BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= 4005867.2.1
(1967 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
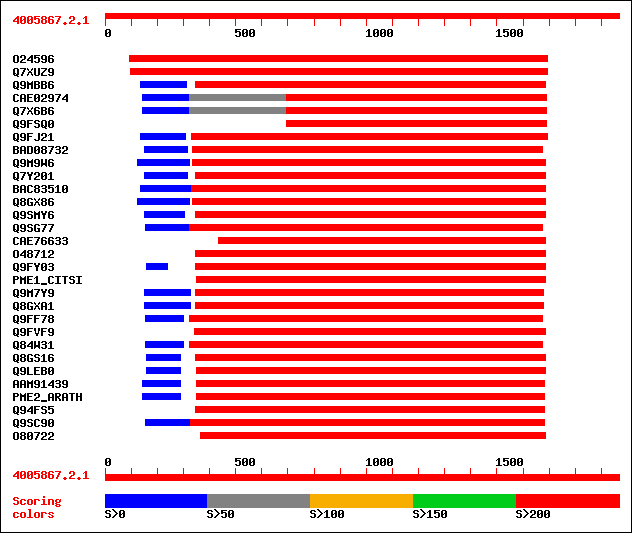
sptr|O24596|O24596 Pectin methylesterase-like protein. 1029 0.0
sptr|Q7XUZ9|Q7XUZ9 OSJNBa0036B21.20 protein. 824 0.0
sptr|Q9MBB6|Q9MBB6 Pectin methylesterase. 493 e-141
sptrnew|CAE02974|CAE02974 OSJNBb0079B02.7 protein. 466 e-130
sptr|Q7X6B6|Q7X6B6 OSJNBb0079B02.8 protein (OSJNBa0063C18.2 prot... 466 e-130
sptr|Q9FSQ0|Q9FSQ0 Putative pectin methylesterase. 466 e-130
sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11) (AT5g4918... 408 e-117
sptrnew|BAD08732|BAD08732 Putative pectinesterase. 405 e-117
sptr|Q9M9W6|Q9M9W6 Putative pectinesterase. 388 e-114
sptr|Q7Y201|Q7Y201 Putative pectin methylesterase. 397 e-113
sptrnew|BAC83510|BAC83510 Putative pectin methylesterase. 383 e-113
sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610). 384 e-112
sptr|Q9SMY6|Q9SMY6 Pectinesterase-like protein. 398 e-112
sptr|Q9SG77|Q9SG77 Putative pectinesterase. 373 e-109
sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11) (F... 394 e-108
sptr|O48712|O48712 Putative pectinesterase. 394 e-108
sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC ... 390 e-107
sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11) ... 390 e-107
sptr|Q9M7Y9|Q9M7Y9 Pectin methylesterase, putative (EC 3.1.1.11)... 369 e-105
sptr|Q8GXA1|Q8GXA1 Putative pectin methylesterase. 369 e-105
sptr|Q9FF78|Q9FF78 Pectinesterase. 368 e-105
sptr|Q9FVF9|Q9FVF9 Putative pectin methylesterase 3 (EC 3.1.1.11... 381 e-104
sptr|Q84W31|Q84W31 Hypothetical protein. 364 e-104
sptr|Q8GS16|Q8GS16 Putative thermostable pectinesterase. 374 e-104
sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11) (Pectines... 377 e-104
sptrnew|AAM91439|AAM91439 At1g53830/T18A20_6. 372 e-103
sw|Q42534|PME2_ARATH Pectinesterase 2 precursor (EC 3.1.1.11) (P... 372 e-103
sptr|Q94FS5|Q94FS5 Putative pectin methylesterase LuPME5 (EC 3.1... 377 e-103
sptr|Q9SC90|Q9SC90 Pectin methyl-esterase PER precursor (EC 3.1.... 370 e-103
sptr|O80722|O80722 Putative pectinesterase. 375 e-102
>sptr|O24596|O24596 Pectin methylesterase-like protein.
Length = 563
Score = 1029 bits (2660), Expect = 0.0
Identities = 508/534 (95%), Positives = 508/534 (95%)
Frame = +1
Query: 94 KKAGDNFTVPGEASLANVRQVGQVPVRAHPVQGVVREDVVPGHQWHREXXXXXXXXXXXX 273
KKAGDNFTVPGEASLANVRQVGQVPVRAHPVQGVVREDVVPGHQWHRE
Sbjct: 30 KKAGDNFTVPGEASLANVRQVGQVPVRAHPVQGVVREDVVPGHQWHREPQGGVPQRGQGG 89
Query: 274 XXXXXXXXXXXQVDRRGQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRS 453
QVDRRGQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRS
Sbjct: 90 AGVGPDGGRAVQVDRRGQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRS 149
Query: 454 DDLETWLTGVMTFMDTCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDL 633
DDLETWLTGVMTFMDTCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDL
Sbjct: 150 DDLETWLTGVMTFMDTCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDL 209
Query: 634 GMFSKDSRRRLLSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAV 813
GMFSKDSRRRLLSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAV
Sbjct: 210 GMFSKDSRRRLLSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAV 269
Query: 814 DAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKT 993
DAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKT
Sbjct: 270 DAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKT 329
Query: 994 ATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQF 1173
ATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYV RRQF
Sbjct: 330 ATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVQPRRQF 389
Query: 1174 FRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRL 1353
FRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHG TDPNMKSGLVIQNCRL
Sbjct: 390 FRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGPTDPNMKSGLVIQNCRL 449
Query: 1354 VPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEY 1533
VPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEY
Sbjct: 450 VPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEY 509
Query: 1534 NNRGPGAGTSKRVNWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILGFKF 1695
NNRGPGAGTSKRVNWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILGFKF
Sbjct: 510 NNRGPGAGTSKRVNWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILGFKF 563
>sptr|Q7XUZ9|Q7XUZ9 OSJNBa0036B21.20 protein.
Length = 568
Score = 824 bits (2128), Expect = 0.0
Identities = 404/536 (75%), Positives = 449/536 (83%), Gaps = 4/536 (0%)
Frame = +1
Query: 97 KAGDNFTVPGEASLANVRQVGQVPVRAHPVQGVVREDVVPGHQWHREXXXXXXXXXXXXX 276
KAGDNFTVPGEA+LA + + + + +
Sbjct: 32 KAGDNFTVPGEANLATSGKSVESLCAPTLYKESCEKTLTTATSGTENPKEVFSTVAKSAL 91
Query: 277 XXXXXXXXXXQVDRRGQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSD 456
+ + SDSMTESAREDCK LLED+ DDLRGM+EMAGGD+KVLFSRSD
Sbjct: 92 ESIKSAVEKSKAIGEAKTSDSMTESAREDCKALLEDSVDDLRGMVEMAGGDVKVLFSRSD 151
Query: 457 DLETWLTGVMTFMDTCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLG 636
DLE WLTGVMTFMDTC DGF DEKLKADMHSV+RNA+ELSSNALAITN+LG I KK+DL
Sbjct: 152 DLEHWLTGVMTFMDTCADGFADEKLKADMHSVLRNASELSSNALAITNTLGAIFKKLDLD 211
Query: 637 MFSKDS--RRRLLSSEQDEKGWPVWMRSPERKLLASG--NQPKPNAIVAKDGSGQFKSIQ 804
MF ++ R L++ ++ G+P WM++P+RKLLASG N+P+PNA+VA+DGSGQFK+IQ
Sbjct: 212 MFKGENPIHRSLIAEQETVGGFPSWMKAPDRKLLASGDRNRPQPNAVVAQDGSGQFKTIQ 271
Query: 805 QAVDAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITT 984
+AV+++PKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPK+SRVTGRKSFADGITT
Sbjct: 272 EAVNSMPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKRSRVTGRKSFADGITT 331
Query: 985 MKTATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHAR 1164
MKTATFSVEA+GFICKNMGFHNTAGAERHQAVALR+ GDL AFYNCRFDAFQDTLYVHAR
Sbjct: 332 MKTATFSVEAAGFICKNMGFHNTAGAERHQAVALRINGDLGAFYNCRFDAFQDTLYVHAR 391
Query: 1165 RQFFRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQN 1344
RQFFRNCV+SGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQN
Sbjct: 392 RQFFRNCVISGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQN 451
Query: 1345 CRLVPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYY 1524
CRLVPDQKLFPDRFKIPSYLGRPWKE+SRLVIMESTIADF+KPEGYMPWNG+FAL TLYY
Sbjct: 452 CRLVPDQKLFPDRFKIPSYLGRPWKEYSRLVIMESTIADFIKPEGYMPWNGEFALNTLYY 511
Query: 1525 AEYNNRGPGAGTSKRVNWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILGFK 1692
AE+NNRGPGAGTSKRVNW GF VIG+KEAE FTAGPF+DG WLK+TG PH LGFK
Sbjct: 512 AEFNNRGPGAGTSKRVNWKGFRVIGQKEAEQFTAGPFVDGGTWLKFTGTPHFLGFK 567
Score = 105 bits (261), Expect = 3e-21
Identities = 52/68 (76%), Positives = 61/68 (89%)
Frame = +2
Query: 125 GRLPLPTSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQ 304
G L TSGKSV+SLCAPTLYKESCEKTL+ AT+GTENPKEVF +VAK ALES+++AVE+
Sbjct: 41 GEANLATSGKSVESLCAPTLYKESCEKTLTTATSGTENPKEVFSTVAKSALESIKSAVEK 100
Query: 305 SKSIGEAK 328
SK+IGEAK
Sbjct: 101 SKAIGEAK 108
>sptr|Q9MBB6|Q9MBB6 Pectin methylesterase.
Length = 596
Score = 493 bits (1270), Expect(2) = e-141
Identities = 240/454 (52%), Positives = 323/454 (71%), Gaps = 7/454 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGFVDE 525
++A +DC +L+EDA ++L+ ++ G DI L S + DL WL+ VM++ TC+DGF +
Sbjct: 146 KAAFDDCLELIEDAKEELKNSVDCIGNDIGKLASNAPDLSNWLSAVMSYQQTCIDGFPEG 205
Query: 526 KLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQ-----DEK 690
KLK+DM + EL+SN+LA+ +SL LK FS RRLL+ EQ D+
Sbjct: 206 KLKSDMEKTFKATRELTSNSLAMVSSLVSFLKNFS---FSGTLNRRLLAEEQNSPSLDKD 262
Query: 691 GWPVWMRSPERKLL--ASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKA 864
G P WM +R++L A ++PKPN VAKDGSG FK+I +A+ A+P ++GRYVI+VK
Sbjct: 263 GVPGWMSHEDRRILKGADKDKPKPNVSVAKDGSGDFKTISEALAAMPAKYEGRYVIFVKQ 322
Query: 865 GLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGF 1044
G+YDE V V K NI MYGDG +++ VTG K+FADG+ T +TATF+V GF+CK MGF
Sbjct: 323 GVYDETVTVTKKMANITMYGDGSQKTIVTGNKNFADGVQTFRTATFAVLGDGFLCKFMGF 382
Query: 1045 HNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNS 1224
NTAG E+HQAVA+RVQ D A F NCRF+ +QDTLY RQF+R+CV++GT+DFIFG++
Sbjct: 383 RNTAGPEKHQAVAIRVQADRAIFLNCRFEGYQDTLYAQTHRQFYRSCVITGTVDFIFGDA 442
Query: 1225 AAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYL 1404
+VFQNCLI R+P++NQQN VTA GR D + +G+V+Q+CR+ PD+ L P + KI SYL
Sbjct: 443 TSVFQNCLITVRKPLENQQNIVTAQGRIDGHETTGIVLQSCRIEPDKDLVPVKNKIRSYL 502
Query: 1405 GRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPG 1584
GRPWKEFSR VIM+STI DF+ P G++PW GDF LKTLYYAEY+N+G GA T+ R+ WPG
Sbjct: 503 GRPWKEFSRTVIMDSTIGDFIHPGGWLPWQGDFGLKTLYYAEYSNKGGGAQTNARIKWPG 562
Query: 1585 FHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
+H+I ++EA FT F G W+ +G+P LG
Sbjct: 563 YHIIKKEEAMKFTIENFYQGD-WISASGSPVHLG 595
Score = 34.7 bits (78), Expect(2) = e-141
Identities = 15/64 (23%), Positives = 33/64 (51%), Gaps = 4/64 (6%)
Frame = +2
Query: 134 PLPTSGKSVKSLCAPTLYKESCEKTLSQATNGTE----NPKEVFHSVAKVALESVQTAVE 301
P+ + +K++C T Y+++C+ TL + G + PK++ K A E + ++
Sbjct: 74 PISHVARVIKTVCNATTYQDTCQNTLEKGVLGKDPSSVQPKDLLKIAIKAADEEIDKVIK 133
Query: 302 QSKS 313
++ S
Sbjct: 134 KASS 137
>sptrnew|CAE02974|CAE02974 OSJNBb0079B02.7 protein.
