BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
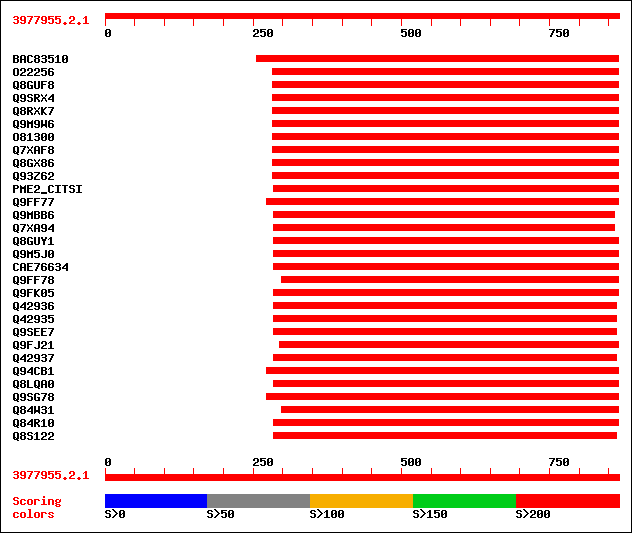
Query= 3977955.2.1
(867 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|BAC83510|BAC83510 Putative pectin methylesterase. 349 3e-95
sptr|O22256|O22256 Putative pectinesterase. 216 5e-55
sptr|Q8GUF8|Q8GUF8 At2g47550/T30B22.15. 216 5e-55
sptr|Q9SRX4|Q9SRX4 F22D16.20 protein. 215 6e-55
sptr|Q8RXK7|Q8RXK7 Hypothetical protein. 215 6e-55
sptr|Q9M9W6|Q9M9W6 Putative pectinesterase. 215 8e-55
sptr|O81300|O81300 T14P8.14 protein (AT4G02330 protein) (Hypothe... 214 1e-54
sptr|Q7XAF8|Q7XAF8 Papillar cell-specific pectin methylesterase-... 213 2e-54
sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610). 213 3e-54
sptr|Q93Z62|Q93Z62 At2g47550/T30B22.15. 212 5e-54
sw|O04887|PME2_CITSI Pectinesterase 2.1 precursor (EC 3.1.1.11) ... 211 1e-53
sptr|Q9FF77|Q9FF77 Pectinesterase. 209 6e-53
sptr|Q9MBB6|Q9MBB6 Pectin methylesterase. 209 6e-53
sptr|Q7XA94|Q7XA94 Pectinesterase (EC 3.1.1.11) (Fragment). 207 1e-52
sptr|Q8GUY1|Q8GUY1 Methyl pectinesterase (Fragment). 207 2e-52
sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC 3.1.1... 206 3e-52
sptrnew|CAE76634|CAE76634 Pectin methylesterase (EC 3.1.1.11) (F... 206 5e-52
sptr|Q9FF78|Q9FF78 Pectinesterase. 205 6e-52
sptr|Q9FK05|Q9FK05 Pectinesterase. 205 6e-52
sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11) (Pectines... 205 8e-52
sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11) (Pectines... 205 8e-52
sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11) (Pectine... 205 8e-52
sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11) (AT5g4918... 205 8e-52
sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment). 205 8e-52
sptr|Q94CB1|Q94CB1 Putative pectinesterase. 203 2e-51
sptr|Q8LQA0|Q8LQA0 Putative pectinesterase. 203 2e-51
sptr|Q9SG78|Q9SG78 Putative pectinesterase. 203 2e-51
sptr|Q84W31|Q84W31 Hypothetical protein. 202 4e-51
sptr|Q84R10|Q84R10 Putative pectinesterase. 202 4e-51
sptr|Q8S122|Q8S122 Putative pectin esterase. 202 4e-51
>sptrnew|BAC83510|BAC83510 Putative pectin methylesterase.
