAAHPDWSPAAIRSALITTAYTVDNSGEPMMDESTGNTSSVMDYGSGHVHPTKAMDPGL 621
Query: 523 VYDAGS 540
VYD S
Sbjct: 622 VYDITS 627
>sptr|Q9LUM3|Q9LUM3 Subtilisin proteinase-like protein.
Length = 775
Score = 218 bits (555), Expect = 4e-56
Identities = 112/186 (60%), Positives = 138/186 (74%), Gaps = 6/186 (3%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYV------QTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGP 162
H++PAT VG GD+IR Y+ ++ PTATIVF+GT +G P AP VA+FS+RGP
Sbjct: 443 HVLPATSVGASGGDEIRRYISESSKSRSSKHPTATIVFKGTRLGIRP-APVVASFSARGP 501
Query: 163 NYRAPEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAAL 342
N PEILKPDVIAPG+NILAAW P+ + D+RR EFNI+SGTSM+CPHVSGLAAL
Sbjct: 502 NPETPEILKPDVIAPGLNILAAWPDRIGPSGVTSDNRRTEFNILSGTSMACPHVSGLAAL 561
Query: 343 LRQAHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRL 522
L+ AHP+WSPAAI+SAL+TTAY +DNSGE + D +TG S+ G+GHV P A+DP L
Sbjct: 562 LKAAHPDWSPAAIRSALITTAYTVDNSGEPMMDESTGNTSSVMDYGSGHVHPTKAMDPGL 621
Query: 523 VYDAGS 540
VYD S
Sbjct: 622 VYDITS 627
>sptr|P93205|P93205 Serine protease, SBT2.
Length = 775
Score = 218 bits (554), Expect = 5e-56
Identities = 110/180 (61%), Positives = 136/180 (75%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PA VG++ G I+ Y S TAT+ F GT +G PS P VAAFSSRGPN+ + E
Sbjct: 457 HLLPAVAVGEREGRAIKLYA-AGRSATATLRFLGTKLGIRPS-PVVAAFSSRGPNFLSLE 514
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD++APGVNILA WTGA P+ L ID RR FNI+SGTSMSCPHVSG+AALL+ HP
Sbjct: 515 ILKPDMVAPGVNILAGWTGALGPSSLPIDQRRTNFNILSGTSMSCPHVSGIAALLKARHP 574
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAGS 540
+WSPAAIKSALMTTAY DN+ +++KD ++ STP+ GAGHV+P A+DP L+YD G+
Sbjct: 575 DWSPAAIKSALMTTAYVHDNTYKSLKDASSVTPSTPYDHGAGHVNPRKAVDPGLIYDIGA 634
>sptr|O49607|O49607 Subtilisin proteinase - like (Putative
subtilisin serine proteinase).
Length = 764
Score = 216 bits (550), Expect = 1e-55
Identities = 104/181 (57%), Positives = 135/181 (74%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PA VG GD+I+ Y + P+P A+I FRGT++G P AP +A+FS RGPN +PE
Sbjct: 438 HLIPACAVGSNEGDRIKAYASSHPNPIASIDFRGTIVGIKP-APVIASFSGRGPNGLSPE 496
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNILAAWT A PT L D R+ EFNI+SGTSM+CPHVSG AALL+ AHP
Sbjct: 497 ILKPDLIAPGVNILAAWTDAVGPTGLPSDPRKTEFNILSGTSMACPHVSGAAALLKSAHP 556
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAGS 540
+WSPA I+SA+MTT +DNS ++ D +TG +TP+ G+GH++ A++P LVYD +
Sbjct: 557 DWSPAVIRSAMMTTTNLVDNSNRSLIDESTGKSATPYDYGSGHLNLGRAMNPGLVYDITN 616
Query: 541 D 543
D
Sbjct: 617 D 617
>sptrnew|BAD08783|BAD08783 Putative subtilisin-like proteinase.