Length = 971
Score = 466 bits (1198), Expect = e-130
Identities = 216/332 (65%), Positives = 264/332 (79%)
Frame = +1
Query: 694 WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLY 873
+P W+ + +R+LL +G Q KP+ +VAKDGSG FK+I +AV+AVPK R+VIYVKAG Y
Sbjct: 639 FPSWVSAHQRRLLQAGTQ-KPDKVVAKDGSGDFKTITEAVNAVPKNSPTRFVIYVKAGEY 697
Query: 874 DEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFHNT 1053
+E V +P NIFMYGDGP ++RV G KS DG+ TM T TFS E +GF+CK+MGF NT
Sbjct: 698 NEYVTIPSSLPNIFMYGDGPTKTRVLGNKSNKDGVATMATRTFSAEGNGFVCKSMGFVNT 757
Query: 1054 AGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAV 1233
AG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC V+GTID+IFGNSAAV
Sbjct: 758 AGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIFGNSAAV 817
Query: 1234 FQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRP 1413
FQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I SYLGRP
Sbjct: 818 FQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIASYLGRP 877
Query: 1414 WKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHV 1593
WKE++R V+MES I DF+KPEG+ W GD LKTLYYAEY N GPGAGTSKRV WPG+ V
Sbjct: 878 WKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVTWPGYRV 937
Query: 1594 IGRKEAEPFTAGPFIDGAMWLKYTGAPHILGF 1689
IG+ EA FTAG FIDG WLK T P+++GF
Sbjct: 938 IGQAEATQFTAGVFIDGLTWLKNTATPNVMGF 969
Score = 88.6 bits (218), Expect(2) = 1e-22
Identities = 55/131 (41%), Positives = 78/131 (59%), Gaps = 1/131 (0%)
Frame = +1
Query: 322 GQGSDS-MTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMD 498
G+ D+ +T+SA E CKKLL+DA +DLRGM + ++ +DL WL+ VMT++
Sbjct: 96 GKDDDAKITKSAIEMCKKLLDDAIEDLRGMASLKPEEVT---KHVNDLRCWLSSVMTYIY 152
Query: 499 TCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSE 678
TC DGF +LK M +++N+TELSSNALAI SLG ++ S + RRLL +
Sbjct: 153 TCADGFDKPELKEAMDKLLQNSTELSSNALAIITSLGELMPAAK-SNGSTGAHRRLLGLQ 211
Query: 679 QDEKGWPVWMR 711
E V +R
Sbjct: 212 GSEAAEGVSLR 222
Score = 42.4 bits (98), Expect(2) = 1e-22
Identities = 19/60 (31%), Positives = 34/60 (56%)
Frame = +2
Query: 143 TSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQSKSIGE 322
T+ + ++C+ T Y CE++L N T +P+ V + +VALE V +A +S +G+
Sbjct: 38 TANVRLSTVCSVTRYPGRCEQSLGPVVNDTIDPESVLRAALQVALEEVTSAFNRSMDVGK 97
>sptr|Q7X6B6|Q7X6B6 OSJNBb0079B02.8 protein (OSJNBa0063C18.2 protein).
Length = 971
Score = 466 bits (1198), Expect = e-130
Identities = 216/332 (65%), Positives = 264/332 (79%)
Frame = +1
Query: 694 WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLY 873
+P W+ + +R+LL +G Q KP+ +VAKDGSG FK+I +AV+AVPK R+VIYVKAG Y
Sbjct: 639 FPSWVSAHQRRLLQAGTQ-KPDKVVAKDGSGDFKTITEAVNAVPKNSPTRFVIYVKAGEY 697
Query: 874 DEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFHNT 1053
+E V +P NIFMYGDGP ++RV G KS DG+ TM T TFS E +GF+CK+MGF NT
Sbjct: 698 NEYVTIPSSLPNIFMYGDGPTKTRVLGNKSNKDGVATMATRTFSAEGNGFVCKSMGFVNT 757
Query: 1054 AGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAV 1233
AG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC V+GTID+IFGNSAAV
Sbjct: 758 AGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIFGNSAAV 817
Query: 1234 FQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRP 1413
FQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I SYLGRP
Sbjct: 818 FQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIASYLGRP 877
Query: 1414 WKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHV 1593
WKE++R V+MES I DF+KPEG+ W GD LKTLYYAEY N GPGAGTSKRV WPG+ V
Sbjct: 878 WKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVTWPGYRV 937
Query: 1594 IGRKEAEPFTAGPFIDGAMWLKYTGAPHILGF 1689
IG+ EA FTAG FIDG WLK T P+++GF
Sbjct: 938 IGQAEATQFTAGVFIDGLTWLKNTATPNVMGF 969
Score = 88.6 bits (218), Expect(2) = 1e-22
Identities = 55/131 (41%), Positives = 78/131 (59%), Gaps = 1/131 (0%)
Frame = +1
Query: 322 GQGSDS-MTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMD 498
G+ D+ +T+SA E CKKLL+DA +DLRGM + ++ +DL WL+ VMT++
Sbjct: 96 GKDDDAKITKSAIEMCKKLLDDAIEDLRGMASLKPEEVT---KHVNDLRCWLSSVMTYIY 152
Query: 499 TCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSE 678
TC DGF +LK M +++N+TELSSNALAI SLG ++ S + RRLL +
Sbjct: 153 TCADGFDKPELKEAMDKLLQNSTELSSNALAIITSLGELMPAAK-SNGSTGAHRRLLGLQ 211
Query: 679 QDEKGWPVWMR 711
E V +R
Sbjct: 212 GSEAAEGVSLR 222
Score = 42.4 bits (98), Expect(2) = 1e-22
Identities = 19/60 (31%), Positives = 34/60 (56%)
Frame = +2
Query: 143 TSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQSKSIGE 322
T+ + ++C+ T Y CE++L N T +P+ V + +VALE V +A +S +G+
Sbjct: 38 TANVRLSTVCSVTRYPGRCEQSLGPVVNDTIDPESVLRAALQVALEEVTSAFNRSMDVGK 97
>sptr|Q9FSQ0|Q9FSQ0 Putative pectin methylesterase.
Length = 717
Score = 466 bits (1198), Expect = e-130
Identities = 216/332 (65%), Positives = 264/332 (79%)
Frame = +1
Query: 694 WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLY 873
+P W+ + +R+LL +G Q KP+ +VAKDGSG FK+I +AV+AVPK R+VIYVKAG Y
Sbjct: 385 FPSWVSAHQRRLLQAGTQ-KPDKVVAKDGSGDFKTITEAVNAVPKNSPTRFVIYVKAGEY 443
Query: 874 DEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFHNT 1053
+E V +P NIFMYGDGP ++RV G KS DG+ TM T TFS E +GF+CK+MGF NT
Sbjct: 444 NEYVTIPSSLPNIFMYGDGPTKTRVLGNKSNKDGVATMATRTFSAEGNGFVCKSMGFVNT 503
Query: 1054 AGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAV 1233
AG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC V+GTID+IFGNSAAV
Sbjct: 504 AGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIFGNSAAV 563
Query: 1234 FQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRP 1413
FQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I SYLGRP
Sbjct: 564 FQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIASYLGRP 623
Query: 1414 WKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHV 1593
WKE++R V+MES I DF+KPEG+ W GD LKTLYYAEY N GPGAGTSKRV WPG+ V
Sbjct: 624 WKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVTWPGYRV 683
Query: 1594 IGRKEAEPFTAGPFIDGAMWLKYTGAPHILGF 1689
IG+ EA FTAG FIDG WLK T P+++GF
Sbjct: 684 IGQAEATQFTAGVFIDGLTWLKNTATPNVMGF 715
>sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11)
(AT5g49180/K21P3_5) (Pectinesterase).
Length = 571
Score = 408 bits (1048), Expect(2) = e-117
Identities = 207/465 (44%), Positives = 293/465 (63%), Gaps = 11/465 (2%)
Frame = +1
Query: 331 SDSMTESAREDCKKLLEDAADDLRGMLEMAGG-DIKVLFSRSDDLETWLTGVMTFMDTCV 507
+D T+ A E C+KL+ DA DDL+ L+ G I + +DL WL+G + + TC+
Sbjct: 114 NDKDTKGALELCEKLMNDATDDLKKCLDNFDGFSIPQIEDFVEDLRVWLSGSIAYQQTCM 173
Query: 508 DGF--VDEKLKADMHSVVRNATELSSNALAITNSLGGILKKM-------DLGMFSKDSRR 660
D F + KL DM + + + EL+SN LA+ ++ +L + DLG ++ R
Sbjct: 174 DTFEETNSKLSQDMQKIFKTSRELTSNGLAMITNISNLLGEFNVTGVTGDLGKYA----R 229
Query: 661 RLLSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQG 840
+LLS+E G P W+ R+L+A+ K N +VA DGSGQ+K+I +A++AVPK +Q
Sbjct: 230 KLLSAED---GIPSWVGPNTRRLMATKGGVKANVVVAHDGSGQYKTINEALNAVPKANQK 286
Query: 841 RYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADG-ITTMKTATFSVEAS 1017
+VIY+K G+Y+E V V K ++ GDGP ++++TG ++ G + T TAT ++
Sbjct: 287 PFVIYIKQGVYNEKVDVTKKMTHVTFIGDGPTKTKITGSLNYYIGKVKTYLTATVAINGD 346
Query: 1018 GFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSG 1197
F KN+GF NTAG E HQAVALRV DLA FYNC+ D +QDTLYVH+ RQFFR+C VSG
Sbjct: 347 NFTAKNIGFENTAGPEGHQAVALRVSADLAVFYNCQIDGYQDTLYVHSHRQFFRDCTVSG 406
Query: 1198 TIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFP 1377
T+DFIFG+ V QNC I+ R+PM +Q +TA GR+D +GLV+QNC + + P
Sbjct: 407 TVDFIFGDGIVVLQNCNIVVRKPMKSQSCMITAQGRSDKRESTGLVLQNCHITGEPAYIP 466
Query: 1378 DRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAG 1557
+ +YLGRPWKEFSR +IM +TI D + P G++PWNGDFAL TLYYAEY N GPG+
Sbjct: 467 VKSINKAYLGRPWKEFSRTIIMGTTIDDVIDPAGWLPWNGDFALNTLYYAEYENNGPGSN 526
Query: 1558 TSKRVNWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILGFK 1692
++RV WPG + K+A FT F+ G +W+ P++ F+
Sbjct: 527 QAQRVKWPGIKKLSPKQALRFTPARFLRGNLWIPPNRVPYMGNFQ 571
Score = 37.7 bits (86), Expect(2) = e-117
Identities = 15/57 (26%), Positives = 36/57 (63%)
Frame = +2
Query: 137 LPTSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQS 307
+ T+ +V+++CAPT YKE+C +L +A+ + P ++ V + S++ +++++
Sbjct: 48 IKTATTAVEAVCAPTDYKETCVNSLMKASPDSTQPLDLIKLGFNVTIRSIEDSIKKA 104
>sptrnew|BAD08732|BAD08732 Putative pectinesterase.
