BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
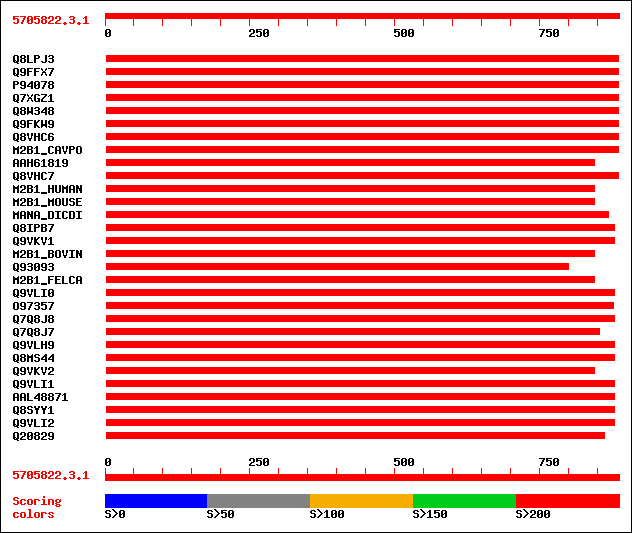
Query= 5705822.3.1
(888 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8LPJ3|Q8LPJ3 Alpha-mannosidase. 486 e-136
sptr|Q9FFX7|Q9FFX7 Alpha-mannosidase. 486 e-136
sptr|P94078|P94078 Alpha-mannosidase (AT3G26720/MLJ15_12). 473 e-132
sptr|Q7XGZ1|Q7XGZ1 Putative alpha-mannosidase. 437 e-121
sptr|Q8W348|Q8W348 Putative alpha-mannosidase. 437 e-121
sptr|Q9FKW9|Q9FKW9 Alpha-mannosidase. 396 e-109
sptr|Q8VHC6|Q8VHC6 Lysosomal alpha-mannosidase (EC 3.2.1.24). 285 8e-76
sw|Q8VHC8|M2B1_CAVPO Lysosomal alpha-mannosidase precursor (EC 3... 285 8e-76
sptrnew|AAH61819|AAH61819 Similar to mannosidase 2, alpha B1. 284 1e-75
sptr|Q8VHC7|Q8VHC7 Lysosomal alpha-mannosidase (EC 3.2.1.24). 281 7e-75
sw|O00754|M2B1_HUMAN Lysosomal alpha-mannosidase precursor (EC 3... 274 1e-72
sw|O09159|M2B1_MOUSE Lysosomal alpha-mannosidase precursor (EC 3... 272 5e-72
sw|P34098|MANA_DICDI Lysosomal alpha-mannosidase precursor (EC 3... 271 7e-72
sptr|Q8IPB7|Q8IPB7 CG6206-PB. 271 9e-72
sptr|Q9VKV1|Q9VKV1 CG6206 protein (BCDNA.GH02419). 271 1e-71
sw|Q29451|M2B1_BOVIN Lysosomal alpha-mannosidase precursor (EC 3... 264 1e-69
sptr|Q93093|Q93093 Truncated lysosomal acid alpha-mannosidase (E... 262 4e-69
sw|O46432|M2B1_FELCA Lysosomal alpha-mannosidase precursor (EC 3... 262 6e-69
sptr|Q9VLI0|Q9VLI0 CG9466 protein (GH02475P). 251 1e-65
sptr|O97357|O97357 Lysosomal acid alpha-mannosidase precursor. 250 2e-65
sptr|Q7Q8J8|Q7Q8J8 AgCP15268 (Fragment). 249 4e-65
sptr|Q7Q8J7|Q7Q8J7 AgCP15371 (Fragment). 248 8e-65
sptr|Q9VLH9|Q9VLH9 CG9468 protein. 244 9e-64
sptr|Q8MS44|Q8MS44 RE08556p. 244 9e-64
sptr|Q9VKV2|Q9VKV2 CG5322 protein. 240 2e-62
sptr|Q9VLI1|Q9VLI1 CG9465 protein. 233 2e-60
sptrnew|AAL48871|AAL48871 RE28991p (Fragment). 233 2e-60
sptr|Q8SYY1|Q8SYY1 RE28991p. 233 2e-60
sptr|Q9VLI2|Q9VLI2 CG9463 protein. 233 2e-60
sptr|Q20829|Q20829 Hypothetical protein. 215 6e-55
>sptr|Q8LPJ3|Q8LPJ3 Alpha-mannosidase.
Length = 1024
Score = 486 bits (1250), Expect = e-136
Identities = 231/295 (78%), Positives = 257/295 (87%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYYVGSNNSIQ ACVQNVLDS++PALL D+NRKFIYVEQAFFQRWW Q++
Sbjct: 48 DVGWLKTVDQYYVGSNNSIQVACVQNVLDSIVPALLADKNRKFIYVEQAFFQRWWNEQSE 107
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
IK VK LI SG+LELINGGMCMHDEAA HYIDMIDQTTLGH++I EF PRIGWQI
Sbjct: 108 EIKRIVKELIHSGQLELINGGMCMHDEAAPHYIDMIDQTTLGHRFIIREFNVTPRIGWQI 167
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHSAVQAYLLGAEVGFD+ +F RIDYQDR+ R K LEV+WRGSK+LGSS+ IFAG
Sbjct: 168 DPFGHSAVQAYLLGAEVGFDSVFFGRIDYQDREKRYKEKTLEVIWRGSKSLGSSSQIFAG 227
Query: 543 IFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHIMF 722
FP NYEPPPG FY+E+ D+SPVVQDDP LFDYNV+ERVN FVAAA+ QANITR NHIMF
Sbjct: 228 AFPTNYEPPPGGFYYEITDDSPVVQDDPDLFDYNVQERVNAFVAAALDQANITRINHIMF 287
Query: 723 TMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAANEQGPLQ 887
TMGTDF+YQYA +W+ MDKLIHYVN DGR++A YSTPSIYTDAK+AANE PL+
Sbjct: 288 TMGTDFRYQYAHTWYRQMDKLIHYVNLDGRVNAFYSTPSIYTDAKHAANEAWPLK 342
>sptr|Q9FFX7|Q9FFX7 Alpha-mannosidase.
Length = 1030
Score = 486 bits (1250), Expect = e-136
Identities = 231/295 (78%), Positives = 257/295 (87%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYYVGSNNSIQ ACVQNVLDS++PALL D+NRKFIYVEQAFFQRWW Q++
Sbjct: 48 DVGWLKTVDQYYVGSNNSIQVACVQNVLDSIVPALLADKNRKFIYVEQAFFQRWWNEQSE 107
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
IK VK LI SG+LELINGGMCMHDEAA HYIDMIDQTTLGH++I EF PRIGWQI
Sbjct: 108 EIKRIVKELIHSGQLELINGGMCMHDEAAPHYIDMIDQTTLGHRFIIREFNVTPRIGWQI 167
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHSAVQAYLLGAEVGFD+ +F RIDYQDR+ R K LEV+WRGSK+LGSS+ IFAG
Sbjct: 168 DPFGHSAVQAYLLGAEVGFDSVFFGRIDYQDREKRYKEKTLEVIWRGSKSLGSSSQIFAG 227
Query: 543 IFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHIMF 722
FP NYEPPPG FY+E+ D+SPVVQDDP LFDYNV+ERVN FVAAA+ QANITR NHIMF
Sbjct: 228 AFPTNYEPPPGGFYYEITDDSPVVQDDPDLFDYNVQERVNAFVAAALDQANITRINHIMF 287
Query: 723 TMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAANEQGPLQ 887
TMGTDF+YQYA +W+ MDKLIHYVN DGR++A YSTPSIYTDAK+AANE PL+
Sbjct: 288 TMGTDFRYQYAHTWYRQMDKLIHYVNLDGRVNAFYSTPSIYTDAKHAANEAWPLK 342
>sptr|P94078|P94078 Alpha-mannosidase (AT3G26720/MLJ15_12).