Length = 796
Score = 216 bits (550), Expect = 1e-55
Identities = 113/180 (62%), Positives = 134/180 (74%), Gaps = 3/180 (1%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYV--QTDPSP-TATIVFRGTVIGKSPSAPRVAAFSSRGPNYR 171
H++PAT VG GDK+R Y+ T +P T TI+F GT +G P AP VAAFS+RGPN +
Sbjct: 463 HVLPATAVGAAAGDKLRKYIGSSTRQAPATGTILFEGTHLGVHP-APVVAAFSARGPNPQ 521
Query: 172 APEILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQ 351
+PEILKPD+IAPG+NILAAW P + D RR EFNI+SGTSM+CPH+SGLAALL+
Sbjct: 522 SPEILKPDLIAPGLNILAAWPSGVGPAGIPSDGRRTEFNILSGTSMACPHISGLAALLKA 581
Query: 352 AHPEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
AHP WSPAAIKSALMTTAY DNS T+ D +TGV + F GAGHVDP A+DP LVYD
Sbjct: 582 AHPTWSPAAIKSALMTTAYIKDNSNGTMVDESTGVVADVFDFGAGHVDPMRAMDPGLVYD 641
>sptr|Q9FLI4|Q9FLI4 Serine protease-like protein (Putative
subtilisin serine protease).
Length = 780
Score = 213 bits (543), Expect = 9e-55
Identities = 107/179 (59%), Positives = 131/179 (73%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PA VG+K G I+ Y T TA++ GT IG PS P VAAFSSRGPN+ + E
Sbjct: 460 HMLPAVAVGEKEGKLIKQYAMTSKKATASLEILGTRIGIKPS-PVVAAFSSRGPNFLSLE 518
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD++APGVNILAAWTG +P+ L D RRV+FNI+SGTSMSCPHVSG+AAL++ HP
Sbjct: 519 ILKPDLLAPGVNILAAWTGDMAPSSLSSDPRRVKFNILSGTSMSCPHVSGVAALIKSRHP 578
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
+WSPAAIKSALMTTAY DN + + D + S+P+ GAGH+DP A DP LVYD G
Sbjct: 579 DWSPAAIKSALMTTAYVHDNMFKPLTDASGAAPSSPYDHGAGHIDPLRATDPGLVYDIG 637
>sptr|O64495|O64495 F20D22.12 protein.
Length = 775
Score = 191 bits (485), Expect = 5e-48
Identities = 100/177 (56%), Positives = 122/177 (68%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
HL+PAT++G ++ YV P A I+F GTVIG+S AP VA FS+RGP+ P
Sbjct: 452 HLLPATLIGYTESVLLKAYVNATVKPKARIIFGGTVIGRS-RAPEVAQFSARGPSLANPS 510
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+IAPGVNI+AAW PT L DSRRV F ++SGTSMSCPHVSG+ AL+R A+P
Sbjct: 511 ILKPDMIAPGVNIIAAWPQNLGPTGLPYDSRRVNFTVMSGTSMSCPHVSGITALIRSAYP 570
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
WSPAAIKSALMTTA D G+ IKD + F GAGHV+P A++P LVY+
Sbjct: 571 NWSPAAIKSALMTTADLYDRQGKAIKD--GNKPAGVFAIGAGHVNPQKAINPGLVYN 625
>sptr|Q9LPD1|Q9LPD1 F22M8.3 protein.