Length = 664
Score = 405 bits (1041), Expect(2) = e-117
Identities = 224/514 (43%), Positives = 300/514 (58%), Gaps = 67/514 (13%)
Frame = +1
Query: 334 DSMTESAREDCKKLLEDAADDL-RGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVD 510
D ++A DCK++ E+A DDL R + + G + L L WL+ V+ +TC+D
Sbjct: 144 DPRVKAAVADCKEIYENAKDDLDRTLAGIDAGGVDGLTKGGYQLRVWLSAVIAHQETCID 203
Query: 511 GFVDEKLKADMHSVVRNATELSSNALAIT------------------------------- 597
GF D LK M + + EL+SNALA+
Sbjct: 204 GFPDGDLKDKMRDAMESGKELTSNALALIGKASSFLAALHLPASSAASHRRLLSFAFDED 263
Query: 598 ------------NSLGGIL---------KKMDLGMFSKDSRRRLLSSEQDEKGWP----- 699
NSL +L K+ ++ S +S RRLLS DE
Sbjct: 264 VTKQPEVNRSSGNSLRRLLSFAFDEDATKQPEVNRSSGNSLRRLLSFAFDENAPKQPKGN 323
Query: 700 -----VWMRSPERKLLASG--NQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYV 858
VW+ ER+LL + N+ KPN +VAKDGSG+FK+I A+ A+PK + GRYVIYV
Sbjct: 324 DDDVLVWVNRQERRLLKAKFQNKLKPNVVVAKDGSGKFKTINDALAAMPKKYTGRYVIYV 383
Query: 859 KAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNM 1038
K G+Y+E V + K N+ MYGDG K++ +TG ++F DG+TT KTATF+ + GF+ +
Sbjct: 384 KEGVYEEYVTITKKMANVTMYGDGAKKTIITGNRNFVDGLTTYKTATFNAQGDGFMGVAL 443
Query: 1039 GFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFG 1218
GF NTA A +HQAVAL VQ D + F NCR + QDTLY H++ QF+RNCV+SGT+DFIFG
Sbjct: 444 GFRNTARAAKHQAVALLVQSDKSIFLNCRMEGHQDTLYAHSKAQFYRNCVISGTVDFIFG 503
Query: 1219 NSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKL-FPDRFKIP 1395
++AAVFQNC+I+ RRP+DNQQN TA GR D +G V+Q+ R + L R +
Sbjct: 504 DAAAVFQNCVIVLRRPLDNQQNIATAQGRADRREATGFVLQHYRFAAESALGDASRPAVR 563
Query: 1396 SYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVN 1575
SYL RPW+E+SR +IM S I FV GY+PW+GDF LKTL+YAEY N+G GA T+ RV+
Sbjct: 564 SYLARPWREYSRTLIMNSDIPAFVDKAGYLPWSGDFGLKTLWYAEYGNKGAGAATAGRVS 623
Query: 1576 WPGF-HVIGRKEAEPFTAGPFIDGAMWLKYTGAP 1674
WPG+ VI +KEA FT F+ W+K TG P
Sbjct: 624 WPGYKKVISKKEATKFTVQNFLHAEPWIKPTGTP 657
Score = 40.0 bits (92), Expect(2) = e-117
Identities = 19/61 (31%), Positives = 39/61 (63%), Gaps = 6/61 (9%)
Frame = +2
Query: 152 KSVKSLCAPTLYKESCEKTLSQ------ATNGTENPKEVFHSVAKVALESVQTAVEQSKS 313
K +K++CA T YK++CEK+L++ A++ + +PK+V + V ++++ A ++S
Sbjct: 80 KIIKAMCAQTDYKDTCEKSLAKAAANASASSSSSSPKDVVRASVAVIGDAIEKAFDKSSV 139
Query: 314 I 316
I
Sbjct: 140 I 140
>sptr|Q9M9W6|Q9M9W6 Putative pectinesterase.
Length = 669
Score = 388 bits (996), Expect(2) = e-114
Identities = 194/457 (42%), Positives = 285/457 (62%), Gaps = 6/457 (1%)
Frame = +1
Query: 334 DSMTESAREDCKKLLEDAADDLRGMLEMAGG-DIKVLFSRSDDLETWLTGVMTFMDTCVD 510
DS T A + CK+L++ A D+L E G + +L +L WL+ ++ +TC++
Sbjct: 116 DSRTRMALDQCKELMDYALDELSNSFEELGKFEFHLLDEALINLRIWLSAAISHEETCLE 175
Query: 511 GFVDEKLKAD--MHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQD 684
GF + A M ++ A EL+ N LAI + + + +M + + RRLL+
Sbjct: 176 GFQGTQGNAGETMKKALKTAIELTHNGLAIISEMSNFVGQMQIPGLNS---RRLLA---- 228
Query: 685 EKGWPVWMRSPERKLL---ASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIY 855
+G+P W+ RKLL A+ + KP+ +VA+DGSGQ+K+I +A+ VPK +V++
Sbjct: 229 -EGFPSWVDQRGRKLLQAAAAYSDVKPDIVVAQDGSGQYKTINEALQFVPKKRNTTFVVH 287
Query: 856 VKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKN 1035
+KAGLY E V V K ++ GDGP ++ ++G K++ DGITT +TAT ++ + FI KN
Sbjct: 288 IKAGLYKEYVQVNKTMSHLVFIGDGPDKTIISGNKNYKDGITTYRTATVAIVGNYFIAKN 347
Query: 1036 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIF 1215
+GF NTAGA +HQAVA+RVQ D + F+NCRFD +QDTLY H+ RQFFR+C +SGTIDF+F
Sbjct: 348 IGFENTAGAIKHQAVAVRVQSDESIFFNCRFDGYQDTLYTHSHRQFFRDCTISGTIDFLF 407
Query: 1216 GNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 1395
G++AAVFQNC ++ R+P+ NQ +TAHGR DP +G V Q C + + +
Sbjct: 408 GDAAAVFQNCTLLVRKPLPNQACPITAHGRKDPRESTGFVFQGCTIAGEPDYLAVKETSK 467
Query: 1396 SYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVN 1575
+YLGRPWKE+SR +IM + I DFV+P+G+ PW GDF LKTL+Y+E N GPG+ + RV
Sbjct: 468 AYLGRPWKEYSRTIIMNTFIPDFVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVT 527
Query: 1576 WPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
W G + ++ FT +I G W+ G P+ G
Sbjct: 528 WAGIKTLSEEDILKFTPAQYIQGDDWIPGKGVPYTTG 564
Score = 47.4 bits (111), Expect(2) = e-114
Identities = 22/66 (33%), Positives = 40/66 (60%)
Frame = +2
Query: 125 GRLPLPTSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQ 304
G+ + S K+VK +CAPT Y+++CE TL + T +P E+ + V ++ + A ++
Sbjct: 47 GKAEVNASVKAVKDVCAPTDYRKTCEDTLIKNGKNTTDPMELVKTAFNVTMKQITDAAKK 106
Query: 305 SKSIGE 322
S++I E
Sbjct: 107 SQTIME 112
>sptr|Q7Y201|Q7Y201 Putative pectin methylesterase.
Length = 614
Score = 397 bits (1019), Expect(2) = e-113
Identities = 204/455 (44%), Positives = 295/455 (64%), Gaps = 8/455 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAG-GDIKVLFSRSDDLETWLTGVMTFMDTCVDGFVD 522
+ A E CK L+EDA ++ L ++ DLE+WL+ VM++ +TC+DGF +
Sbjct: 172 KDAIEQCKLLVEDAKEETVASLNKINVTEVNSFEKVVPDLESWLSAVMSYQETCLDGFEE 231
Query: 523 EKLKADMHSVVRNATELSSNALAI----TNSLGGILKKMDLGMFSKDSRRRLLSSEQDEK 690
LK+++ + V ++ L+SN+LA+ T +L ++K ++ R LL
Sbjct: 232 GNLKSEVKTSVNSSQVLTSNSLALIKTFTENLSPVMKVVE---------RHLLD------ 276
Query: 691 GWPVWMRSPERKLLASGNQP--KPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKA 864
G P W+ + +R++L + + KPNA VAKDGSG F +I A+ A+P+ ++GRY+IYVK
Sbjct: 277 GIPSWVSNDDRRMLRAVDVKALKPNATVAKDGSGDFTTINDALRAMPEKYEGRYIIYVKQ 336
Query: 865 GLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGF 1044
G+YDE V V K K N+ M GDG +++ VTG KS A I T TATF + GF+ ++MGF
Sbjct: 337 GIYDEYVTVDKKKANLTMVGDGSQKTIVTGNKSHAKKIRTFLTATFVAQGEGFMAQSMGF 396
Query: 1045 HNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNS 1224
NTAG+E HQAVA+RVQ D + F NCRF+ +QDTLY + RQ++R+CV+ GTIDFIFG++
Sbjct: 397 RNTAGSEGHQAVAIRVQSDRSIFLNCRFEGYQDTLYAYTHRQYYRSCVIVGTIDFIFGDA 456
Query: 1225 AAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYL 1404
AA+FQNC I R+ + Q+N+VTA GR D +G V+ NC++ ++ L P + + SYL
Sbjct: 457 AAIFQNCNIFIRKGLPGQKNTVTAQGRVDKFQTTGFVVHNCKIAANEDLKPVKEEYKSYL 516
Query: 1405 GRPWKEFSRLVIMESTIADFVKPEGYMPW-NGDFALKTLYYAEYNNRGPGAGTSKRVNWP 1581
GRPWK +SR +IMES I + + P G++ W DFA+ TLYYAEYNN+G T+ RV WP
Sbjct: 517 GRPWKNYSRTIIMESKIENVIDPVGWLRWQETDFAIDTLYYAEYNNKGSSGDTTSRVKWP 576
Query: 1582 GFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
GF VI ++EA +T GPF+ G W+ +G+P LG
Sbjct: 577 GFKVINKEEALNYTVGPFLQGD-WISASGSPVKLG 610
Score = 35.4 bits (80), Expect(2) = e-113
Identities = 18/58 (31%), Positives = 30/58 (51%), Gaps = 3/58 (5%)
Frame = +2
Query: 152 KSVKSLCAPTLYKESCEKTLSQATN---GTENPKEVFHSVAKVALESVQTAVEQSKSI 316
K +++LC+ TLY + CEKTL T+ +NP S + E + +E+ S+
Sbjct: 107 KIIQTLCSSTLYMQICEKTLKNRTDKGFALDNPTTFLKSAIEAVNEDLDLVLEKVLSL 164
>sptrnew|BAC83510|BAC83510 Putative pectin methylesterase.
Length = 566
Score = 383 bits (983), Expect(2) = e-113
Identities = 205/463 (44%), Positives = 274/463 (59%), Gaps = 11/463 (2%)
Frame = +1
Query: 331 SDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRS-DDLETWLTGVMTFMDTCV 507
+D T A ++CK+LLE A DDL+ E GG F ++ DDL TWL+ +T+ TC+
Sbjct: 103 NDKRTSGALQNCKELLEYAVDDLKTSFEKLGGFEMTNFHKAVDDLRTWLSAALTYQGTCL 162
Query: 508 DGFVDEKLKA--DMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQ 681
DGF++ A M S + ++ EL+ + LA+ + L +++G RRRLL+ +
Sbjct: 163 DGFLNTTTDAADKMKSALNSSQELTEDILAVVDQFSATLGSLNIG------RRRLLADD- 215
Query: 682 DEKGWPVWMRSPERKLLASGNQP-------KPNAIVAKDGSGQFKSIQQAVDAVPKGHQG 840
G PVWM R+ L P KP+ VA DGSG K+I +AV VP ++
Sbjct: 216 ---GMPVWMSEGGRRQLLEAAGPEAGPVEFKPDVTVAADGSGDVKTIGEAVAKVPPKNKE 272
Query: 841 RYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASG 1020
RY IYVKAG Y+E V V + N+ M GDG ++ +TG K+F +TT TAT +G
Sbjct: 273 RYTIYVKAGTYNEYVSVGRPATNVNMIGDGIGKTIITGNKNFKMNLTTKDTATMEAIGNG 332
Query: 1021 FICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGT 1200
F + + NTAG E HQAVALR Q D+A FY C FD +QDTLY HA+RQFFR+C VSGT
Sbjct: 333 FFMRGITVENTAGPENHQAVALRAQSDMAVFYQCEFDGYQDTLYPHAQRQFFRDCTVSGT 392
Query: 1201 IDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPD 1380
IDFIFGNS V QNCL+ R+PMDNQ N +TA GR + G VI NC + P L
Sbjct: 393 IDFIFGNSQVVLQNCLLQPRKPMDNQVNIITAQGRREKRSAGGTVIHNCTVAPHPDLEKF 452
Query: 1381 RFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGT 1560
K+ +YL RPWKE+SR + +++ I V P G++ WNG+FAL TLYYAE +N GPGA
Sbjct: 453 TDKVKTYLARPWKEYSRTIFVQNEIGAVVDPVGWLEWNGNFALDTLYYAEVDNHGPGADM 512
Query: 1561 SKRVNWPGFHVIGRKEAE-PFTAGPFIDGAMWLKYTGAPHILG 1686
SKR W G + ++ + FT FI G ++ G P+I G
Sbjct: 513 SKRAKWKGVQSLTYQDVQKEFTVEAFIQGQEFIPKFGVPYIPG 555
Score = 49.3 bits (116), Expect(2) = e-113
Identities = 24/65 (36%), Positives = 40/65 (61%), Gaps = 1/65 (1%)
Frame = +2
Query: 137 LPTSGKSVKSLCAPTLYKESCEKTLSQAT-NGTENPKEVFHSVAKVALESVQTAVEQSKS 313
L TS KSVK+ C PT Y+++CE+ L +A NG +P ++ ++ V E + A+ +S +
Sbjct: 38 LSTSVKSVKAFCQPTDYQQTCEEELGKAAGNGASSPTDLAKAMFAVTSEKISKAISESST 97
Query: 314 IGEAK 328
+ E K
Sbjct: 98 LEELK 102
>sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610).