Length = 566
Score = 349 bits (896), Expect = 3e-95
Identities = 157/204 (76%), Positives = 181/204 (88%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
YQC FDGYQDTLY HAQRQFFRDC V+GTIDFIFGNSQVVLQNCL+QPRKPM NQ NIIT
Sbjct: 364 YQCEFDGYQDTLYPHAQRQFFRDCTVSGTIDFIFGNSQVVLQNCLLQPRKPMDNQVNIIT 423
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGRR+KRS GGTV+HNCT+ PHPD E+ K++TYLARPWKEYSRTI++QN+IG +D
Sbjct: 424 AQGRREKRSAGGTVIHNCTVAPHPDL-EKFTDKVKTYLARPWKEYSRTIFVQNEIGAVVD 482
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWLEWNG+F L+TL+YAEVDN G GADMSKRAKW G++++TY++ QKEFTVE FIQGQ
Sbjct: 483 PVGWLEWNGNFALDTLYYAEVDNHGPGADMSKRAKWKGVQSLTYQDVQKEFTVEAFIQGQ 542
Query: 326 QFIPKFGVPFIPGLLPQEQKGRTH 255
+FIPKFGVP+IPGLLPQ Q+GR H
Sbjct: 543 EFIPKFGVPYIPGLLPQTQQGRMH 566
>sptr|O22256|O22256 Putative pectinesterase.
Length = 560
Score = 216 bits (549), Expect = 5e-55
Identities = 99/195 (50%), Positives = 131/195 (67%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C V GT+DFIFGN+ VVLQNC + PR+P Q+N +T
Sbjct: 367 YSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAAVVLQNCNLYPRQPRKGQSNEVT 426
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D GT +H CTI P D + ++TYL RPWKEYSRT+ +Q I GF++
Sbjct: 427 AQGRTDPNQNTGTAIHGCTIRPADDL-ATSNYTVKTYLGRPWKEYSRTVVMQTYIDGFLE 485
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW W+GDF L TL+YAE +N G G+D + R W G + +A FTV F+ G+
Sbjct: 486 PSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGYHVINATDA-SNFTVTNFLVGE 544
Query: 326 QFIPKFGVPFIPGLL 282
+I + GVPF+ GL+
Sbjct: 545 GWIGQTGVPFVGGLI 559
>sptr|Q8GUF8|Q8GUF8 At2g47550/T30B22.15.
Length = 345
Score = 216 bits (549), Expect = 5e-55
Identities = 99/195 (50%), Positives = 131/195 (67%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C V GT+DFIFGN+ VVLQNC + PR+P Q+N +T
Sbjct: 152 YSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAAVVLQNCNLYPRQPRKGQSNEVT 211
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D GT +H CTI P D + ++TYL RPWKEYSRT+ +Q I GF++
Sbjct: 212 AQGRTDPNQNTGTAIHGCTIRPADDL-ATSNYTVKTYLGRPWKEYSRTVVMQTYIDGFLE 270
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW W+GDF L TL+YAE +N G G+D + R W G + +A FTV F+ G+
Sbjct: 271 PSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGYHVINATDA-SNFTVTNFLVGE 329
Query: 326 QFIPKFGVPFIPGLL 282
+I + GVPF+ GL+
Sbjct: 330 GWIGQTGVPFVGGLI 344
>sptr|Q9SRX4|Q9SRX4 F22D16.20 protein.
Length = 579
Score = 215 bits (548), Expect = 6e-55
Identities = 102/195 (52%), Positives = 130/195 (66%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C V GT+DFIFGN+ VV QNC + PRKPM NQ N IT
Sbjct: 386 YSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAAVVFQNCNLYPRKPMPNQFNAIT 445
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D GT + NCTI+P D + ++TYL RPWKEYSRT+Y+Q+ I GF++
Sbjct: 446 AQGRSDPNQNTGTSIQNCTIKPADDL-VSSNYTVKTYLGRPWKEYSRTVYMQSYIDGFVE 504
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW EWNGDF L TL+YAE +N G G++ + R W G + +A FTV
Sbjct: 505 PVGWREWNGDFALSTLYYAEYNNTGPGSNTTNRVTWPGYHVINSTDA-ANFTVTGLFIEA 563
Query: 326 QFIPKFGVPFIPGLL 282
+I K GVP+ GL+
Sbjct: 564 DWIWKTGVPYTSGLI 578
>sptr|Q8RXK7|Q8RXK7 Hypothetical protein.