Length = 756
Score = 183 bits (464), Expect = 1e-45
Identities = 98/183 (53%), Positives = 124/183 (67%), Gaps = 3/183 (1%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PA +G G + Y+ + TA++ FRGT G + AP VAAFSSRGP+ PE
Sbjct: 435 HVLPAVSLGFSDGKTLLNYLAGAANATASVRFRGTAYGAT--APMVAAFSSRGPSVAGPE 492
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
I KPD+ APG+NILA W+ +SP+ L D RRV+FNIISGTSM+CPH+SG+AAL++ H
Sbjct: 493 IAKPDIAAPGLNILAGWSPFSSPSLLRSDPRRVQFNIISGTSMACPHISGIAALIKSVHG 552
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDL-ATGVES--TPFVRGAGHVDPNAALDPRLVYD 531
+WSPA IKSA+MTTA DN I D A G ES T F GAG+VDP A+DP LVYD
Sbjct: 553 DWSPAMIKSAIMTTARITDNRNRPIGDRGAAGAESAATAFAFGAGNVDPTRAVDPGLVYD 612
Query: 532 AGS 540
+
Sbjct: 613 TST 615
>sptr|Q84WS0|Q84WS0 Putative subtilisin-like serine protease.
Length = 774
Score = 183 bits (464), Expect = 1e-45
Identities = 98/183 (53%), Positives = 124/183 (67%), Gaps = 3/183 (1%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PA +G G + Y+ + TA++ FRGT G + AP VAAFSSRGP+ PE
Sbjct: 453 HVLPAVSLGFSDGKTLLNYLAGAANATASVRFRGTAYGAT--APMVAAFSSRGPSVAGPE 510
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
I KPD+ APG+NILA W+ +SP+ L D RRV+FNIISGTSM+CPH+SG+AAL++ H
Sbjct: 511 IAKPDIAAPGLNILAGWSPFSSPSLLRSDPRRVQFNIISGTSMACPHISGIAALIKSVHG 570
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDL-ATGVES--TPFVRGAGHVDPNAALDPRLVYD 531
+WSPA IKSA+MTTA DN I D A G ES T F GAG+VDP A+DP LVYD
Sbjct: 571 DWSPAMIKSAIMTTARITDNRNRPIGDRGAAGAESAATAFAFGAGNVDPTRAVDPGLVYD 630
Query: 532 AGS 540
+
Sbjct: 631 TST 633
>sptr|Q8S896|Q8S896 Subtilisin-like serine protease AIR3 (Fragment).
Length = 755
Score = 181 bits (459), Expect = 5e-45
Identities = 93/179 (51%), Positives = 126/179 (70%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PAT + K + Y+ P A I T +G P AP +A+FSS+GP+ AP+
Sbjct: 461 HVLPATQLTSKDSFAVSRYISQTKKPIAHITPSRTDLGLKP-APVMASFSSKGPSIVAPQ 519
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+ APGV+++AA+TGA SPT+ D RR+ FN ISGTSMSCPH+SG+A LL+ +P
Sbjct: 520 ILKPDITAPGVSVIAAYTGAVSPTNEQFDPRRLLFNAISGTSMSCPHISGIAGLLKTRYP 579
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
WSPAAI+SA+MTTA +D+ I++ AT +++TPF GAGHV PN A++P LVYD G
Sbjct: 580 SWSPAAIRSAIMTTATIMDDIPGPIQN-ATNMKATPFSFGAGHVQPNLAVNPGLVYDLG 637
>sptr|Q9ZSP5|Q9ZSP5 Subtilisin-like protease.
Length = 772
Score = 181 bits (459), Expect = 5e-45
Identities = 93/179 (51%), Positives = 126/179 (70%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PAT + K + Y+ P A I T +G P AP +A+FSS+GP+ AP+
Sbjct: 461 HVLPATQLTSKDSFAVSRYISQTKKPIAHITPSRTDLGLKP-APVMASFSSKGPSIVAPQ 519
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+ APGV+++AA+TGA SPT+ D RR+ FN ISGTSMSCPH+SG+A LL+ +P
Sbjct: 520 ILKPDITAPGVSVIAAYTGAVSPTNEQFDPRRLLFNAISGTSMSCPHISGIAGLLKTRYP 579
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
WSPAAI+SA+MTTA +D+ I++ AT +++TPF GAGHV PN A++P LVYD G
Sbjct: 580 SWSPAAIRSAIMTTATIMDDIPGPIQN-ATNMKATPFSFGAGHVQPNLAVNPGLVYDLG 637
>sptr|Q8S8B0|Q8S8B0 Subtilisin-like serine protease AIR3 (Fragment).