Length = 669
Score = 384 bits (986), Expect(2) = e-112
Identities = 192/457 (42%), Positives = 284/457 (62%), Gaps = 6/457 (1%)
Frame = +1
Query: 334 DSMTESAREDCKKLLEDAADDLRGMLEMAGG-DIKVLFSRSDDLETWLTGVMTFMDTCVD 510
DS T A + CK+L++ A D+L E G + +L +L WL+ ++ +TC++
Sbjct: 116 DSRTRMALDQCKELMDYALDELSNSFEELGKFEFHLLDEALINLRIWLSAAISHEETCLE 175
Query: 511 GFVDEKLKAD--MHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQD 684
GF + A M ++ A EL+ N LAI + + + +M + + RRLL+
Sbjct: 176 GFQGTQGNAGETMKKALKTAIELTHNGLAIISEMSNFVGQMQIPGLNS---RRLLA---- 228
Query: 685 EKGWPVWMRSPERKLL---ASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIY 855
+G+P W+ RKLL A+ + KP+ +VA+DGSGQ+K+I +A+ VPK +V++
Sbjct: 229 -EGFPSWVDQRGRKLLQAAAAYSDVKPDIVVAQDGSGQYKTINEALQFVPKKRNTTFVVH 287
Query: 856 VKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKN 1035
+KAGLY E V V K ++ GDGP ++ ++G K++ DGIT +TAT ++ + FI KN
Sbjct: 288 IKAGLYKEYVQVNKTMSHLVFIGDGPDKTIISGNKNYKDGITAYRTATVAIVGNYFIAKN 347
Query: 1036 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIF 1215
+GF NTAGA +HQAVA+RVQ D + F+NCRFD +Q+TLY H+ RQFFR+C +SGTIDF+F
Sbjct: 348 IGFENTAGAIKHQAVAVRVQSDESIFFNCRFDGYQNTLYTHSHRQFFRDCTISGTIDFLF 407
Query: 1216 GNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 1395
G++AAVFQNC ++ R+P+ NQ +TAHGR DP +G V Q C + + +
Sbjct: 408 GDAAAVFQNCTLLVRKPLPNQACPITAHGRKDPRESTGFVFQGCTIAGEPDYLAVKETSK 467
Query: 1396 SYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVN 1575
+YLGRPWKE+SR +IM + I DFV+P+G+ PW GDF LKTL+Y+E N GPG+ + RV
Sbjct: 468 AYLGRPWKEYSRTIIMNTFIPDFVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVT 527
Query: 1576 WPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
W G + ++ FT +I G W+ G P+ G
Sbjct: 528 WAGIKTLSEEDILKFTPAQYIQGDDWIPGKGVPYTTG 564
Score = 47.4 bits (111), Expect(2) = e-112
Identities = 22/66 (33%), Positives = 40/66 (60%)
Frame = +2
Query: 125 GRLPLPTSGKSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQ 304
G+ + S K+VK +CAPT Y+++CE TL + T +P E+ + V ++ + A ++
Sbjct: 47 GKAEVNASVKAVKDVCAPTDYRKTCEDTLIKNGKNTTDPMELVKTAFNVTMKQITDAAKK 106
Query: 305 SKSIGE 322
S++I E
Sbjct: 107 SQTIME 112
>sptr|Q9SMY6|Q9SMY6 Pectinesterase-like protein.
Length = 609
Score = 398 bits (1023), Expect(2) = e-112
Identities = 206/454 (45%), Positives = 290/454 (63%), Gaps = 7/454 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDL-RGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGFVD 522
+ A CK L+++A ++L M + ++ DL++WL+ VM++ +TCVDGF +
Sbjct: 158 KDAIAQCKLLVDEAKEELGTSMKRINDSEVNNFAKIVPDLDSWLSAVMSYQETCVDGFEE 217
Query: 523 EKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLL---SSEQDEKG 693
KLK ++ ++ L+SN+LA+ SL G L + K R LL SS ++
Sbjct: 218 GKLKTEIRKNFNSSQVLTSNSLAMIKSLDGYLSSVP-----KVKTRLLLEARSSAKETDH 272
Query: 694 WPVWMRSPERKLLASGNQP--KPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAG 867
W+ + ER++L + + KPNA VAKDGSG F +I A+ A+P +QGRY IY+K G
Sbjct: 273 ITSWLSNKERRMLKAVDVKALKPNATVAKDGSGNFTTINAALKAMPAKYQGRYTIYIKHG 332
Query: 868 LYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFH 1047
+YDE V++ K K N+ M GDG +++ VTG KS A I T TATF + GF+ ++MGF
Sbjct: 333 IYDESVIIDKKKPNVTMVGDGSQKTIVTGNKSHAKKIRTFLTATFVAQGEGFMAQSMGFR 392
Query: 1048 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSA 1227
NTAG E HQAVA+RVQ D + F NCRF+ +QDTLY + RQ++R+CV+ GT+DFIFG++A
Sbjct: 393 NTAGPEGHQAVAIRVQSDRSVFLNCRFEGYQDTLYAYTHRQYYRSCVIIGTVDFIFGDAA 452
Query: 1228 AVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 1407
A+FQNC I R+ + Q+N+VTA GR D +G VI NC + P++ L P + + SYLG
Sbjct: 453 AIFQNCDIFIRKGLPGQKNTVTAQGRVDKFQTTGFVIHNCTVAPNEDLKPVKAQFKSYLG 512
Query: 1408 RPWKEFSRLVIMESTIADFVKPEGYMPW-NGDFALKTLYYAEYNNRGPGAGTSKRVNWPG 1584
RPWK SR V+MESTI D + P G++ W DFA+ TL YAEY N GP T+ RV WPG
Sbjct: 513 RPWKPHSRTVVMESTIEDVIDPVGWLRWQETDFAIDTLSYAEYKNDGPSGATAARVKWPG 572
Query: 1585 FHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
F V+ ++EA FT GPF+ G W++ G+P LG
Sbjct: 573 FRVLNKEEAMKFTVGPFLQGE-WIQAIGSPVKLG 605
Score = 32.7 bits (73), Expect(2) = e-112
Identities = 17/51 (33%), Positives = 29/51 (56%)
Frame = +2
Query: 152 KSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQ 304
K +++LC TLYK +C+ TL T + P+ S+ K A+ +V ++Q
Sbjct: 93 KIIQTLCNSTLYKPTCQNTLKNETK-KDTPQTDPRSLLKSAIVAVNDDLDQ 142
>sptr|Q9SG77|Q9SG77 Putative pectinesterase.
Length = 561
Score = 373 bits (958), Expect(2) = e-109
Identities = 203/452 (44%), Positives = 277/452 (61%), Gaps = 1/452 (0%)
Frame = +1
Query: 322 GQGSDSMTESAREDCKKLLEDAADDLRGML-EMAGGDIKVLFSRSDDLETWLTGVMTFMD 498
G +++T +A C +LL+ D+L L + GD+ V DDL TWL+ T+
Sbjct: 125 GDEKNNITMNA---CAELLDLTIDNLNNTLTSSSNGDVTVP-ELVDDLRTWLSSAGTYQR 180
Query: 499 TCVDGFVDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSE 678
TCV+ + ++ S ++N+TEL+SNALAI LG I L RRRLL++
Sbjct: 181 TCVETLAPD-MRPFGESHLKNSTELTSNALAIITWLGKIADSFKL-------RRRLLTTA 232
Query: 679 QDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYV 858
E V + R L ++ + + +VAKDGSG++++I++A+ VP+ + R +IYV
Sbjct: 233 DVE----VDFHAGRRLLQSTDLRKVADIVVAKDGSGKYRTIKRALQDVPEKSEKRTIIYV 288
Query: 859 KAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNM 1038
K G+Y E V V K N+ + GDG +S V+GR + DG T KTATF+V GF+ ++M
Sbjct: 289 KKGVYFENVKVEKKMWNVIVVGDGESKSIVSGRLNVIDGTPTFKTATFAVFGKGFMARDM 348
Query: 1039 GFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFG 1218
GF NTAG +HQAVAL V DL AFY C +A+QDTLYVHA+RQF+R C + GT+DFIFG
Sbjct: 349 GFINTAGPSKHQAVALMVSADLTAFYRCTMNAYQDTLYVHAQRQFYRECTIIGTVDFIFG 408
Query: 1219 NSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPS 1398
NSA+V Q+C I+ RRPM QQN++TA GRTDPNM +G+ I C + P D + +
Sbjct: 409 NSASVLQSCRILPRRPMKGQQNTITAQGRTDPNMNTGISIHRCNISP----LGDLTDVMT 464
Query: 1399 YLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNW 1578
+LGRPWK FS VIM+S + F+ +G++PW GD A T++Y EY N GPGA T RV W
Sbjct: 465 FLGRPWKNFSTTVIMDSYLHGFIDRKGWLPWTGDSAPDTIFYGEYKNTGPGASTKNRVKW 524
Query: 1579 PGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAP 1674
G + KEA FT PFIDG WL T P
Sbjct: 525 KGLRFLSTKEANRFTVKPFIDGGRWLPATKVP 556
Score = 45.8 bits (107), Expect(2) = e-109
Identities = 24/60 (40%), Positives = 36/60 (60%), Gaps = 2/60 (3%)
Frame = +2
Query: 155 SVKSLCAPTLYKESCEKTLSQATNGTE-NPKEVFHSVAKVALESVQTAVEQ-SKSIGEAK 328
SVK++C TL+KE C +TL A N + NP+E+F K+ + V A+ S S+G+ K
Sbjct: 69 SVKAVCDVTLHKEKCFETLGSAPNASSLNPEELFRYAVKITIAEVSKAINAFSSSLGDEK 128
>sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11)
(Fragment).
Length = 426
Score = 394 bits (1013), Expect = e-108
Identities = 205/425 (48%), Positives = 272/425 (64%), Gaps = 7/425 (1%)
Frame = +1
Query: 433 KVLFSRSDDLETWLTGVMTFMDTCVDGF----VDEKLKADMHSVVRNATELSSNALAITN 600
K L+ +DDL+T ++ +T TC+DGF D++++ + + + SNALA+T
Sbjct: 6 KTLYQHADDLKTLISAAITNQVTCLDGFSHDGADKQVRKVLEQGQVHVEHMCSNALAMTK 65
Query: 601 SLGGILKKMDLGMFSKDSRR--RLLSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAK 774
++ D+ F +++ + R L E++ WP W+ + +R+LL G K + +VA
Sbjct: 66 NM----TDKDIAKFEENNNKKNRKLLEEENGVNWPEWISAGDRRLL-QGAAVKADVVVAA 120
Query: 775 DGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTG 954
DGSG FK++ +AV P RYVI +KAG+Y E V VPK K NI GDG K + +T
Sbjct: 121 DGSGNFKTVSEAVAGAPLKSSKRYVIKIKAGVYKENVEVPKKKSNIMFLGDGKKNTIITA 180
Query: 955 RKSFADGITTMKTATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDA 1134
++ DG TT +AT +V F+ +++ F NTAG +HQAVALRV GDL+AFYNC A
Sbjct: 181 SRNVVDGSTTFHSATVAVVGGNFLARDITFQNTAGPSKHQAVALRVGGDLSAFYNCDIIA 240
Query: 1135 FQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDP 1314
+QDTLYVH RQFF NC +SGT+DFIFGNSA VFQNC I R+P Q+N VTA GR DP
Sbjct: 241 YQDTLYVHTNRQFFVNCFISGTVDFIFGNSAVVFQNCDIHARKPDSGQKNMVTAQGRVDP 300
Query: 1315 NMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWN 1494
N +G+VIQ CR+ + L + P+YLGRPWKE+SR VIM+S+I+D + P G+ WN
Sbjct: 301 NQNTGIVIQKCRIGATKDLEGLKGTFPTYLGRPWKEYSRTVIMQSSISDVIDPIGWHEWN 360
Query: 1495 GDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHVI-GRKEAEPFTAGPFIDGAMWLKYTGA 1671
G+FAL TL Y EY N GPGAGTSKRVNW GF VI EA+ FT G FI G+ WL TG
Sbjct: 361 GNFALNTLVYREYQNTGPGAGTSKRVNWKGFKVITSASEAQTFTPGNFIGGSTWLGSTGF 420
Query: 1672 PHILG 1686
P LG
Sbjct: 421 PFSLG 425
>sptr|O48712|O48712 Putative pectinesterase.