Length = 1019
Score = 473 bits (1216), Expect = e-132
Identities = 222/295 (75%), Positives = 255/295 (86%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYYVGSNNSI+GACVQNVLDS+I +LL DENRKFIYVE AFFQRWWR Q++
Sbjct: 49 DVGWLKTVDQYYVGSNNSIRGACVQNVLDSVIASLLDDENRKFIYVEMAFFQRWWRQQSN 108
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
K VK L+ SG+LE INGGMCMHDEA HYIDMIDQTTLGH++IK EFGQ+PR+GWQI
Sbjct: 109 AKKVKVKKLVDSGQLEFINGGMCMHDEATPHYIDMIDQTTLGHQFIKTEFGQVPRVGWQI 168
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHSAVQAYLLGAE GFD+ +F RIDYQDR R K LEV+W+GSK+LGSS+ IF G
Sbjct: 169 DPFGHSAVQAYLLGAEFGFDSLFFARIDYQDRAKRLREKTLEVIWQGSKSLGSSSQIFTG 228
Query: 543 IFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHIMF 722
+FP++Y+PP G F FE++D S +QDDPLLFDYNV+ERVNDFVAAA+AQ N+TRTNHIM+
Sbjct: 229 VFPRHYDPPEG-FTFEINDVSAPIQDDPLLFDYNVQERVNDFVAAALAQVNVTRTNHIMW 287
Query: 723 TMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAANEQGPLQ 887
MGTDF+YQYA SWF +DK IHYVNKDGR++ LYSTPSIYTDAKYAANE PL+
Sbjct: 288 LMGTDFRYQYAYSWFRQIDKFIHYVNKDGRLNVLYSTPSIYTDAKYAANESWPLK 342
>sptr|Q7XGZ1|Q7XGZ1 Putative alpha-mannosidase.
Length = 640
Score = 437 bits (1123), Expect = e-121
Identities = 209/306 (68%), Positives = 246/306 (80%), Gaps = 11/306 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKT+DQY+VG+NNSIQGACV N LDS++ AL+ D RKF++ EQAFFQRWW ++
Sbjct: 49 DVGWLKTIDQYFVGTNNSIQGACVMNTLDSVVDALILDPARKFVFAEQAFFQRWWAEKSP 108
Query: 183 MIKDTVKGLISSGRLELI-----------NGGMCMHDEAAVHYIDMIDQTTLGHKYIKEE 329
I+ V L+ SG+LE I NGG CMHDEAAVHYIDMIDQTTLGH+ IK++
Sbjct: 109 KIQAIVHKLVDSGQLEFISLRKFGNFKFRNGGWCMHDEAAVHYIDMIDQTTLGHRVIKKQ 168
Query: 330 FGQIPRIGWQIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSK 509
F +IPR GWQIDPFGHSAVQ YLLGAE+GFD+ +F RIDYQDR RKG K LEV+WRGS+
Sbjct: 169 FNKIPRAGWQIDPFGHSAVQGYLLGAELGFDSMHFARIDYQDRAKRKGDKGLEVIWRGSR 228
Query: 510 TLGSSADIFAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQ 689
T GSS+ IF FP +Y PP G F FE+ D+ VQDD LLFDYN++ERVNDFVAAA+ Q
Sbjct: 229 TFGSSSQIFTNAFPVHYSPPDG-FGFEIFDDFVPVQDDMLLFDYNLKERVNDFVAAALKQ 287
Query: 690 ANITRTNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAAN 869
AN+TRTNHIM+TMG DF YQYAESWF NMD+LI+YVNKDGR+HALYSTPSIYTDAK+A+N
Sbjct: 288 ANVTRTNHIMWTMGDDFNYQYAESWFRNMDRLINYVNKDGRVHALYSTPSIYTDAKHASN 347
Query: 870 EQGPLQ 887
E PL+
Sbjct: 348 ESWPLK 353
>sptr|Q8W348|Q8W348 Putative alpha-mannosidase.
Length = 640
Score = 437 bits (1123), Expect = e-121
Identities = 209/306 (68%), Positives = 246/306 (80%), Gaps = 11/306 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKT+DQY+VG+NNSIQGACV N LDS++ AL+ D RKF++ EQAFFQRWW ++
Sbjct: 49 DVGWLKTIDQYFVGTNNSIQGACVMNTLDSVVDALILDPARKFVFAEQAFFQRWWAEKSP 108
Query: 183 MIKDTVKGLISSGRLELI-----------NGGMCMHDEAAVHYIDMIDQTTLGHKYIKEE 329
I+ V L+ SG+LE I NGG CMHDEAAVHYIDMIDQTTLGH+ IK++
Sbjct: 109 KIQAIVHKLVDSGQLEFISLRKFGNFKFRNGGWCMHDEAAVHYIDMIDQTTLGHRVIKKQ 168
Query: 330 FGQIPRIGWQIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSK 509
F +IPR GWQIDPFGHSAVQ YLLGAE+GFD+ +F RIDYQDR RKG K LEV+WRGS+
Sbjct: 169 FNKIPRAGWQIDPFGHSAVQGYLLGAELGFDSMHFARIDYQDRAKRKGDKGLEVIWRGSR 228
Query: 510 TLGSSADIFAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQ 689
T GSS+ IF FP +Y PP G F FE+ D+ VQDD LLFDYN++ERVNDFVAAA+ Q
Sbjct: 229 TFGSSSQIFTNAFPVHYSPPDG-FGFEIFDDFVPVQDDMLLFDYNLKERVNDFVAAALKQ 287
Query: 690 ANITRTNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAAN 869
AN+TRTNHIM+TMG DF YQYAESWF NMD+LI+YVNKDGR+HALYSTPSIYTDAK+A+N
Sbjct: 288 ANVTRTNHIMWTMGDDFNYQYAESWFRNMDRLINYVNKDGRVHALYSTPSIYTDAKHASN 347
Query: 870 EQGPLQ 887
E PL+
Sbjct: 348 ESWPLK 353
>sptr|Q9FKW9|Q9FKW9 Alpha-mannosidase.