Length = 573
Score = 215 bits (548), Expect = 6e-55
Identities = 99/195 (50%), Positives = 133/195 (68%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C + GT+DFIFGN+ VV Q+C + PR+PM NQ N IT
Sbjct: 380 YSCSFEAYQDTLYTHSLRQFYRECDIYGTVDFIFGNAAVVFQDCNLYPRQPMQNQFNAIT 439
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D+ G +HNCTI+P D + ++TYL RPWKEYSRT+++Q+ I ++
Sbjct: 440 AQGRTDQNQNTGISIHNCTIKPADDL-VSSNYTVKTYLGRPWKEYSRTVFMQSYIDEVVE 498
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW EWNGDF L TL+YAE +N G G+ + R W G + +A FTVE F+ G
Sbjct: 499 PVGWREWNGDFALSTLYYAEYNNTGSGSSTTDRVVWPGYHVINSTDA-NNFTVENFLLGD 557
Query: 326 QFIPKFGVPFIPGLL 282
++ + GVP+I GLL
Sbjct: 558 GWMVQSGVPYISGLL 572
>sptr|Q9M9W6|Q9M9W6 Putative pectinesterase.
Length = 669
Score = 215 bits (547), Expect = 8e-55
Identities = 105/198 (53%), Positives = 132/198 (66%), Gaps = 3/198 (1%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
+ C FDGYQDTLYTH+ RQFFRDC ++GTIDF+FG++ V QNC + RKP+ NQA IT
Sbjct: 374 FNCRFDGYQDTLYTHSHRQFFRDCTISGTIDFLFGDAAAVFQNCTLLVRKPLPNQACPIT 433
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDF---KEEAGGKIRTYLARPWKEYSRTIYIQNDIGG 516
A GR+D R G V CTI PD+ KE + + YL RPWKEYSRTI + I
Sbjct: 434 AHGRKDPRESTGFVFQGCTIAGEPDYLAVKETS----KAYLGRPWKEYSRTIIMNTFIPD 489
Query: 515 FIDPKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFI 336
F+ P+GW W GDFGL+TLFY+EV N G G+ ++ R W GIKT++ E+ K FT +I
Sbjct: 490 FVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVTWAGIKTLSEEDILK-FTPAQYI 548
Query: 335 QGQQFIPKFGVPFIPGLL 282
QG +IP GVP+ GLL
Sbjct: 549 QGDDWIPGKGVPYTTGLL 566
>sptr|O81300|O81300 T14P8.14 protein (AT4G02330 protein)
(Hypothetical protein).
Length = 573
Score = 214 bits (546), Expect = 1e-54
Identities = 99/195 (50%), Positives = 132/195 (67%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C + GT+DFIFGN+ VV Q+C + PR+PM NQ N IT
Sbjct: 380 YSCSFEAYQDTLYTHSLRQFYRECDIYGTVDFIFGNAAVVFQDCNLYPRQPMQNQFNAIT 439
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D G +HNCTI+P D + ++TYL RPWKEYSRT+++Q+ I ++
Sbjct: 440 AQGRTDPNQNTGISIHNCTIKPADDL-VSSNYTVKTYLGRPWKEYSRTVFMQSYIDEVVE 498
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW EWNGDF L TL+YAE +N G G+ + R W G + +A FTVE F+ G
Sbjct: 499 PVGWREWNGDFALSTLYYAEYNNTGSGSSTTDRVVWPGYHVINSTDA-NNFTVENFLLGD 557
Query: 326 QFIPKFGVPFIPGLL 282
++ + GVP+I GLL
Sbjct: 558 GWMVQSGVPYISGLL 572
>sptr|Q7XAF8|Q7XAF8 Papillar cell-specific pectin
methylesterase-like protein.
Length = 562
Score = 213 bits (543), Expect = 2e-54
Identities = 99/195 (50%), Positives = 130/195 (66%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C V GT+DFIFGN+ VVLQ C + PR+P QAN +T
Sbjct: 369 YSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAAVVLQKCNLYPRQPRQGQANEVT 428
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D GTVLH CTI P D + ++TYL RPWKEYSRT+ +Q I GF+D
Sbjct: 429 AQGRTDPNQNTGTVLHGCTIRPADDL-ASSNYTVKTYLGRPWKEYSRTVVMQTYIDGFLD 487
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW W+G+F L TL+YAE +N G G+ + R W G + +A FTV F+ G+
Sbjct: 488 PTGWNAWSGNFALSTLYYAEYNNTGPGSSTTNRVTWPGYHVINATDA-SNFTVTNFLVGE 546
Query: 326 QFIPKFGVPFIPGLL 282
+I + GVPF+ G++
Sbjct: 547 GWIGQTGVPFVGGMI 561
>sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610).