Length = 578
Score = 181 bits (459), Expect = 5e-45
Identities = 93/179 (51%), Positives = 126/179 (70%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PAT + K + Y+ P A I T +G P AP +A+FSS+GP+ AP+
Sbjct: 267 HVLPATQLTSKDSFAVSRYISQTKKPIAHITPSRTDLGLKP-APVMASFSSKGPSIVAPQ 325
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+ APGV+++AA+TGA SPT+ D RR+ FN ISGTSMSCPH+SG+A LL+ +P
Sbjct: 326 ILKPDITAPGVSVIAAYTGAVSPTNEQFDPRRLLFNAISGTSMSCPHISGIAGLLKTRYP 385
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
WSPAAI+SA+MTTA +D+ I++ AT +++TPF GAGHV PN A++P LVYD G
Sbjct: 386 SWSPAAIRSAIMTTATIMDDIPGPIQN-ATNMKATPFSFGAGHVQPNLAVNPGLVYDLG 443
>sptr|Q93W36|Q93W36 At2g04160/T16B23.1.
Length = 421
Score = 181 bits (459), Expect = 5e-45
Identities = 93/179 (51%), Positives = 126/179 (70%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PAT + K + Y+ P A I T +G P AP +A+FSS+GP+ AP+
Sbjct: 110 HVLPATQLTSKDSFAVSRYISQTKKPIAHITPSRTDLGLKP-APVMASFSSKGPSIVAPQ 168
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+ APGV+++AA+TGA SPT+ D RR+ FN ISGTSMSCPH+SG+A LL+ +P
Sbjct: 169 ILKPDITAPGVSVIAAYTGAVSPTNEQFDPRRLLFNAISGTSMSCPHISGIAGLLKTRYP 228
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
WSPAAI+SA+MTTA +D+ I++ AT +++TPF GAGHV PN A++P LVYD G
Sbjct: 229 SWSPAAIRSAIMTTATIMDDIPGPIQN-ATNMKATPFSFGAGHVQPNLAVNPGLVYDLG 286
>sptr|Q9LWA3|Q9LWA3 Subtilisin-like protease.
Length = 746
Score = 180 bits (457), Expect = 9e-45
Identities = 99/182 (54%), Positives = 127/182 (69%), Gaps = 1/182 (0%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PA V G +I Y + +PTA I F+GT+IG +AP VA+FSSRGPN +P
Sbjct: 436 HVLPALEVSAADGIRILTYTNSISNPTAKITFQGTIIGDE-NAPIVASFSSRGPNKPSPG 494
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSR-RVEFNIISGTSMSCPHVSGLAALLRQAH 357
ILKPD+I PGVNILAAW PT +D + + + FNIISGTSMSCPH+SG+AALL+ H
Sbjct: 495 ILKPDIIGPGVNILAAW-----PTSVDDNKKTKSTFNIISGTSMSCPHLSGVAALLKSTH 549
Query: 358 PEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
P+WSPAAIKSA+MTTAY L+ + I D + + F GAGHV+P++A DP LVYD
Sbjct: 550 PDWSPAAIKSAIMTTAYTLNLASSPILDERL-LPADIFAIGAGHVNPSSANDPGLVYDTP 608
Query: 538 SD 543
S+
Sbjct: 609 SE 610
>sptr|O82440|O82440 Subtilisin-like protease (Fragment).