Length = 1047
Score = 396 bits (1018), Expect = e-109
Identities = 184/295 (62%), Positives = 231/295 (78%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYYVGSNN IQ ACV+NVLDS++ +LL+D NRKF++ E AFF RWW Q+
Sbjct: 58 DVGWLKTVDQYYVGSNNRIQNACVRNVLDSVVDSLLRDPNRKFVFAEMAFFTRWWEEQSP 117
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
++ V+ L+ SG+LE +NGG M+DEA HYIDMIDQTT GH++IK++F PR WQI
Sbjct: 118 ERQEQVRRLVKSGQLEFVNGGWAMNDEATCHYIDMIDQTTKGHRFIKQQFNTTPRAAWQI 177
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHS+VQAYLLGAE+G D+ +F RIDYQDR+ RK K LEV+WRGSKTL SS+ IF
Sbjct: 178 DPFGHSSVQAYLLGAELGLDSVHFARIDYQDREKRKAEKSLEVIWRGSKTLDSSSQIFTN 237
Query: 543 IFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHIMF 722
IF +Y PP G F++EV D+ +QD+P YN++E VNDFV A++ AN++R NH+M+
Sbjct: 238 IFFVHYGPPTG-FHYEVTDDYVPLQDNPRFDGYNIKEAVNDFVNASLVYANVSRGNHVMW 296
Query: 723 TMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAANEQGPLQ 887
TMG DF+YQ+AESWF MD+LIHYVNKDGR++ALYSTPS+Y DAK AN PL+
Sbjct: 297 TMGDDFQYQFAESWFRQMDRLIHYVNKDGRVNALYSTPSLYVDAKNVANVTWPLK 351
>sptr|Q8VHC6|Q8VHC6 Lysosomal alpha-mannosidase (EC 3.2.1.24).
Length = 1007
Score = 285 bits (728), Expect = 8e-76
Identities = 147/302 (48%), Positives = 196/302 (64%), Gaps = 7/302 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G +N +Q A VQ +LDS+I ALL + R+F+YVE AFF RWW Q +
Sbjct: 72 DVGWLKTVDQYYWGIHNDLQQAGVQYILDSVISALLAEPTRRFVYVEMAFFSRWWHQQTN 131
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY ++DQ TLG +++++ FG PR+ W
Sbjct: 132 ETQEVVRRLVRQGRLEFANGGWVMNDEAATHYGAIVDQMTLGLRFLEDTFGSDGRPRVAW 191
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD +F RIDYQD+ RK +E+E+VWR S +L +AD+
Sbjct: 192 HIDPFGHSREQASLF-AQMGFDGVFFGRIDYQDKLVRKKRREMELVWRASASLKAPAADL 250
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P NY PP G ++V P V DDP +YN ++ V+ F+ A AQ RTNH
Sbjct: 251 FTGVLPNNYGPPEG-LCWDVLCADPPVVDDPRSPEYNAKKLVSYFLQLATAQGRYYRTNH 309
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG+DF+Y+ A +WF N+DKLI VN R+H LYSTP+ Y AN P
Sbjct: 310 TVMTMGSDFQYENANTWFKNLDKLIQLVNMQQANGSRVHVLYSTPACYLWELNKANLTWP 369
Query: 882 LQ 887
++
Sbjct: 370 VK 371
>sw|Q8VHC8|M2B1_CAVPO Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1007
Score = 285 bits (728), Expect = 8e-76
Identities = 147/302 (48%), Positives = 196/302 (64%), Gaps = 7/302 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G +N +Q A VQ +LDS+I ALL + R+F+YVE AFF RWW Q +
Sbjct: 72 DVGWLKTVDQYYWGIHNDLQQAGVQYILDSVISALLAEPTRRFVYVEMAFFSRWWHQQTN 131
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY ++DQ TLG +++++ FG PR+ W
Sbjct: 132 ETQEVVRRLVRQGRLEFANGGWVMNDEAATHYGAIVDQMTLGLRFLEDTFGSDGRPRVAW 191
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD +F RIDYQD+ RK +E+E+VWR S +L +AD+
Sbjct: 192 HIDPFGHSREQASLF-AQMGFDGVFFGRIDYQDKLVRKKRREMELVWRASASLKAPAADL 250
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P NY PP G ++V P V DDP +YN ++ V+ F+ A AQ RTNH
Sbjct: 251 FTGVLPNNYGPPEG-LCWDVLCADPPVVDDPRSPEYNAKKLVSYFLQLATAQGRYYRTNH 309
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG+DF+Y+ A +WF N+DKLI VN R+H LYSTP+ Y AN P
Sbjct: 310 TVMTMGSDFQYENANTWFKNLDKLIQLVNMQQANGSRVHVLYSTPACYLWELNKANLTWP 369
Query: 882 LQ 887
++
Sbjct: 370 VK 371
>sptrnew|AAH61819|AAH61819 Similar to mannosidase 2, alpha B1.
Length = 1009
Score = 284 bits (726), Expect = 1e-75
Identities = 142/288 (49%), Positives = 191/288 (66%), Gaps = 7/288 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G + +Q A VQ +LDS+I +LL D R+FIYVE AFF RWW+ Q +
Sbjct: 74 DVGWLKTVDQYYYGIMSDVQHASVQYILDSVIYSLLNDPTRRFIYVEMAFFSRWWKQQTN 133
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
+ +D V+ L+ GRLE +NGG M+DEAA HY ++DQ TLG +++++ FG +PR+ W
Sbjct: 134 VTQDAVRNLVRQGRLEFVNGGWVMNDEAATHYGAIVDQMTLGLRFLQDTFGSDGLPRVAW 193
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD F+ RIDYQD+ RK ++E +WR S +L +AD+
Sbjct: 194 HIDPFGHSREQASLF-AQMGFDGFFLGRIDYQDKFNRKRKLKMEELWRASASLKPPAADL 252
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P NY PP D ++V P V DDP ++N + V+ F+ A +Q RTNH
Sbjct: 253 FTGVLPNNYNPPK-DLCWDVLCTDPPVVDDPTSPEFNANKLVDYFLNLASSQKKYYRTNH 311
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVNKD----GRIHALYSTPSIY 845
+ TMG+DF+Y+ A WF NMDKLI VN+ ++H LYSTPS Y
Sbjct: 312 TVMTMGSDFQYENANMWFKNMDKLIRLVNEQQANGSKVHVLYSTPSCY 359
>sptr|Q8VHC7|Q8VHC7 Lysosomal alpha-mannosidase (EC 3.2.1.24).