Length = 496
Score = 394 bits (1011), Expect = e-108
Identities = 203/455 (44%), Positives = 293/455 (64%), Gaps = 8/455 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAG-GDIKVLFSRSDDLETWLTGVMTFMDTCVDGFVD 522
+ A E CK L+EDA ++ L ++ DLE+WL+ VM++ +TC+DGF +
Sbjct: 54 KDAIEQCKLLVEDAKEETVASLNKINVTEVNSFEKVVPDLESWLSAVMSYQETCLDGFEE 113
Query: 523 EKLKADMHSVVRNATELSSNALAI----TNSLGGILKKMDLGMFSKDSRRRLLSSEQDEK 690
LK+++ + V ++ L+SN+LA+ T +L ++K ++ R LL
Sbjct: 114 GNLKSEVKTSVNSSQVLTSNSLALIKTFTENLSPVMKVVE---------RHLLDDI---- 160
Query: 691 GWPVWMRSPERKLLASGNQP--KPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKA 864
P W+ + +R++L + + KPNA VAKDGSG F +I A+ A+P+ ++GRY+IYVK
Sbjct: 161 --PSWVSNDDRRMLRAVDVKALKPNATVAKDGSGDFTTINDALRAMPEKYEGRYIIYVKQ 218
Query: 865 GLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGF 1044
G+YDE V V K K N+ M GDG +++ VTG KS A I T TATF + GF+ ++MGF
Sbjct: 219 GIYDEYVTVDKKKANLTMVGDGSQKTIVTGNKSHAKKIRTFLTATFVAQGEGFMAQSMGF 278
Query: 1045 HNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNS 1224
NTAG E HQAVA+RVQ D + F NCRF+ +QDTLY + RQ++R+CV+ GTIDFIFG++
Sbjct: 279 RNTAGPEGHQAVAIRVQSDRSIFLNCRFEGYQDTLYAYTHRQYYRSCVIVGTIDFIFGDA 338
Query: 1225 AAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYL 1404
AA+FQNC I R+ + Q+N+VTA GR D +G V+ NC++ ++ L P + + SYL
Sbjct: 339 AAIFQNCNIFIRKGLPGQKNTVTAQGRVDKFQTTGFVVHNCKIAANEDLKPVKEEYKSYL 398
Query: 1405 GRPWKEFSRLVIMESTIADFVKPEGYMPW-NGDFALKTLYYAEYNNRGPGAGTSKRVNWP 1581
GRPWK +SR +IMES I + + P G++ W DFA+ TLYYAEYNN+G T+ RV WP
Sbjct: 399 GRPWKNYSRTIIMESKIENVIDPVGWLRWQETDFAIDTLYYAEYNNKGSSGDTTSRVKWP 458
Query: 1582 GFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
GF VI ++EA +T GPF+ G W+ +G+P LG
Sbjct: 459 GFKVINKEEALNYTVGPFLQGD-WISASGSPVKLG 492
>sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC
3.1.1.11).
Length = 579
Score = 390 bits (1002), Expect(2) = e-107
Identities = 206/455 (45%), Positives = 289/455 (63%), Gaps = 8/455 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLR-GMLEMAG-GDIKVLFSRSDDLETWLTGVMTFMDTCVDGF- 516
++A DC + +++ D+L ++++G + K L ++D+L T L+ +T +TC+DGF
Sbjct: 129 KTALHDCLETIDETLDELHEAQVDISGYPNKKSLKEQADNLITLLSSAITNQETCLDGFS 188
Query: 517 ---VDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQ-D 684
D+K++ + + ++ SNALA+ ++ D+ +++ R+L ++ +
Sbjct: 189 HDGADKKVRKALLKGQTHVEKMCSNALAMIKNM----TDTDIANELQNTNRKLKEEKEGN 244
Query: 685 EKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKA 864
E+ WP WM +R+LL S + PN +VA DGSG +K++ +AV A PK RY+I +KA
Sbjct: 245 ERVWPEWMSVADRRLLQSSSVT-PNVVVAADGSGDYKTVSEAVAAAPKKSSKRYIIQIKA 303
Query: 865 GLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGF 1044
G+Y E V VPKDK NI GDG K + +T ++ DG TT K+AT + GF+ + + F
Sbjct: 304 GVYRENVEVPKDKHNIMFLGDGRKTTIITASRNVVDGSTTFKSATVAAVGQGFLARGVTF 363
Query: 1045 HNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNS 1224
NTAG +HQAVALRV DL+AFY C A+QDTLYVH+ RQFF NC V+GT+DFIFGN+
Sbjct: 364 ENTAGPSKHQAVALRVGSDLSAFYECDMLAYQDTLYVHSNRQFFINCFVAGTVDFIFGNA 423
Query: 1225 AAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYL 1404
AAVFQ+C RRP Q+N VTA GRTDPN +G+VIQ R+ L P + P+YL
Sbjct: 424 AAVFQDCDYHARRPDSGQKNMVTAQGRTDPNQNTGIVIQKSRIGATSDLLPVQSSFPTYL 483
Query: 1405 GRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPG 1584
GRPWKE+SR VIM+S+I D ++P G+ W+G FAL TL+YAEY N G GAGTS RV W G
Sbjct: 484 GRPWKEYSRTVIMQSSITDVIQPAGWHEWSGSFALSTLFYAEYQNSGAGAGTSSRVKWEG 543
Query: 1585 FHVI-GRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
+ VI EA+ F G FI G+ WL T P LG
Sbjct: 544 YKVITSATEAQAFAPGNFIAGSSWLGSTSFPFSLG 578
Score = 23.9 bits (50), Expect(2) = e-107
Identities = 10/27 (37%), Positives = 14/27 (51%)
Frame = +2
Query: 158 VKSLCAPTLYKESCEKTLSQATNGTEN 238
+KS C+ TLY E C ++ T N
Sbjct: 63 LKSACSSTLYPELCYSAIATVPGVTGN 89
>sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 584
Score = 390 bits (1002), Expect = e-107
Identities = 205/452 (45%), Positives = 279/452 (61%), Gaps = 7/452 (1%)
Frame = +1
Query: 352 AREDCKKLLEDAADDLRGMLEMAGG--DIKVLFSRSDDLETWLTGVMTFMDTCVDGFVDE 525
A DC + +++ D+L +E + K L +DDL+T ++ MT TC+DGF +
Sbjct: 137 ALHDCLETIDETLDELHKAVEDLEEYPNKKSLSQHADDLKTLMSAAMTNQGTCLDGFSHD 196
Query: 526 KLKADMHSVVRNAT----ELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQDEKG 693
+ + + ++ SNALA+ ++ D+ + + R+L G
Sbjct: 197 DANKHVRDALSDGQVHVEKMCSNALAMIKNM----TDTDMMIMRTSNNRKLTEETSTVDG 252
Query: 694 WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLY 873
WP W+ +R+LL S + PNA+VA DGSG FK++ AV A P+G RY+I +KAG+Y
Sbjct: 253 WPAWLSPGDRRLLQSSSVT-PNAVVAADGSGNFKTVAAAVAAAPQGGTKRYIIRIKAGVY 311
Query: 874 DEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFHNT 1053
E V V K NI GDG ++ +TG ++ DG TT K+AT +V GF+ +++ F NT
Sbjct: 312 RENVEVTKKHKNIMFIGDGRTRTIITGSRNVVDGSTTFKSATAAVVGEGFLARDITFQNT 371
Query: 1054 AGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAV 1233
AG +HQAVALRV DL+AFYNC A+QDTLYVH+ RQFF NC+++GT+DFIFGN+AAV
Sbjct: 372 AGPSKHQAVALRVGADLSAFYNCDMLAYQDTLYVHSNRQFFVNCLIAGTVDFIFGNAAAV 431
Query: 1234 FQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRP 1413
QNC I R+P Q+N VTA GRTDPN +G+VIQ R+ L P + P+YLGRP
Sbjct: 432 LQNCDIHARKPNSGQKNMVTAQGRTDPNQNTGIVIQKSRIGATSDLKPVQGSFPTYLGRP 491
Query: 1414 WKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHV 1593
WKE+SR VIM+S+I D + P G+ W+G+FAL TL+Y E+ N G GAGTS RV W GF V
Sbjct: 492 WKEYSRTVIMQSSITDLIHPAGWHEWDGNFALNTLFYGEHQNSGAGAGTSGRVKWKGFRV 551
Query: 1594 I-GRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
I EA+ FT G FI G+ WL TG P LG
Sbjct: 552 ITSATEAQAFTPGSFIAGSSWLGSTGFPFSLG 583
>sptr|Q9M7Y9|Q9M7Y9 Pectin methylesterase, putative (EC 3.1.1.11)
(Pectinesterase).
Length = 562
Score = 369 bits (946), Expect(2) = e-105
Identities = 191/451 (42%), Positives = 280/451 (62%), Gaps = 7/451 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGF--V 519
+ A E C+KL+ DA DDL+ ++ G + + +DL WL+G + F TC+D F +
Sbjct: 115 KGAFELCEKLMIDAIDDLKKCMDH-GFSVDQIEVFVEDLRVWLSGSIAFQQTCMDSFGEI 173
Query: 520 DEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQDEKGWP 699
L DM + + + ELSSN+LA+ + ++ +L + R+LLS+E P
Sbjct: 174 KSNLMQDMLKIFKTSRELSSNSLAMVTRISTLIPNSNL---TAKYARKLLSTEDSI---P 227
Query: 700 VWMRSPERKLLAS-GNQPKP---NAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAG 867
W+ R+L+A+ G P P NA+VA+DG+GQFK+I A++AVPKG++ ++I++K G
Sbjct: 228 TWVGPEARRLMAAQGGGPGPVKANAVVAQDGTGQFKTITDALNAVPKGNKVPFIIHIKEG 287
Query: 868 LYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADG-ITTMKTATFSVEASGFICKNMGF 1044
+Y E V V K ++ GDGP ++ +TG +F G + T TAT ++E F KN+G
Sbjct: 288 IYKEKVTVTKKMPHVTFIGDGPNKTLITGSLNFGIGKVKTFLTATITIEGDHFTAKNIGI 347
Query: 1045 HNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNS 1224
NTAG E QAVALRV D A F++C+ D QDTLYVH+ RQF+R+C VSGT+DFIFG++
Sbjct: 348 ENTAGPEGGQAVALRVSADYAVFHSCQIDGHQDTLYVHSHRQFYRDCTVSGTVDFIFGDA 407
Query: 1225 AAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYL 1404
+ QNC I+ R+P Q VTA GR++ +GLV+ C + D P + +YL
Sbjct: 408 KCILQNCKIVVRKPNKGQTCMVTAQGRSNVRESTGLVLHGCHITGDPAYIPMKSVNKAYL 467
Query: 1405 GRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPG 1584
GRPWKEFSR +IM++TI D + P G++PW+GDFALKTLYYAE+ N GPG+ ++RV WPG
Sbjct: 468 GRPWKEFSRTIIMKTTIDDVIDPAGWLPWSGDFALKTLYYAEHMNTGPGSNQAQRVKWPG 527
Query: 1585 FHVIGRKEAEPFTAGPFIDGAMWLKYTGAPH 1677
+ ++A +T F+ G W+ T P+
Sbjct: 528 IKKLTPQDALLYTGDRFLRGDTWIPQTQVPY 558
Score = 39.3 bits (90), Expect(2) = e-105
Identities = 17/59 (28%), Positives = 39/59 (66%)
Frame = +2
Query: 152 KSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQSKSIGEAK 328
K+V+++CAPT +K++C +L A+ +++P ++ KV ++S+ ++E++ +AK
Sbjct: 49 KAVQAVCAPTDFKDTCVNSLMGASPDSDDPVDLIKLGFKVTIKSINESLEKASGDIKAK 107
>sptr|Q8GXA1|Q8GXA1 Putative pectin methylesterase.