Length = 669
Score = 213 bits (542), Expect = 3e-54
Identities = 104/198 (52%), Positives = 132/198 (66%), Gaps = 3/198 (1%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
+ C FDGYQ+TLYTH+ RQFFRDC ++GTIDF+FG++ V QNC + RKP+ NQA IT
Sbjct: 374 FNCRFDGYQNTLYTHSHRQFFRDCTISGTIDFLFGDAAAVFQNCTLLVRKPLPNQACPIT 433
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDF---KEEAGGKIRTYLARPWKEYSRTIYIQNDIGG 516
A GR+D R G V CTI PD+ KE + + YL RPWKEYSRTI + I
Sbjct: 434 AHGRKDPRESTGFVFQGCTIAGEPDYLAVKETS----KAYLGRPWKEYSRTIIMNTFIPD 489
Query: 515 FIDPKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFI 336
F+ P+GW W GDFGL+TLFY+EV N G G+ ++ R W GIKT++ E+ K FT +I
Sbjct: 490 FVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVTWAGIKTLSEEDILK-FTPAQYI 548
Query: 335 QGQQFIPKFGVPFIPGLL 282
QG +IP GVP+ GLL
Sbjct: 549 QGDDWIPGKGVPYTTGLL 566
>sptr|Q93Z62|Q93Z62 At2g47550/T30B22.15.
Length = 345
Score = 212 bits (540), Expect = 5e-54
Identities = 98/195 (50%), Positives = 130/195 (66%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C+F+ YQDTLYTH+ RQF+R+C V GT+DFIFGN+ VVLQNC + PR+P Q+N +T
Sbjct: 152 YSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAAVVLQNCNLYPRQPRKGQSNEVT 211
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR GT +H CTI P D + ++TYL RPWKEYSRT+ +Q I GF++
Sbjct: 212 AQGRTYPNQNTGTAIHGCTIRPADDL-ATSNYTVKTYLGRPWKEYSRTVVMQTYIDGFLE 270
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW W+GDF L TL+YAE +N G G+D + R W G + +A FTV F+ G+
Sbjct: 271 PSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGYHVINATDA-SNFTVTNFLVGE 329
Query: 326 QFIPKFGVPFIPGLL 282
+I + GVPF+ GL+
Sbjct: 330 GWIGQTGVPFVGGLI 344
>sw|O04887|PME2_CITSI Pectinesterase 2.1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 510
Score = 211 bits (536), Expect = 1e-53
Identities = 96/194 (49%), Positives = 127/194 (65%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F+GYQDTLY H+QRQF+R+C + GT+DFIFGN+ VVLQNC I RKP N+ N +T
Sbjct: 319 YRCSFEGYQDTLYVHSQRQFYRECDIYGTVDFIFGNAAVVLQNCNIFARKP-PNRTNTLT 377
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D G ++HNC + D K ++T+L RPWK+YSRT+YI+ + I+
Sbjct: 378 AQGRTDPNQSTGIIIHNCRVTAASDLKP-VQSSVKTFLGRPWKQYSRTVYIKTFLDSLIN 436
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW+EW+GDF L TL+YAE N G G+ + R KW G +T +FTV FI G
Sbjct: 437 PAGWMEWSGDFALNTLYYAEYMNTGPGSSTANRVKWRGYHVLTSPSQVSQFTVGNFIAGN 496
Query: 326 QFIPKFGVPFIPGL 285
++P VPF GL
Sbjct: 497 SWLPATNVPFTSGL 510
>sptr|Q9FF77|Q9FF77 Pectinesterase.