Length = 1007
Score = 281 bits (720), Expect = 7e-75
Identities = 146/302 (48%), Positives = 195/302 (64%), Gaps = 7/302 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G +N +Q A VQ +LDS+I ALL + R+F+YVE AFF RWW Q +
Sbjct: 72 DVGWLKTVDQYYWGIHNDLQQAGVQYILDSVISALLAEPTRRFVYVEMAFFSRWWHQQTN 131
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY ++DQ TLG +++++ FG PR+ W
Sbjct: 132 ETQEVVRRLVRQGRLEFANGGWVMNDEAATHYGAIVDQMTLGLRFLEDTFGSDGRPRVAW 191
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD +F RIDYQD+ K +E+E+VWR S +L +AD+
Sbjct: 192 HIDPFGHSREQASLF-AQMGFDGVFFGRIDYQDKLVWKKRREMELVWRASASLKAPAADL 250
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P NY PP G ++V P V DDP +YN ++ V+ F+ A AQ RTNH
Sbjct: 251 FTGVLPNNYGPPEG-LCWDVLCADPPVVDDPRSPEYNAKKLVSYFLQLATAQGRYYRTNH 309
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG+DF+Y+ A +WF N+DKLI VN R+H LYSTP+ Y AN P
Sbjct: 310 TVMTMGSDFQYENANTWFKNLDKLIQLVNMQQANGSRVHVLYSTPACYLWELNKANLTWP 369
Query: 882 LQ 887
++
Sbjct: 370 VK 371
>sw|O00754|M2B1_HUMAN Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1010
Score = 274 bits (701), Expect = 1e-72
Identities = 142/288 (49%), Positives = 186/288 (64%), Gaps = 7/288 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G N IQ A VQ +LDS+I ALL D R+FIYVE AFF RWW Q +
Sbjct: 73 DVGWLKTVDQYFYGIKNDIQHAGVQYILDSVISALLADPTRRFIYVEIAFFSRWWHQQTN 132
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY ++DQ TLG +++++ FG PR+ W
Sbjct: 133 ATQEVVRDLVRQGRLEFANGGWVMNDEAATHYGAIVDQMTLGLRFLEDTFGNDGRPRVAW 192
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD F+F R+DYQD+ R E+E VWR S +L +AD+
Sbjct: 193 HIDPFGHSREQASLF-AQMGFDGFFFGRLDYQDKWVRMQKLEMEQVWRASTSLKPPTADL 251
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P Y PP + + + P+V +DP +YN +E V+ F+ A AQ RTNH
Sbjct: 252 FTGVLPNGYNPPRNLCWDVLCVDQPLV-EDPRSPEYNAKELVDYFLNVATAQGRYYRTNH 310
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIY 845
+ TMG+DF+Y+ A WF N+DKLI VN K +H LYSTP+ Y
Sbjct: 311 TVMTMGSDFQYENANMWFKNLDKLIRLVNAQQAKGSSVHVLYSTPACY 358
>sw|O09159|M2B1_MOUSE Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1013
Score = 272 bits (695), Expect = 5e-72
Identities = 138/288 (47%), Positives = 184/288 (63%), Gaps = 7/288 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G + +Q A VQ +LDS++ +LL+ R+FIYVE AFF RWW+ Q
Sbjct: 74 DVGWLKTVDQYYYGILSDVQHASVQYILDSVVSSLLEKPTRRFIYVEMAFFSRWWKQQTS 133
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
+D V+ L+ GRLE +NGG M+DEAA HY ++DQ TLG +++++ FG +PR+ W
Sbjct: 134 ATQDAVRNLVRQGRLEFVNGGWVMNDEAATHYGAIVDQMTLGLRFLQDTFGSDGLPRVAW 193
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD F+ RIDYQD+ RK +E +WR S +L +AD+
Sbjct: 194 HIDPFGHSREQASLF-AQMGFDGFFLGRIDYQDKLNRKKKLRMEELWRASDSLEPPAADL 252
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P NY PP ++V P V D+P ++N + VN F+ A +Q RTNH
Sbjct: 253 FTGVLPNNYNPPK-YLCWDVLCTDPPVVDNPRSPEFNAKTLVNYFLKLASSQKGFYRTNH 311
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIY 845
+ TMG+DF Y+ A WF NMDKLI VN +H LYSTP+ Y
Sbjct: 312 TVMTMGSDFHYENANMWFKNMDKLIRLVNAQQANGSLVHVLYSTPTCY 359
>sw|P34098|MANA_DICDI Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Alpha-D-mannoside mannohydrolase) (Laman).
Length = 1005
Score = 271 bits (694), Expect = 7e-72
Identities = 142/292 (48%), Positives = 188/292 (64%), Gaps = 3/292 (1%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVD+YY GSN SI A VQ LD+ I LL + RKFIYVE AFFQRWW Q+
Sbjct: 53 DVGWLKTVDEYYYGSNMSIAFAGVQYTLDTAITCLLANPERKFIYVEIAFFQRWWDEQST 112
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
+++ VKGL+ +E INGG CM+DEA +Y D IDQ TLGH+++ E FG +P+IGW I
Sbjct: 113 TMQNIVKGLVGKWSIEFINGGYCMNDEATTYYDDTIDQMTLGHQFLWENFGVMPKIGWHI 172
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHSA QA + G ++GFDAF R+DYQD + R K++E +WR +++ + +F
Sbjct: 173 DPFGHSATQARIFG-QLGFDAFIIGRMDYQDIEARLENKQMEFMWRSTQSTPEN-QVFTS 230
Query: 543 IFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHIMF 722
+ Y P G F FE D+ +QDDP LFD NV+ R F A+ A RTN+++
Sbjct: 231 VLRAMYCTPDG-FNFEQGDDP--IQDDPNLFDNNVDSRAEQFTQVALEYATHYRTNNVLI 287
Query: 723 TMGTDFKYQYAESWFGNMDKLIHYVNKDG---RIHALYSTPSIYTDAKYAAN 869
G DF Y A+ ++ N+DKLI ++N + ++ LYSTPSIY DA AN
Sbjct: 288 PFGCDFAYLNAQMYYKNIDKLIAHINSNPDKYGLNLLYSTPSIYIDAVNDAN 339
>sptr|Q8IPB7|Q8IPB7 CG6206-PB.
Length = 1080
Score = 271 bits (693), Expect = 9e-72
Identities = 139/299 (46%), Positives = 187/299 (62%), Gaps = 6/299 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY GS IQ A VQ ++DS++ ALL+D ++FIYVE AFF +WW+ Q
Sbjct: 58 DVGWLKTVDQYYYGSETKIQKAGVQYIIDSVVEALLRDPEKRFIYVESAFFFKWWKEQKP 117
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
+++ VK L+ GRLE I G M+DEA HY +IDQ + G + + + FG+ PR+GW
Sbjct: 118 KVQEAVKMLVEQGRLEFIGGAWSMNDEATTHYQSVIDQFSWGLRLLNDTFGECGRPRVGW 177
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS A + A++GFD +F R+DYQD+D R TK E++W GS LG AD+F
Sbjct: 178 QIDPFGHSREMASMF-AQMGFDGMFFGRLDYQDKDERLMTKNAEMIWHGSANLGEEADLF 236
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
+G NY+ P G F F++ N + D D NV+ERV+ F+A A RT ++
Sbjct: 237 SGALYNNYQAPDG-FCFDILCNDAPIIDGKHSPDNNVKERVDAFLAYVTEMAEHFRTPNV 295
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG DF YQ A+ W+ N+DKLI Y N+ I+ LYSTPS Y + + A P
Sbjct: 296 ILTMGEDFHYQNADMWYKNLDKLIKYGNERQANGSNINLLYSTPSCYLKSLHDAGITWP 354
>sptr|Q9VKV1|Q9VKV1 CG6206 protein (BCDNA.GH02419).