Length = 568
Score = 369 bits (946), Expect(2) = e-105
Identities = 193/455 (42%), Positives = 283/455 (62%), Gaps = 11/455 (2%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGF--V 519
+ A E C+KL+ DA DDL+ ++ G + + +DL WL+G + F TC+D F +
Sbjct: 115 KGAFELCEKLMIDAIDDLKKCMDH-GFSVDQIEVFVEDLRVWLSGSIAFQQTCMDSFGEI 173
Query: 520 DEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDL----GMFSKDSRRRLLSSEQDE 687
L DM + + + ELSSN+LA+ + ++ +L G +K +R+ LLS+E
Sbjct: 174 KSNLMQDMLKIFKTSRELSSNSLAMVTRISTLIPNSNLTGLTGALAKYARK-LLSTEDSI 232
Query: 688 KGWPVWMRSPERKLLAS-GNQPKP---NAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIY 855
P W+ R+L+A+ G P P NA+VA+DG+GQFK+I A++AVPKG++ ++I+
Sbjct: 233 ---PTWVGPEARRLMAAQGGGPGPVKANAVVAQDGTGQFKTITDALNAVPKGNKVPFIIH 289
Query: 856 VKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADG-ITTMKTATFSVEASGFICK 1032
+K G+Y E V V K ++ GDGP ++ +TG +F G + T TAT ++E F K
Sbjct: 290 IKEGIYKEKVTVTKKMPHVTFIGDGPNKTLITGSLNFGIGKVKTFLTATITIEGDHFTAK 349
Query: 1033 NMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFI 1212
N+G NTAG E QAVALRV D A F++C+ D QDTLYVH+ RQF+R+C VSGT+DFI
Sbjct: 350 NIGIENTAGPEGGQAVALRVSADYAVFHSCQIDGHQDTLYVHSHRQFYRDCTVSGTVDFI 409
Query: 1213 FGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKI 1392
FG++ + QNC I+ R+P Q VTA GR++ +GLV+ C + D P +
Sbjct: 410 FGDAKCILQNCKIVVRKPNKGQTCMVTAQGRSNVRESTGLVLHGCHITGDPAYIPMKSVN 469
Query: 1393 PSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRV 1572
+YLGRPWKEFSR +IM++TI D + P G++PW+GDFALKTLYYAE+ N GPG+ ++RV
Sbjct: 470 KAYLGRPWKEFSRTIIMKTTIDDVIDPAGWLPWSGDFALKTLYYAEHMNTGPGSNQAQRV 529
Query: 1573 NWPGFHVIGRKEAEPFTAGPFIDGAMWLKYTGAPH 1677
WPG + ++A +T F+ G W+ T P+
Sbjct: 530 KWPGIKKLTPQDALLYTGDRFLRGDTWIPQTQVPY 564
Score = 38.5 bits (88), Expect(2) = e-105
Identities = 18/59 (30%), Positives = 39/59 (66%)
Frame = +2
Query: 152 KSVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQSKSIGEAK 328
K+V+++CAPT +K++C +L A+ +++P ++ KV ++S+ ++E K+ G+ K
Sbjct: 49 KAVQAVCAPTDFKDTCVNSLMGASPDSDDPVDLIKLGFKVTIKSINESLE--KASGDIK 105
>sptr|Q9FF78|Q9FF78 Pectinesterase.
Length = 564
Score = 368 bits (945), Expect(2) = e-105
Identities = 202/456 (44%), Positives = 276/456 (60%), Gaps = 5/456 (1%)
Frame = +1
Query: 322 GQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDT 501
G+ D+ T +A C +L+ A D L + + DDL TWL+ V T+ +T
Sbjct: 121 GEHMDNATSAAMGACVELIGLAVDQLNETMTSS-------LKNFDDLRTWLSSVGTYQET 173
Query: 502 CVDGFVDEK---LKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLS 672
C+D V+ L + ++N+TE++SNALAI LG I + K RRRLL
Sbjct: 174 CMDALVEANKPSLTTFGENHLKNSTEMTSNALAIITWLGKIADTV------KFRRRRLLE 227
Query: 673 SEQDEKGWPVWMRSPERKLLASGN-QPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYV 849
+ + R+LL SG+ + K +VAKDGSG++++I +A+ V + ++ +
Sbjct: 228 TGNAKVVVADLPMMEGRRLLESGDLKKKATIVVAKDGSGKYRTIGEALAEVEEKNEKPTI 287
Query: 850 IYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFIC 1029
IYVK G+Y E V V K K N+ M GDG ++ V+ +F DG T +TATF+V GF+
Sbjct: 288 IYVKKGVYLENVRVEKTKWNVVMVGDGQSKTIVSAGLNFIDGTPTFETATFAVFGKGFMA 347
Query: 1030 KNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDF 1209
++MGF NTAG +HQAVAL V DL+ FY C DAFQDT+Y HA+RQF+R+CV+ GT+DF
Sbjct: 348 RDMGFINTAGPAKHQAVALMVSADLSVFYKCTMDAFQDTMYAHAQRQFYRDCVILGTVDF 407
Query: 1210 IFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFK 1389
IFGN+A VFQ C I+ RRPM QQN++TA GR DPN +G+ I NC + P L
Sbjct: 408 IFGNAAVVFQKCEILPRRPMKGQQNTITAQGRKDPNQNTGISIHNCTIKPLDNL----TD 463
Query: 1390 IPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKR 1569
I ++LGRPWK+FS VIM+S + F+ P+G++PW GD A T++YAEY N GPGA T R
Sbjct: 464 IQTFLGRPWKDFSTTVIMKSFMDKFINPKGWLPWTGDTAPDTIFYAEYLNSGPGASTKNR 523
Query: 1570 VNWPGFHV-IGRKEAEPFTAGPFIDGAMWLKYTGAP 1674
V W G + +KEA FT PFIDG WL T P
Sbjct: 524 VKWQGLKTSLTKKEANKFTVKPFIDGNNWLPATKVP 559
Score = 38.9 bits (89), Expect(2) = e-105
Identities = 19/50 (38%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Frame = +2
Query: 155 SVKSLCAPTLYKESCEKTLSQATNGT-ENPKEVFHSVAKVALESVQTAVE 301
SVK+LC TL+KE C +TL A N + +P+E+F KV + + ++
Sbjct: 67 SVKALCDVTLHKEKCFETLGSAPNASRSSPEELFKYAVKVTITELSKVLD 116
>sptr|Q9FVF9|Q9FVF9 Putative pectin methylesterase 3 (EC 3.1.1.11)
(Pectinesterase).
Length = 555
Score = 381 bits (979), Expect = e-104
Identities = 207/454 (45%), Positives = 274/454 (60%), Gaps = 6/454 (1%)
Frame = +1
Query: 343 TESAREDCKKLLEDAADDLRGMLEMAGG--DIKVLFSRSDDLETWLTGVMTFMDTCVDGF 516
+ SA +DC + + + D+L L + K + +DDL+T L+ T +TC+DGF
Sbjct: 108 SRSALKDCVETMSTSLDELHVALAELDEYPNKKSITRHADDLKTLLSAATTNQETCLDGF 167
Query: 517 VDEKLKADMHSVVRNATELSSNALA--ITNSLGGILKKMDLGMFSKDSRRRLLSSEQDEK 690
+ D VR E + N+LG I+ + M S + +++E
Sbjct: 168 SHD----DSEKKVRKTLETGPVRVEKMCGNALGMIVNMTETDMASATNA---VNTEGGSS 220
Query: 691 G-WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAG 867
G WP+WM+ +R+LL +G PN +VA DGSG+++ + +AV A P RYVI +KAG
Sbjct: 221 GSWPIWMKGGDRRLLQAGTTVTPNVVVAADGSGKYRRVSEAVAAAPSKSSKRYVIRIKAG 280
Query: 868 LYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFH 1047
+Y E V VPKDK NI GDG + +TG K+ DG TT +AT +V GF+ +++ F
Sbjct: 281 IYRENVEVPKDKTNIMFVGDGRSNTIITGNKNVVDGSTTFNSATVAVVGQGFLARDITFQ 340
Query: 1048 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSA 1227
NTAG +HQAVALRV DLAAFY C F A+QDTLYVH+ RQFF NC+V GT+DFIFGNSA
Sbjct: 341 NTAGPSKHQAVALRVGADLAAFYRCDFLAYQDTLYVHSNRQFFINCLVVGTVDFIFGNSA 400
Query: 1228 AVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 1407
AVFQNC I RRP Q+N +TAHGRTDPN +G+VIQ R+ L + +YLG
Sbjct: 401 AVFQNCDIHARRPNPGQKNMLTAHGRTDPNQNTGIVIQKSRIAATSDLQSVKGSFGTYLG 460
Query: 1408 RPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGF 1587
RPWK ++R VIM+STI+D V P G+ W+G+FAL TL+Y E+ N G G+G + RV W G
Sbjct: 461 RPWKAYARTVIMQSTISDVVHPAGWHEWDGNFALNTLFYGEHKNSGAGSGVNGRVKWKGH 520
Query: 1588 HVIGR-KEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
VI EA FT G FI G WL T P LG
Sbjct: 521 KVISSDAEAAGFTPGRFIAGGSWLGSTTFPFTLG 554
>sptr|Q84W31|Q84W31 Hypothetical protein.
Length = 564
Score = 364 bits (935), Expect(2) = e-104
Identities = 200/456 (43%), Positives = 275/456 (60%), Gaps = 5/456 (1%)
Frame = +1
Query: 322 GQGSDSMTESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDT 501
G+ D+ T +A C +L+ A D L + + DDL TWL+ V T+ +T
Sbjct: 121 GEHMDNATSAAMGACVELIGLAVDQLNETMTSS-------LKNFDDLRTWLSSVGTYQET 173
Query: 502 CVDGFVDEK---LKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLS 672
C+D V+ L + ++N+TE++SNALAI LG I + K RRRLL
Sbjct: 174 CMDALVEANKPSLTTFGENHLKNSTEMTSNALAIITWLGKIADTV------KFRRRRLLE 227
Query: 673 SEQDEKGWPVWMRSPERKLLASGN-QPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYV 849
+ + R+LL SG+ + K +VAKDGSG++++I +A+ V + ++ +
Sbjct: 228 TGNAKVVVADLPMMEGRRLLESGDLKKKATIVVAKDGSGKYRTIGEALAEVEEKNEKPTI 287
Query: 850 IYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFIC 1029
IYVK G+Y E V V K K N+ M GDG ++ V+ +F DG T +TATF+V GF+
Sbjct: 288 IYVKKGVYLENVRVEKTKWNVVMVGDGQSKTIVSAGLNFIDGTPTFETATFAVFGKGFMA 347
Query: 1030 KNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDF 1209
++MGF NTAG +HQAVAL V DL+ FY C DAFQDT+Y HA+RQF+R+CV+ GT+DF
Sbjct: 348 RDMGFINTAGPAKHQAVALMVSADLSVFYKCTMDAFQDTMYAHAQRQFYRDCVILGTVDF 407
Query: 1210 IFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFK 1389
IFGN+A VFQ C I+ RRPM QQN++TA GR DPN +G+ I NC + P L +
Sbjct: 408 IFGNAAVVFQKCEILPRRPMKGQQNTITAQGRKDPNQNTGISIHNCTIKPLDNLTDTQ-- 465
Query: 1390 IPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKR 1569
++L RPWK+FS VIM+S + F+ P+G++PW GD A T++YAEY N GPGA T R
Sbjct: 466 --TFLDRPWKDFSTTVIMKSFMDKFINPKGWLPWTGDTAPDTIFYAEYLNSGPGASTKNR 523
Query: 1570 VNWPGFHV-IGRKEAEPFTAGPFIDGAMWLKYTGAP 1674
V W G + +KEA FT PFIDG WL T P
Sbjct: 524 VKWQGLKTSLTKKEANKFTVKPFIDGNNWLPATKVP 559
Score = 38.9 bits (89), Expect(2) = e-104
Identities = 19/50 (38%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Frame = +2
Query: 155 SVKSLCAPTLYKESCEKTLSQATNGT-ENPKEVFHSVAKVALESVQTAVE 301
SVK+LC TL+KE C +TL A N + +P+E+F KV + + ++
Sbjct: 67 SVKALCDVTLHKEKCFETLGSAPNASRSSPEELFKYAVKVTITELSKVLD 116
>sptr|Q8GS16|Q8GS16 Putative thermostable pectinesterase.