Length = 758
Score = 180 bits (456), Expect = 1e-44
Identities = 92/179 (51%), Positives = 126/179 (70%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++P+T + K + Y+ P A I T +G P AP +A+FSS+GP+ AP+
Sbjct: 447 HVLPSTQLTSKDSFAVSRYMTQTKKPIAHITPSRTDLGLKP-APVMASFSSKGPSIVAPQ 505
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAHP 360
ILKPD+ APGV+++AA+TGA SPT+ D RR+ FN ISGTSMSCPH+SG+A LL+ +P
Sbjct: 506 ILKPDITAPGVSVIAAYTGAVSPTNEQFDPRRLLFNAISGTSMSCPHISGIAGLLKTRYP 565
Query: 361 EWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
WSPAAI+SA+MTTA +D+ I++ AT +++TPF GAGHV PN A++P LVYD G
Sbjct: 566 SWSPAAIRSAIMTTATTMDDIPGPIQN-ATNMKATPFSFGAGHVQPNLAVNPGLVYDLG 623
>sptr|Q8H8B6|Q8H8B6 Putatvie subtilisin-like serine protease.
Length = 663
Score = 179 bits (453), Expect = 3e-44
Identities = 98/180 (54%), Positives = 121/180 (67%), Gaps = 1/180 (0%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQ-TDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAP 177
+ +P V + G +I Y++ + PTA T++GKSP AP VA FSSRGP+ +P
Sbjct: 311 NFLPTVHVDLRQGTRILDYIRGSSRPPTARFSPSTTLVGKSP-APAVAYFSSRGPSSISP 369
Query: 178 EILKPDVIAPGVNILAAWTGAASPTDLDIDSRRVEFNIISGTSMSCPHVSGLAALLRQAH 357
ILKPDV APGVNILAAW +SPT + +D R V +N SGTSMSCPHVSG+ A++R H
Sbjct: 370 HILKPDVTAPGVNILAAWPPMSSPTVIPLDKRSVTWNFDSGTSMSCPHVSGIVAVVRAVH 429
Query: 358 PEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYDAG 537
P WSPAAIKSALMTTAY D++ + + T + F GAGHVDP ALDP LVYDAG
Sbjct: 430 PTWSPAAIKSALMTTAYMYDDTSDVMLAGGTLKAADAFDVGAGHVDPLRALDPGLVYDAG 489
>sptr|Q9ZR46|Q9ZR46 P69C protein.
Length = 666
Score = 177 bits (450), Expect = 6e-44
Identities = 98/178 (55%), Positives = 123/178 (69%), Gaps = 1/178 (0%)
Frame = +1
Query: 1 HLVPATMVGQKFGDKIRYYVQTDPSPTATIVFRGTVIGKSPSAPRVAAFSSRGPNYRAPE 180
H++PA V G KI Y+ + +P ATI F+GT+IG +AP VAAFSSRGP+ +P
Sbjct: 435 HVLPALDVSDADGTKILAYMNSTSNPVATIAFQGTIIGDK-NAPMVAAFSSRGPSRASPG 493
Query: 181 ILKPDVIAPGVNILAAWTGAASPTDLDIDS-RRVEFNIISGTSMSCPHVSGLAALLRQAH 357
ILKPD+I PGVNILAAW PT +D + + FNIISGTSMSCPH+SG+AALL+ H
Sbjct: 494 ILKPDIIGPGVNILAAW-----PTSVDDNKDTKSTFNIISGTSMSCPHLSGVAALLKSTH 548
Query: 358 PEWSPAAIKSALMTTAYNLDNSGETIKDLATGVESTPFVRGAGHVDPNAALDPRLVYD 531
P+WSPAAIKSA+MTTA L+ + I D + + F GAGHV+P+ A DP LVYD
Sbjct: 549 PDWSPAAIKSAIMTTADTLNLANSPILDERL-LPADIFATGAGHVNPSRANDPGLVYD 605
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 332,801,608
Number of Sequences: 1395590
Number of extensions: 6326902
Number of successful extensions: 28571
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 27055
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 28019
length of database: 442,889,342
effective HSP length: 115
effective length of database: 282,396,492
effective search space used: 18638168472
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)