Length = 1080
Score = 271 bits (692), Expect = 1e-71
Identities = 139/299 (46%), Positives = 189/299 (63%), Gaps = 6/299 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY GS IQ A VQ ++DS++ ALL+D ++FIYVE AFF +WW+ Q
Sbjct: 58 DVGWLKTVDQYYYGSETKIQKAGVQYIIDSVVEALLRDPEKRFIYVESAFFFKWWKEQKP 117
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
+++ VK L+ GRLE I G M+DEA HY +IDQ + G + + + FG+ PR+GW
Sbjct: 118 KVQEAVKMLVEQGRLEFIGGAWSMNDEATTHYQSVIDQFSWGLRLLNDTFGECGRPRVGW 177
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS A + A++GFD +F R+DYQD+D R TK E++W GS LG AD+F
Sbjct: 178 QIDPFGHSREMASMF-AQMGFDGMFFGRLDYQDKDERLMTKNAEMIWHGSANLGEEADLF 236
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
+G NY+ P G F F++ N + D D NV+ER++ F+ A Q+ RTN+I
Sbjct: 237 SGALYNNYQAPDG-FCFDILCNDAPIIDGKHSPDNNVKERIDTFLDFAKTQSQYYRTNNI 295
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG DF YQ A+ ++ N+DKLI Y N+ I+ LYSTPS Y + + A P
Sbjct: 296 IVTMGGDFTYQAAQVYYKNLDKLIRYGNERQANGSNINLLYSTPSCYLKSLHDAGITWP 354
>sw|Q29451|M2B1_BOVIN Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 999
Score = 264 bits (675), Expect = 1e-69
Identities = 140/291 (48%), Positives = 186/291 (63%), Gaps = 10/291 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G N+IQ A VQ +LDS+I +LL + R+FIYVE AFF RWWR Q +
Sbjct: 75 DVGWLKTVDQYFYGIYNNIQPAGVQYILDSVISSLLANPTRRFIYVEIAFFSRWWRQQTN 134
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
+ V+ L+ GRLE NGG M+DEA HY +IDQ TL ++++E FG PR+ W
Sbjct: 135 ATQKIVRELVRQGRLEFANGGWVMNDEATTHYGAIIDQMTLRLRFLEETFGSDGRPRVAW 194
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD F+F R+DYQD+ RK T ++E VWR S +L +AD+
Sbjct: 195 HIDPFGHSREQASLF-AQMGFDGFFFGRLDYQDKKVRKKTLQMEQVWRASTSLKPPTADL 253
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQD--DPLLFDYNVEERVNDFVAAAVAQANITRT 707
F + P Y PP G + + + PVV+D P +YN +E V F+ A Q + RT
Sbjct: 254 FTSVLPNMYNPPEGLCWDMLCADKPVVEDTRSP---EYNAKELVRYFLKLATDQGKLYRT 310
Query: 708 NHIMFTMGTDFKYQYAESWFGNMDKLIHYVNKDG-----RIHALYSTPSIY 845
H + TMG+DF+Y+ A +WF N+DKLI VN R++ LYSTP+ Y
Sbjct: 311 KHTVMTMGSDFQYENANTWFKNLDKLIQLVNAQQRANGIRVNVLYSTPACY 361
>sptr|Q93093|Q93093 Truncated lysosomal acid alpha-mannosidase (EC
3.2.1.24).
Length = 323
Score = 262 bits (670), Expect = 4e-69
Identities = 134/269 (49%), Positives = 177/269 (65%), Gaps = 3/269 (1%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G N IQ A VQ +LDS+I ALL+D R+FIYVE AFF RWW Q +
Sbjct: 51 DVGWLKTVDQYFYGIKNDIQHAGVQYILDSVISALLEDPTRRFIYVEIAFFSRWWHQQTN 110
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY ++DQ TLG +++++ FG PR+ W
Sbjct: 111 ATQEVVRDLVRQGRLEFANGGWVMNDEAATHYGAIVDQMTLGLRFLEDTFGNDGRPRVAW 170
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD F+F R+DYQD+ R E+E VWR S +L +AD+
Sbjct: 171 HIDPFGHSREQASLF-AQMGFDGFFFGRLDYQDKWVRMQKLEMEQVWRASTSLKPPTADL 229
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
F G+ P Y PP + + + P+V +DP +YN +E V+ F+ A AQ RTNH
Sbjct: 230 FTGVLPNGYNPPRNLCWDVLCVDQPLV-EDPRSPEYNAKELVDYFLNVATAQGRYYRTNH 288
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN 800
+ TMG+DF+Y+ A WF N+DKLI VN
Sbjct: 289 TVMTMGSDFQYENANMWFKNLDKLIRLVN 317
>sw|O46432|M2B1_FELCA Lysosomal alpha-mannosidase precursor (EC
3.2.1.24) (Mannosidase, alpha B) (Lysosomal acid
alpha-mannosidase) (Laman) (Mannosidase alpha class 2B
member 1).
Length = 1007
Score = 262 bits (669), Expect = 6e-69
Identities = 138/291 (47%), Positives = 185/291 (63%), Gaps = 10/291 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G +N +Q A VQ +LDS+I +LL + R+FIYVE AFF RWW Q +
Sbjct: 75 DVGWLKTVDQYFYGIHNDVQHAGVQYILDSVISSLLVEPTRRFIYVEIAFFSRWWHQQTN 134
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQ--IPRIGW 356
++ V+ L+ GRLE NGG M+DEAA HY +IDQ TLG +++++ FG+ PR+ W
Sbjct: 135 ATQEVVRDLVRQGRLEFANGGWVMNDEAATHYGAIIDQMTLGLRFLEDTFGKDGRPRVAW 194
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTL-GSSADI 533
IDPFGHS QA L A++GFD +F R+DYQD+ R+ LE VWR S +L +AD+
Sbjct: 195 HIDPFGHSREQASLF-AQMGFDGLFFGRLDYQDKRVREENLGLEQVWRASASLKPPAADL 253
Query: 534 FAGIFPKNYEPPPGDFYFEVDDNSPVVQD--DPLLFDYNVEERVNDFVAAAVAQANITRT 707
F + P Y PP + + + P V+D P +YN EE VN F+ A AQ RT
Sbjct: 254 FTSVLPNIYNPPEKLCWDTLCADKPFVEDRRSP---EYNAEELVNYFLQLATAQGQHFRT 310
Query: 708 NHIMFTMGTDFKYQYAESWFGNMDKLIHYVN-----KDGRIHALYSTPSIY 845
NH + TMG+DF+Y+ A WF N+D+LI VN R++ LYSTP+ Y
Sbjct: 311 NHTIMTMGSDFQYENANMWFRNLDRLIQLVNAQQQANGSRVNVLYSTPACY 361
>sptr|Q9VLI0|Q9VLI0 CG9466 protein (GH02475P).