Length = 631
Score = 374 bits (960), Expect(2) = e-104
Identities = 201/458 (43%), Positives = 285/458 (62%), Gaps = 11/458 (2%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDL-RGMLEMAG----GDIKVLFSRSDDLETWLTGVMTFMDTCVD 510
+++ DC +++++ D+L + E+ G + K + ++D+L+ ++ MT +TC+D
Sbjct: 180 KTSLHDCLEMVDETLDELYKTEHELQGYPAAANNKSIAEQADELKILVSAAMTNQETCLD 239
Query: 511 GF----VDEKLKADMHSVVRNATELSSNALA-ITNSLGGILKKMDLGMFSKDSRRRLLSS 675
GF D+K++ ++ + + SNALA I N G + K + +SK RRL
Sbjct: 240 GFSHERADKKIREELMEGQMHVFHMCSNALAMIKNMTDGDIGKDIVDHYSK--ARRL--- 294
Query: 676 EQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIY 855
DE WP W+ + +R+LL + P+ VA DGSG + ++ AV A P+G RY+I
Sbjct: 295 -DDETKWPEWLSAGDRRLLQA-TTVVPDVTVAADGSGNYLTVAAAVAAAPEGSSRRYIIR 352
Query: 856 VKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKN 1035
+KAG Y E V VPK K+N+ GDG + +TG ++ DG TT +AT +V GF+ ++
Sbjct: 353 IKAGEYRENVEVPKKKINLMFIGDGRTTTIITGSRNVVDGSTTFNSATVAVVGDGFLARD 412
Query: 1036 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIF 1215
+ F NTAG +HQAVALRV DL+AFY C A+QDTLYVH+ RQF+ +C+++GT+DFIF
Sbjct: 413 ITFQNTAGPSKHQAVALRVGSDLSAFYRCDMLAYQDTLYVHSLRQFYTSCIIAGTVDFIF 472
Query: 1216 GNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 1395
GN+AAVFQNC I RRP NQ+N VTA GR DPN +G+VIQ CR+ L +
Sbjct: 473 GNAAAVFQNCDIHARRPNPNQRNMVTAQGRDDPNQNTGIVIQKCRIGATSDLLAVKGSFE 532
Query: 1396 SYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVN 1575
+YLGRPWK +SR V+M+S I+D + P G+ W+G+FAL TL+YAEY N G GA TS RV
Sbjct: 533 TYLGRPWKRYSRTVVMQSDISDVINPAGWYEWSGNFALDTLFYAEYQNTGAGADTSNRVK 592
Query: 1576 WPGFHVI-GRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
W F VI EA+ +TA FI G+ WL TG P LG
Sbjct: 593 WSTFKVITSAAEAQTYTAANFIAGSTWLGSTGFPFSLG 630
Score = 28.1 bits (61), Expect(2) = e-104
Identities = 15/48 (31%), Positives = 24/48 (50%), Gaps = 4/48 (8%)
Frame = +2
Query: 158 VKSLCAPTLYKESCEKTLS----QATNGTENPKEVFHSVAKVALESVQ 289
++S C+ TLY E C LS A + +PK+V + + +VQ
Sbjct: 111 IRSSCSATLYPELCFSALSAAPAAAVSKVNSPKDVIRLSLNLTITAVQ 158
>sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase).
Length = 579
Score = 377 bits (969), Expect(2) = e-104
Identities = 198/452 (43%), Positives = 280/452 (61%), Gaps = 7/452 (1%)
Frame = +1
Query: 352 AREDCKKLLEDAADDLRGMLEMAG--GDIKVLFSRSDDLETWLTGVMTFMDTCVDGF--- 516
A DC + +++ D+L ++ + K L + +DDL+T ++ +T +TC+DGF
Sbjct: 131 ALHDCLETIDETLDELHTAIKDLELYPNKKSLKAHADDLKTLISSAITNQETCLDGFSHD 190
Query: 517 -VDEKLKADMHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSEQDEKG 693
D+K++ + ++ ++ SNALA+ ++ + + + R+L +D
Sbjct: 191 DADKKVRKALLKGQKHVEKMCSNALAMICNMTDTDIANEQKLKGTTTNRKL---REDNSE 247
Query: 694 WPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAGLY 873
WP W+ + +R+LL S +P+ +VA DGSG FK++ +AV P+ RYVI +KAG+Y
Sbjct: 248 WPEWLSAGDRRLLQSSTV-RPDVVVAADGSGNFKTVSEAVAKAPEKSSKRYVIRIKAGVY 306
Query: 874 DEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFHNT 1053
E V VPK K NI GDG + +TG ++ DG TT +AT + F+ +++ F NT
Sbjct: 307 RENVDVPKKKTNIMFMGDGRSNTIITGSRNVKDGSTTFHSATVAAVGEKFLARDITFQNT 366
Query: 1054 AGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSAAV 1233
AGA +HQAVALRV DL+AFY C A+QD+LYVH+ RQ+F C+++GT+DFIFGN+AAV
Sbjct: 367 AGAAKHQAVALRVGSDLSAFYRCDILAYQDSLYVHSNRQYFVQCLIAGTVDFIFGNAAAV 426
Query: 1234 FQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLGRP 1413
QNC I RRP Q+N VTA GR+DPN +G+VIQ CR+ L P + P+YLGRP
Sbjct: 427 LQNCDIHARRPGSGQKNMVTAQGRSDPNQNTGIVIQKCRIGATSDLRPVQKSFPTYLGRP 486
Query: 1414 WKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGFHV 1593
WKE+SR VIM+S+I D + G+ WNG+FAL TL+Y EY N G GAGTS RV W GF V
Sbjct: 487 WKEYSRTVIMQSSITDVINSAGWHEWNGNFALNTLFYGEYQNTGAGAGTSGRVKWKGFKV 546
Query: 1594 I-GRKEAEPFTAGPFIDGAMWLKYTGAPHILG 1686
I EA+ +T G FI G WL TG P LG
Sbjct: 547 ITSATEAQAYTPGRFIAGGSWLSSTGFPFSLG 578
Score = 24.3 bits (51), Expect(2) = e-104
Identities = 13/47 (27%), Positives = 24/47 (51%), Gaps = 3/47 (6%)
Frame = +2
Query: 158 VKSLCAPTLYKESCEKTLSQATNGTE---NPKEVFHSVAKVALESVQ 289
VKS C TL+ E C T++ ++ ++ + K+V + +VQ
Sbjct: 62 VKSACENTLHPELCYSTIASVSDFSKKVTSQKDVIELSLNITCRAVQ 108
>sptrnew|AAM91439|AAM91439 At1g53830/T18A20_6.
Length = 587
Score = 372 bits (956), Expect(2) = e-103
Identities = 205/463 (44%), Positives = 282/463 (60%), Gaps = 18/463 (3%)
Frame = +1
Query: 349 SAREDCKKLLEDAADDLRGMLEMAGGDI------KVLFSRSDDLETWLTGVMTFMDTCVD 510
+A DC + +++ D+L +E D+ K L +DDL+T ++ +T TC+D
Sbjct: 128 TALHDCLETIDETLDELHVAVE----DLHQYPKQKSLRKHADDLKTLISSAITNQGTCLD 183
Query: 511 GF----VDEKLKADMHSVVRNATELSSNALA-ITNSLGGILKKMDL----GMFSKDSRRR 663
GF D K++ + + + SNALA I N + +L F+ ++ R+
Sbjct: 184 GFSYDDADRKVRKALLKGQVHVEHMCSNALAMIKNMTETDIANFELRDKSSTFTNNNNRK 243
Query: 664 L--LSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQ 837
L ++ + D GWP W+ +R+LL G+ K +A VA DGSG F ++ AV A P+
Sbjct: 244 LKEVTGDLDSDGWPKWLSVGDRRLL-QGSTIKADATVADDGSGDFTTVAAAVAAAPEKSN 302
Query: 838 GRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEAS 1017
R+VI++KAG+Y E V V K K NI GDG ++ +TG ++ DG TT +AT +
Sbjct: 303 KRFVIHIKAGVYRENVEVTKKKTNIMFLGDGRGKTIITGSRNVVDGSTTFHSATVAAVGE 362
Query: 1018 GFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSG 1197
F+ +++ F NTAG +HQAVALRV D +AFY C A+QDTLYVH+ RQFF C ++G
Sbjct: 363 RFLARDITFQNTAGPSKHQAVALRVGSDFSAFYQCDMFAYQDTLYVHSNRQFFVKCHITG 422
Query: 1198 TIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFP 1377
T+DFIFGN+AAV Q+C I RRP Q+N VTA GR+DPN +G+VIQNCR+ L
Sbjct: 423 TVDFIFGNAAAVLQDCDINARRPNSGQKNMVTAQGRSDPNQNTGIVIQNCRIGGTSDLLA 482
Query: 1378 DRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAG 1557
+ P+YLGRPWKE+SR VIM+S I+D ++PEG+ W+G FAL TL Y EY NRG GAG
Sbjct: 483 VKGTFPTYLGRPWKEYSRTVIMQSDISDVIRPEGWHEWSGSFALDTLTYREYLNRGGGAG 542
Query: 1558 TSKRVNWPGFHVI-GRKEAEPFTAGPFIDGAMWLKYTGAPHIL 1683
T+ RV W G+ VI EA+PFTAG FI G WL TG P L
Sbjct: 543 TANRVKWKGYKVITSDTEAQPFTAGQFIGGGGWLASTGFPFSL 585
Score = 28.9 bits (63), Expect(2) = e-103
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Frame = +2
Query: 143 TSGKSVKSLCAPTLYKESCEKTLSQATNGTE--NPKEVFHSVAKVALESVQ 289
TS +KS+C+ TLY E C ++ AT G E + KEV + + ++V+
Sbjct: 57 TSHAILKSVCSSTLYPELCFSAVA-ATGGKELTSQKEVIEASLNLTTKAVK 106
>sw|Q42534|PME2_ARATH Pectinesterase 2 precursor (EC 3.1.1.11) (Pectin
methylesterase 2) (PE 2).