Length = 982
Score = 251 bits (641), Expect = 1e-65
Identities = 134/300 (44%), Positives = 194/300 (64%), Gaps = 7/300 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY G N+IQ A VQ ++D++I L+K+ +R+FI VE +FF +WW Q++
Sbjct: 48 DVGWLKTVDQYYYGHRNNIQHAGVQYIIDTVISELIKNPDRRFIQVETSFFSKWWNEQSE 107
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
++ VK L++ GRL+ ING M+DEAAV+Y +IDQ T+G K++ + FG PR+GW
Sbjct: 108 TMRAVVKMLVNEGRLQFINGAWSMNDEAAVNYQSVIDQFTVGLKFLDDTFGVCGRPRVGW 167
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS QA + A++GFD +F R+D+ D+ R LE++W S++L S +F
Sbjct: 168 QIDPFGHSREQASIY-AQMGFDGEFFSRMDHNDKGRRMNDLALEMIWDASESL-SQVKLF 225
Query: 537 AGIFPKNYEPPPGDFYFEVD-DNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
G+ Y PG F F+V + P++ D +D NV+ RV+DF+A A A RTNH
Sbjct: 226 TGLLYTFYWDTPG-FCFDVHCSDDPIIDSDS--YDNNVKSRVDDFIAYAAQVAEKFRTNH 282
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANEQGP 881
IM MG DF+Y+ AE + NMDKLI YVN+ + + +YSTP+ Y ++ + + + P
Sbjct: 283 IMIPMGGDFQYEDAEVNYKNMDKLIKYVNERQSSGSKYNIIYSTPTCYLNSVHKSVQSYP 342
>sptr|O97357|O97357 Lysosomal acid alpha-mannosidase precursor.
Length = 977
Score = 250 bits (639), Expect = 2e-65
Identities = 132/301 (43%), Positives = 176/301 (58%), Gaps = 9/301 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
D+GWLKTV+QY G NN+IQ A V ++ S+I LL + RKF YVE FF RWW+ Q +
Sbjct: 36 DMGWLKTVEQYQYGLNNTIQVADVNGIISSVIAGLLLNPRRKFTYVEIGFFSRWWKEQGE 95
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQIPRIGWQI 362
+++TV+ L++ G+L+ NGG CMHDEA H+IDMIDQTT+GH+++ E +PR+GWQ+
Sbjct: 96 EMRNTVRNLVAKGQLQFANGGWCMHDEATTHFIDMIDQTTIGHRWLWRELKVVPRVGWQV 155
Query: 363 DPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIFAG 542
DPFGHSA QA +L A G +F R+DY+D + R T + W+ S +L FA
Sbjct: 156 DPFGHSATQAAMLTARAGLVGTFFARVDYRDFEYRASTGRRQFWWQPSPSL-PELQTFAE 214
Query: 543 IFPKNYEPPPGDFYFEVDD--NSPVVQDDPLLF-------DYNVEERVNDFVAAAVAQAN 695
I PP F ++V D S V DPL F +YN+ + F
Sbjct: 215 INLHQTYCPPSKFSWDVVDYWMSAVRNTDPLNFVEDKSSENYNIPFILELFKEEVRRNVE 274
Query: 696 ITRTNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNKDGRIHALYSTPSIYTDAKYAANEQ 875
TR +IM+TMG DF Y +E WFG MD+LI VN DG YSTP Y AK + +
Sbjct: 275 QTRGKNIMWTMGCDFNYFASELWFGKMDRLIEIVNADGEFEVRYSTPYEYAMAKREEHSK 334
Query: 876 G 878
G
Sbjct: 335 G 335
>sptr|Q7Q8J8|Q7Q8J8 AgCP15268 (Fragment).
Length = 694
Score = 249 bits (636), Expect = 4e-65
Identities = 134/299 (44%), Positives = 184/299 (61%), Gaps = 6/299 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY GS ++IQ A VQ +LDS+I LLKD R+FI+VE AF +W+ Q++
Sbjct: 21 DVGWLKTVDQYYYGSRSTIQKAGVQYILDSVIQELLKDAARRFIFVESAFMHKWYTEQSE 80
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
++ TV+ L+S GRLE I G M+DEAAVHY +IDQ T G +++ + FG+ P GW
Sbjct: 81 EMRQTVRDLVSEGRLEFIGGAWSMNDEAAVHYQSVIDQFTWGLRFLNDTFGECGRPMAGW 140
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS Q L ++G+D + R+D+QD+ R + +E++W+ S +L S
Sbjct: 141 QIDPFGHSREQGSLF-VQMGYDGLFQGRLDFQDKSNRIAQRTMEMMWKTSDSLEQSELFT 199
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
+G++ Y PPPG F F+V N + D P D NV+E+V+ F+ + TN+I
Sbjct: 200 SGLY-YIYAPPPG-FCFDVLCNDDPIIDGPYSADNNVKEKVDLFLDTVHNMSKTFLTNNI 257
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
+ TMG DF YQ A WF N+DKLI Y N + ++ YSTPS Y A AN P
Sbjct: 258 ILTMGEDFHYQDANMWFKNLDKLIRYTNQRQSEGSMVNVFYSTPSCYLKAVNEANVHWP 316
>sptr|Q7Q8J7|Q7Q8J7 AgCP15371 (Fragment).
Length = 1175
Score = 248 bits (633), Expect = 8e-65
Identities = 134/290 (46%), Positives = 176/290 (60%), Gaps = 6/290 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY GS +IQ A VQ +LDS+I +LL D RKFIYVE AFF +W+ Q
Sbjct: 167 DVGWLKTVDQYYYGSKTTIQKAGVQYILDSVIQSLLSDPERKFIYVESAFFFKWYDEQTA 226
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
++ V+ L++ GRLE I G M+DEAA HY ++DQ T G + + FG+ PRIGW
Sbjct: 227 ELQQQVRMLVNEGRLEFIGGAWSMNDEAAAHYHSIVDQFTWGLAKLNDTFGECGRPRIGW 286
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS QA L A++G+D +F R+DYQD+ R K E++W+ S L + D+F
Sbjct: 287 QIDPFGHSREQASLF-AQMGYDGLFFGRLDYQDKRERMTHKRAEMIWKTSDNL-ADGDLF 344
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
G+ Y+ PPG F F++ + D P + NV+ +V F+ QA RTN+I
Sbjct: 345 TGVLYNLYQAPPG-FCFDILCSDEPFMDSPYSAENNVKAKVEKFLYYVNLQAESYRTNNI 403
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDA 854
+ TMG DF Y A +F NMDKLI Y N ++ YSTPS Y A
Sbjct: 404 LLTMGGDFTYMDANVYFKNMDKLIKYTNALQSNGSNVNVFYSTPSCYLKA 453
>sptr|Q9VLH9|Q9VLH9 CG9468 protein.
Length = 1007
Score = 244 bits (624), Expect = 9e-64
Identities = 130/300 (43%), Positives = 193/300 (64%), Gaps = 7/300 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G ++IQ A VQ ++D++I L+KD R+FI VE +FF +WW Q++
Sbjct: 49 DVGWLKTVDQYFYGHRSNIQHAGVQYIIDTVIAELIKDPARRFIQVETSFFAKWWAEQSE 108
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
K V+ L++ GRLE G M+DEAAV+Y +IDQ TLG K++ + FG PRIGW
Sbjct: 109 TAKQVVRKLVNEGRLEFTGGAWSMNDEAAVNYQSVIDQFTLGLKFLDDTFGSCARPRIGW 168
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS QA + A++G+D +F R+D++D++ R +E++W S +L S+ +IF
Sbjct: 169 QIDPFGHSREQASIF-AQMGYDGEFFARMDHRDKNDRIDNLGMEMIWDASDSL-SNDEIF 226
Query: 537 AGIFPKNYEPPPGDFYFEVD-DNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
G+ ++Y PPG + F+V + P++ D +D NV+ RV+DF++ + A R+NH
Sbjct: 227 TGLLYRHYSAPPG-YCFDVHCGDDPII--DTKSYDNNVKSRVDDFLSYVTSVAQHYRSNH 283
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
IM MG DF+Y+ A+ F NMDKLI YVN + + YSTP+ Y ++ + + P
Sbjct: 284 IMIPMGDDFQYEDAQVNFKNMDKLIKYVNERQAEGSTFNLFYSTPACYLNSLHEGLQTWP 343
>sptr|Q8MS44|Q8MS44 RE08556p.
Length = 1007
Score = 244 bits (624), Expect = 9e-64
Identities = 130/300 (43%), Positives = 193/300 (64%), Gaps = 7/300 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQY+ G ++IQ A VQ ++D++I L+KD R+FI VE +FF +WW Q++
Sbjct: 49 DVGWLKTVDQYFYGHRSNIQHAGVQYIIDTVIAELIKDPARRFIQVETSFFAKWWAEQSE 108
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
K V+ L++ GRLE G M+DEAAV+Y +IDQ TLG K++ + FG PRIGW
Sbjct: 109 TAKQVVRKLVNEGRLEFTGGAWSMNDEAAVNYQSVIDQFTLGLKFLDDTFGSCARPRIGW 168
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS QA + A++G+D +F R+D++D++ R +E++W S +L S+ +IF
Sbjct: 169 QIDPFGHSREQASIF-AQMGYDGEFFARMDHRDKNDRIDNLGMEMIWDASDSL-SNDEIF 226
Query: 537 AGIFPKNYEPPPGDFYFEVD-DNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
G+ ++Y PPG + F+V + P++ D +D NV+ RV+DF++ + A R+NH
Sbjct: 227 TGLLYRHYSAPPG-YCFDVHCGDDPII--DTKSYDNNVKSRVDDFLSYVTSVAQHYRSNH 283
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIYTDAKYAANEQGP 881
IM MG DF+Y+ A+ F NMDKLI YVN + + YSTP+ Y ++ + + P
Sbjct: 284 IMIPMGDDFQYEDAQVNFKNMDKLIKYVNERQAEGSTFNLFYSTPACYLNSLHEGLQTWP 343
>sptr|Q9VKV2|Q9VKV2 CG5322 protein.
Length = 950
Score = 240 bits (613), Expect = 2e-62
Identities = 126/288 (43%), Positives = 184/288 (63%), Gaps = 7/288 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQYY GS N IQ A VQ +LD+++ LLKD +R+FI VE FF +W+ Q +
Sbjct: 13 DVGWLKTVDQYYYGSQNKIQHAGVQYILDTVVEELLKDSSRRFIQVETFFFAKWYSEQTE 72
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
++ VK L++ GRLE G M+DEA VHY ++DQ LG +Y+K+ FG P +GW
Sbjct: 73 TVQRAVKKLVAQGRLEFAGGAWSMNDEATVHYQSVVDQFNLGLRYLKDTFGDCGRPTVGW 132
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS A + A++ F+ +F R+DY D+ R E+E++W+ S++L +S +IF
Sbjct: 133 QIDPFGHSREMASIF-AQMAFNGEFFARMDYVDKKQRMLDLEMEMIWQSSESLKNS-NIF 190
Query: 537 AGIFPKNYEPPPGDFYFEVD-DNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNH 713
G+ +Y PPG F F+++ +++P++ + +D NV+ RV++F+ + R+ H
Sbjct: 191 TGMLYNHYSAPPG-FCFDINCEDAPIIDGES--YDNNVDARVSEFIDYVKNMSKSYRSTH 247
Query: 714 IMFTMGTDFKYQYAESWFGNMDKLIHYVN----KDGRIHALYSTPSIY 845
IM MG DF+Y+ A F NMDKLI YVN +++ YSTPS Y
Sbjct: 248 IMVPMGDDFQYEDAAVNFKNMDKLIKYVNDRQSTGSQVNVFYSTPSCY 295
>sptr|Q9VLI1|Q9VLI1 CG9465 protein.
Length = 942
Score = 233 bits (595), Expect = 2e-60
Identities = 124/299 (41%), Positives = 183/299 (61%), Gaps = 6/299 (2%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
DVGWLKTVDQ Y G N I A VQ +LDS++ LL+D R+FI VE AFF +WW Q++
Sbjct: 13 DVGWLKTVDQSYYGYKNKIHHAGVQYILDSVVSQLLQDPKRRFIQVETAFFAKWWNVQSE 72
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
+K V L++ GR E G M+DEA V+Y +IDQ T+G K++ E FG PR+GW
Sbjct: 73 SMKKVVTKLVNEGRFEFTGGAWSMNDEANVNYQGVIDQFTVGLKFLNETFGSCGRPRVGW 132
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QID FGHS QA + A +G+D +F R+D+ D+DTR +E++W S+++ S +F
Sbjct: 133 QIDTFGHSREQAAIF-AHMGYDGQFFSRMDHNDKDTRMNDLTMEMIWDASESM-SGVSLF 190
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
+G+ Y PG + + + P++ D + NV++RV+DF++ A + + TNHI
Sbjct: 191 SGMLYNFYSDTPGFCFDILCTDQPIIDGDS--EENNVKQRVDDFISYAKNVSEVYITNHI 248
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANEQGP 881
M MG DF+Y+ A+ + NMDKLI Y+N+ + + YSTPS Y ++ + + E P
Sbjct: 249 MVPMGGDFQYEDAKVNYKNMDKLIKYINERQASGSKYNIFYSTPSCYLNSLHQSLESWP 307
>sptrnew|AAL48871|AAL48871 RE28991p (Fragment).
Length = 1016
Score = 233 bits (595), Expect = 2e-60
Identities = 129/303 (42%), Positives = 183/303 (60%), Gaps = 10/303 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
D+GWLKTVDQYY G+ ++IQ A VQ + D+++ L KD R+FI VE FF +WW +Q++
Sbjct: 61 DLGWLKTVDQYYYGNKHNIQHAGVQYIFDTVVAELSKDSRRRFIQVETGFFAKWWEDQSE 120
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
K VK L++ GRLE G M+DEAAV+Y +IDQ +G K++ + FG PR+GW
Sbjct: 121 TKKKLVKTLVNEGRLEFTGGAWSMNDEAAVNYQSVIDQFAVGLKFLNDNFGVCGRPRVGW 180
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QID FGHS QA + A++ FD +F R+D D++ R LE+VW S+TL D F
Sbjct: 181 QIDAFGHSREQASMF-AQMAFDGQFFARMDQNDKNKRVDNLGLEMVWDASETL-EEMDFF 238
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLL----FDYNVEERVNDFVAAAVAQANITR 704
G+ +Y PPG F F+ + QDDP++ +D NV+ RV+DF+ A A R
Sbjct: 239 TGMLYNHYSAPPG-FCFD-----SLCQDDPIIDGKSYDNNVKSRVDDFLNYASNLAGYYR 292
Query: 705 TNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANE 872
+NHIM MG DF+Y+ A + NMDKLI YVN+ + + YST Y ++ + + +
Sbjct: 293 SNHIMVPMGDDFQYENAYMNYKNMDKLIKYVNERQASGSKYNIFYSTAGCYLNSLHKSLQ 352
Query: 873 QGP 881
P
Sbjct: 353 SFP 355
>sptr|Q8SYY1|Q8SYY1 RE28991p.