Length = 587
Score = 372 bits (956), Expect(2) = e-103
Identities = 205/463 (44%), Positives = 282/463 (60%), Gaps = 18/463 (3%)
Frame = +1
Query: 349 SAREDCKKLLEDAADDLRGMLEMAGGDI------KVLFSRSDDLETWLTGVMTFMDTCVD 510
+A DC + +++ D+L +E D+ K L +DDL+T ++ +T TC+D
Sbjct: 128 TALHDCLETIDETLDELHVAVE----DLHQYPKQKSLRKHADDLKTLISSAITNQGTCLD 183
Query: 511 GF----VDEKLKADMHSVVRNATELSSNALA-ITNSLGGILKKMDL----GMFSKDSRRR 663
GF D K++ + + + SNALA I N + +L F+ ++ R+
Sbjct: 184 GFSYDDADRKVRKALLKGQVHVEHMCSNALAMIKNMTETDIANFELRDKSSTFTNNNNRK 243
Query: 664 L--LSSEQDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQ 837
L ++ + D GWP W+ +R+LL G+ K +A VA DGSG F ++ AV A P+
Sbjct: 244 LKEVTGDLDSDGWPKWLSVGDRRLL-QGSTIKADATVADDGSGDFTTVAAAVAAAPEKSN 302
Query: 838 GRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEAS 1017
R+VI++KAG+Y E V V K K NI GDG ++ +TG ++ DG TT +AT +
Sbjct: 303 KRFVIHIKAGVYRENVEVTKKKTNIMFLGDGRGKTIITGSRNVVDGSTTFHSATVAAVGE 362
Query: 1018 GFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSG 1197
F+ +++ F NTAG +HQAVALRV D +AFY C A+QDTLYVH+ RQFF C ++G
Sbjct: 363 RFLARDITFQNTAGPSKHQAVALRVGSDFSAFYQCDMFAYQDTLYVHSNRQFFVKCHITG 422
Query: 1198 TIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFP 1377
T+DFIFGN+AAV Q+C I RRP Q+N VTA GR+DPN +G+VIQNCR+ L
Sbjct: 423 TVDFIFGNAAAVLQDCDINARRPNSGQKNMVTAQGRSDPNQNTGIVIQNCRIGGTSDLLA 482
Query: 1378 DRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAG 1557
+ P+YLGRPWKE+SR VIM+S I+D ++PEG+ W+G FAL TL Y EY NRG GAG
Sbjct: 483 VKGTFPTYLGRPWKEYSRTVIMQSDISDVIRPEGWHEWSGSFALDTLTYREYLNRGGGAG 542
Query: 1558 TSKRVNWPGFHVI-GRKEAEPFTAGPFIDGAMWLKYTGAPHIL 1683
T+ RV W G+ VI EA+PFTAG FI G WL TG P L
Sbjct: 543 TANRVKWKGYKVITSDTEAQPFTAGQFIGGGGWLASTGFPFSL 585
Score = 28.9 bits (63), Expect(2) = e-103
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Frame = +2
Query: 143 TSGKSVKSLCAPTLYKESCEKTLSQATNGTE--NPKEVFHSVAKVALESVQ 289
TS +KS+C+ TLY E C ++ AT G E + KEV + + ++V+
Sbjct: 57 TSHAILKSVCSSTLYPELCFSAVA-ATGGKELTSQKEVIEASLNLTTKAVK 106
>sptr|Q94FS5|Q94FS5 Putative pectin methylesterase LuPME5 (EC
3.1.1.11).
Length = 553
Score = 377 bits (968), Expect = e-103
Identities = 197/453 (43%), Positives = 277/453 (61%), Gaps = 7/453 (1%)
Frame = +1
Query: 346 ESAREDCKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGFV-- 519
++A DC +L + D+ +E G K R +DL+T L+ MT +TC+DGF
Sbjct: 104 KAALADCIELCGETMDEPVKTIEELHGKKKSAAERGEDLKTLLSAAMTNQETCLDGFSHD 163
Query: 520 --DEKLKADMHSVVRNATELSSNALAITNSLGG--ILKKMDLGMFSKDSRRRLLSSEQDE 687
D+K++ + + N + SN+LA+ ++ + ++ F + R+ ++E
Sbjct: 164 KGDKKVRELLAAGQTNVGRMCSNSLAMVENITEEEVFREGKTASFLSEGRKM----GEEE 219
Query: 688 KGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYVKAG 867
GWP W+ + +R+LL +G PN +VA DGSG F+++ QAV A P+G RYVI +KAG
Sbjct: 220 DGWPRWISAGDRRLLQAGTVT-PNVVVAADGSGNFRTVSQAVAAAPEGSTSRYVIRIKAG 278
Query: 868 LYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNMGFH 1047
+Y E ++VPK K N+ GDG + +TG + DG TT +AT +V F+ +++ F
Sbjct: 279 VYRETLVVPKKKTNLMFVGDGRTSTIITGSMNVVDGSTTFNSATVAVVGDRFMARDLTFQ 338
Query: 1048 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFGNSA 1227
NTAG +HQAVALRV D AFY C A+QDTLYVH+ RQF+ +C ++GT+DFIFGN+A
Sbjct: 339 NTAGPSKHQAVALRVNADFTAFYRCDMLAYQDTLYVHSLRQFYVSCFIAGTVDFIFGNAA 398
Query: 1228 AVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 1407
V QNC I RRP Q+N VTA GR DPN +G+VIQ CR+ Q L + + SYLG
Sbjct: 399 VVLQNCDIHARRPNSGQRNMVTAQGRDDPNQNTGIVIQKCRIGATQDLLQVQSSVESYLG 458
Query: 1408 RPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNWPGF 1587
RPWK +SR VIM++ I++ ++P G+ W+G+FAL TL Y EY N G G+GTS RV W G+
Sbjct: 459 RPWKMYSRTVIMQTDISNVIRPAGWFMWDGNFALATLTYREYANTGAGSGTSGRVRWGGY 518
Query: 1588 HVI-GRKEAEPFTAGPFIDGAMWLKYTGAPHIL 1683
VI EA+PF FI GA WL TG P L
Sbjct: 519 KVITSASEAQPFAPRSFIGGASWLPATGFPFSL 551
>sptr|Q9SC90|Q9SC90 Pectin methyl-esterase PER precursor (EC
3.1.1.11).
Length = 602
Score = 370 bits (949), Expect(2) = e-103
Identities = 196/461 (42%), Positives = 277/461 (60%), Gaps = 9/461 (1%)
Frame = +1
Query: 328 GSDSMTESAREDCKKLLEDAADDLRGMLEMAGG-DIKVLFSRSDDLETWLTGVMTFMDTC 504
G D MT+ A E C ++L+ A D + + DI + S DL+ WLTG ++ TC
Sbjct: 99 GQDKMTKQAMEVCNEVLDYAVDGIHKSVGAVDKFDINKIHEYSYDLKVWLTGTLSHQQTC 158
Query: 505 VDGFVDEKLKAD--MHSVVRNATELSSNALAITNSLGGILKKMDLGMFSKDSRRRLLSSE 678
+DGF + KA M + + +LSSNA+ + +++ + +++RRLLS +
Sbjct: 159 LDGFANTTTKAGETMARALNTSIQLSSNAIDMVDAVYDLT----------NAKRRLLSLD 208
Query: 679 QDEKGWPVWMRSPERKLLASGNQPKPNAIVAKDGSGQFKSIQQAVDAVPKGHQGRYVIYV 858
G+P+W+ +R+LLA KPN +VA+DGSGQFK++ A+ VP + +VIYV
Sbjct: 209 N---GYPLWVSEGQRRLLAEATV-KPNVVVAQDGSGQFKTLTDAIKTVPANNAQNFVIYV 264
Query: 859 KAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFADGITTMKTATFSVEASGFICKNM 1038
K G+Y+E V VPKD + + GDGP +++ TG ++ADG+ TATF V F+ K++
Sbjct: 265 KEGVYNETVNVPKDMAFVTIIGDGPAKTKFTGSLNYADGLLPYNTATFGVNGENFMAKDI 324
Query: 1039 GFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVVSGTIDFIFG 1218
NTAG E+HQAVALRV D A FYNC+ D +Q TL+ ++RQF+R+C +SGTID I+G
Sbjct: 325 SIENTAGPEKHQAVALRVTADKAIFYNCQIDGYQATLFAESQRQFYRDCSISGTIDMIYG 384
Query: 1219 NSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPS 1398
++ AVFQNC +I R+P++ QQ V A GRT + SG V Q+C + ++ KI +
Sbjct: 385 DAFAVFQNCKLIVRKPLEEQQCFVAADGRTKSDSSSGFVFQSCHFTGEPEVAKIDPKI-A 443
Query: 1399 YLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVNW 1578
YLGRPWK +S++VIM+S I D PEGYMPW G T + EYNN+GPGA TSKRV W
Sbjct: 444 YLGRPWKSYSKVVIMDSNIDDIFDPEGYMPWMGSAFKDTCTFYEYNNKGPGADTSKRVKW 503
Query: 1579 PGFHVIGRKEAEPFTAGPFI------DGAMWLKYTGAPHIL 1683
PG I EA F G F D W+ +G P+ L
Sbjct: 504 PGVKSISSTEAAAFYPGKFFEIANATDRDTWIVKSGVPYSL 544
Score = 29.3 bits (64), Expect(2) = e-103
Identities = 14/56 (25%), Positives = 28/56 (50%)
Frame = +2
Query: 155 SVKSLCAPTLYKESCEKTLSQATNGTENPKEVFHSVAKVALESVQTAVEQSKSIGE 322
+V+ LC T Y+++C ++L++A T K++ + E + + S I E
Sbjct: 42 NVQILCESTQYQQTCHQSLAKAPAETAGVKDLIKAAFSATSEELLKHINSSSLIQE 97
>sptr|O80722|O80722 Putative pectinesterase.
Length = 588
Score = 375 bits (964), Expect = e-102
Identities = 202/481 (41%), Positives = 288/481 (59%), Gaps = 40/481 (8%)
Frame = +1
Query: 364 CKKLLEDAADDLRGMLEMAGGDIKVLFSRSDDLETWLTGVMTFMDTCVDGFVDEKLKADM 543
CK++ A +DL ++E G D+ + S+ D L+ WL GV + C+D ++ L+ +
Sbjct: 110 CKRVFMYALEDLATIIEEMGEDLSQIGSKIDQLKQWLIGVYNYQTDCLDDIEEDDLRKAI 169
Query: 544 HSVVRNATELSSNALAITNSLGGILKKM------------------------DLGMFSKD 651
+ N+ L++NA+ I +++ + K+ + + D
Sbjct: 170 GEGIANSKILTTNAIDIFHTVVSAMAKINNKVDDLKNMTGGIPTPGAPPVVDESPVADPD 229
Query: 652 SRRRLLSSEQDEKGWPVWMRSPERKLLAS-----------GNQPKPNAIVAKDGSGQFKS 798
R L + DE G P W+ +RKL+A G + + N +VAKDGSGQFK+
Sbjct: 230 GPARRLLEDIDETGIPTWVSGADRKLMAKAGRGRRGGRGGGARVRTNFVVAKDGSGQFKT 289
Query: 799 IQQAVDAVPKGHQGRYVIYVKAGLYDEIVMVPKDKVNIFMYGDGPKQSRVTGRKSFA--D 972
+QQAVDA P+ ++GR +IY+KAGLY E V++PK K NIFM+GDG +++ ++ +S A
Sbjct: 290 VQQAVDACPENNRGRCIIYIKAGLYREQVIIPKKKNNIFMFGDGARKTVISYNRSVALSR 349
Query: 973 GITTMKTATFSVEASGFICKNMGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLY 1152
G TT +AT VE+ GF+ K MGF NTAG HQA A+RV GD A +NCRFD +QDTLY
Sbjct: 350 GTTTSLSATVQVESEGFMAKWMGFKNTAGPMGHQAAAIRVNGDRAVIFNCRFDGYQDTLY 409
Query: 1153 VHARRQFFRNCVVSGTIDFIFGNSAAVFQNCLIITRRPMDNQQNSVTAHG-RTDPNMKSG 1329
V+ RQF+RNCVVSGT+DFIFG SA V QN LI+ R+ Q N+VTA G MK G
Sbjct: 410 VNNGRQFYRNCVVSGTVDFIFGKSATVIQNTLIVVRKGSKGQYNTVTADGNELGLGMKIG 469
Query: 1330 LVIQNCRLVPDQKLFPDRFKIPSYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFAL 1509
+V+QNCR+VPD+KL P+R + +YLGRPWK+FS VIM + + D ++PEG+ W+G+
Sbjct: 470 IVLQNCRIVPDRKLTPERLTVATYLGRPWKKFSTTVIMSTEMGDLIRPEGWKIWDGESFH 529
Query: 1510 KTLYYAEYNNRGPGAGTSKRVNWPGFHVIGRKEAE--PFTAGPFIDGAMWLKYTGAPHIL 1683
K+ Y EYNNRGPGA ++RVNW + R AE FTA ++ W++ P +
Sbjct: 530 KSCRYVEYNNRGPGAFANRRVNWA---KVARSAAEVNGFTAANWLGPINWIQEANVPVTI 586
Query: 1684 G 1686
G
Sbjct: 587 G 587
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,291,829,513
Number of Sequences: 1395590
Number of extensions: 27649182
Number of successful extensions: 103524
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 92650
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 102816
length of database: 442,889,342
effective HSP length: 130
effective length of database: 261,462,642
effective search space used: 137267887050
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