Length = 1003
Score = 233 bits (595), Expect = 2e-60
Identities = 129/303 (42%), Positives = 183/303 (60%), Gaps = 10/303 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
D+GWLKTVDQYY G+ ++IQ A VQ + D+++ L KD R+FI VE FF +WW +Q++
Sbjct: 48 DLGWLKTVDQYYYGNKHNIQHAGVQYIFDTVVAELSKDSRRRFIQVETGFFAKWWEDQSE 107
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
K VK L++ GRLE G M+DEAAV+Y +IDQ +G K++ + FG PR+GW
Sbjct: 108 TKKKLVKTLVNEGRLEFTGGAWSMNDEAAVNYQSVIDQFAVGLKFLNDNFGVCGRPRVGW 167
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QID FGHS QA + A++ FD +F R+D D++ R LE+VW S+TL D F
Sbjct: 168 QIDAFGHSREQASMF-AQMAFDGQFFARMDQNDKNKRVDNLGLEMVWDASETL-EEMDFF 225
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLL----FDYNVEERVNDFVAAAVAQANITR 704
G+ +Y PPG F F+ + QDDP++ +D NV+ RV+DF+ A A R
Sbjct: 226 TGMLYNHYSAPPG-FCFD-----SLCQDDPIIDGKSYDNNVKSRVDDFLNYASNLAGYYR 279
Query: 705 TNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANE 872
+NHIM MG DF+Y+ A + NMDKLI YVN+ + + YST Y ++ + + +
Sbjct: 280 SNHIMVPMGDDFQYENAYMNYKNMDKLIKYVNERQASGSKYNIFYSTAGCYLNSLHKSLQ 339
Query: 873 QGP 881
P
Sbjct: 340 SFP 342
>sptr|Q9VLI2|Q9VLI2 CG9463 protein.
Length = 1003
Score = 233 bits (595), Expect = 2e-60
Identities = 129/303 (42%), Positives = 183/303 (60%), Gaps = 10/303 (3%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
D+GWLKTVDQYY G+ ++IQ A VQ + D+++ L KD R+FI VE FF +WW +Q++
Sbjct: 48 DLGWLKTVDQYYYGNKHNIQHAGVQYIFDTVVAELSKDSRRRFIQVETGFFAKWWEDQSE 107
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
K VK L++ GRLE G M+DEAAV+Y +IDQ +G K++ + FG PR+GW
Sbjct: 108 TKKKLVKTLVNEGRLEFTGGAWSMNDEAAVNYQSVIDQFAVGLKFLNDNFGVCGRPRVGW 167
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QID FGHS QA + A++ FD +F R+D D++ R LE+VW S+TL D F
Sbjct: 168 QIDAFGHSREQASMF-AQMAFDGQFFARMDQNDKNKRVDNLGLEMVWDASETL-EEMDFF 225
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLL----FDYNVEERVNDFVAAAVAQANITR 704
G+ +Y PPG F F+ + QDDP++ +D NV+ RV+DF+ A A R
Sbjct: 226 TGMLYNHYSAPPG-FCFD-----SLCQDDPIIDGKSYDNNVKSRVDDFLNYASNLAGYYR 279
Query: 705 TNHIMFTMGTDFKYQYAESWFGNMDKLIHYVNK----DGRIHALYSTPSIYTDAKYAANE 872
+NHIM MG DF+Y+ A + NMDKLI YVN+ + + YST Y ++ + + +
Sbjct: 280 SNHIMVPMGDDFQYENAYMNYKNMDKLIKYVNERQASGSKYNIFYSTAGCYLNSLHKSLQ 339
Query: 873 QGP 881
P
Sbjct: 340 SFP 342
>sptr|Q20829|Q20829 Hypothetical protein.
Length = 955
Score = 215 bits (548), Expect = 6e-55
Identities = 113/291 (38%), Positives = 173/291 (59%), Gaps = 4/291 (1%)
Frame = +3
Query: 3 DVGWLKTVDQYYVGSNNSIQGACVQNVLDSLIPALLKDENRKFIYVEQAFFQRWWRNQND 182
D+GW+KTVDQY+ G+ + VQ + +++I LLK+ +R+F + E F RW+ + +D
Sbjct: 47 DLGWIKTVDQYFWGAKPELVPVGVQYIYNTVIDELLKNPDRRFSFAETGFLWRWYTSNSD 106
Query: 183 MIKDTVKGLISSGRLELINGGMCMHDEAAVHYIDMIDQTTLGHKYIKEEFGQI--PRIGW 356
+ ++ L+ +G++E+I GG +DEA HY+D+IDQ TLG + +++ FG+ P GW
Sbjct: 107 FDRHQLQKLVKNGQIEIIGGGWVQNDEATSHYVDIIDQMTLGLQRLEQIFGECGKPVTGW 166
Query: 357 QIDPFGHSAVQAYLLGAEVGFDAFYFFRIDYQDRDTRKGTKELEVVWRGSKTLGSSADIF 536
QIDPFGHS A + E+G+ + YF RI Y ++ R K LE +W S + +
Sbjct: 167 QIDPFGHSREMANIY-REMGYSSVYFARIHYLEKQIRLKNKTLEFMWNTSDDITDNKLFT 225
Query: 537 AGIFPKNYEPPPGDFYFEVDDNSPVVQDDPLLFDYNVEERVNDFVAAAVAQANITRTNHI 716
F NY PP G + + + P++ D+ + YNV+E+V+ FV QA TN +
Sbjct: 226 GAFFNDNYGPPEGFCWDSLCGDDPIM-DNLNIEGYNVKEKVDAFVDHVKNQAAHQSTNQV 284
Query: 717 MFTMGTDFKYQYAESWFGNMDKLIHYVNKD--GRIHALYSTPSIYTDAKYA 863
M MG+DF+Y A SW+ N+DKLI YVN D ++ +YSTP+ YT A A
Sbjct: 285 MLLMGSDFQYTNANSWYVNLDKLIKYVNADTSKKVRVIYSTPACYTKAVQA 335
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 783,133,803
Number of Sequences: 1381838
Number of extensions: 18206021
Number of successful extensions: 51995
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 48378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 51816
length of database: 439,479,560
effective HSP length: 122
effective length of database: 270,895,324
effective search space used: 46864891052
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