Length = 624
Score = 209 bits (531), Expect = 6e-53
Identities = 100/198 (50%), Positives = 126/198 (63%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F+GYQDTLY H+ RQF+R+C + GTIDFIFGN+ + QNC I RKPMANQ N +T
Sbjct: 429 YRCSFEGYQDTLYVHSLRQFYRECDIYGTIDFIFGNAAAIFQNCNIYARKPMANQKNAVT 488
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
A GR D G + NCTI PD + + T+L RPWK YSRT+YIQ+ I +
Sbjct: 489 AHGRTDPNQKTGISIINCTIGAAPDLAADPKSTM-TFLGRPWKPYSRTVYIQSYISDVVQ 547
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWLEWNG GL+T+ Y E DN G GAD SKR +W G + +A FTV F G
Sbjct: 548 PVGWLEWNGTTGLDTISYGEYDNFGPGADTSKRVQWSGYSLLNLVQAM-NFTVYNFTLGD 606
Query: 326 QFIPKFGVPFIPGLLPQE 273
++P+ +PF GLL E
Sbjct: 607 TWLPQTDIPFYGGLLHTE 624
>sptr|Q9MBB6|Q9MBB6 Pectin methylesterase.
Length = 596
Score = 209 bits (531), Expect = 6e-53
Identities = 104/192 (54%), Positives = 125/192 (65%)
Frame = -2
Query: 860 CTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITAQ 681
C F+GYQDTLY RQF+R C +TGT+DFIFG++ V QNCLI RKP+ NQ NI+TAQ
Sbjct: 408 CRFEGYQDTLYAQTHRQFYRSCVITGTVDFIFGDATSVFQNCLITVRKPLENQQNIVTAQ 467
Query: 680 GRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDPK 501
GR D G VL +C IEP D KIR+YL RPWKE+SRT+ + + IG FI P
Sbjct: 468 GRIDGHETTGIVLQSCRIEPDKDL-VPVKNKIRSYLGRPWKEFSRTVIMDSTIGDFIHPG 526
Query: 500 GWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQF 321
GWL W GDFGL+TL+YAE N+G GA + R KW G + EEA K FT+E F QG +
Sbjct: 527 GWLPWQGDFGLKTLYYAEYSNKGGGAQTNARIKWPGYHIIKKEEAMK-FTIENFYQG-DW 584
Query: 320 IPKFGVPFIPGL 285
I G P GL
Sbjct: 585 ISASGSPVHLGL 596
>sptr|Q7XA94|Q7XA94 Pectinesterase (EC 3.1.1.11) (Fragment).
Length = 193
Score = 207 bits (528), Expect = 1e-52
Identities = 93/192 (48%), Positives = 125/192 (65%)
Frame = -2
Query: 860 CTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITAQ 681
C+F GYQDTLY ++QRQF+RDC + GT+DFIFG++ +LQNC I RKP +NQ N +TAQ
Sbjct: 3 CSFKGYQDTLYVYSQRQFYRDCNIYGTVDFIFGDASAILQNCNIYVRKPSSNQINTVTAQ 62
Query: 680 GRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDPK 501
RRD G ++HNC I PD + G RTYL RPW++YSR + +++++ G I P+
Sbjct: 63 SRRDPNENTGIIIHNCRITAAPDLR-AVQGSFRTYLGRPWQKYSRVVIMKSNLDGLIAPQ 121
Query: 500 GWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQF 321
GW W+G FGL+TL+Y E N G GA R KW G + +T +FTV F+ G +
Sbjct: 122 GWFPWSGSFGLDTLYYGEYMNTGAGAATGGRVKWPGFRVITSATEAGKFTVGNFLAGDAW 181
Query: 320 IPKFGVPFIPGL 285
+P GVPF GL
Sbjct: 182 LPGTGVPFEAGL 193
>sptr|Q8GUY1|Q8GUY1 Methyl pectinesterase (Fragment).
Length = 226
Score = 207 bits (526), Expect = 2e-52
Identities = 94/194 (48%), Positives = 125/194 (64%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F GYQDTLY H+ RQFFR+C + GTIDF+FGNS VVLQ+C + R+P+A+Q+NI T
Sbjct: 34 YRCSFVGYQDTLYVHSLRQFFRECDIYGTIDFVFGNSAVVLQSCNLYARRPLASQSNIYT 93
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D G + C + D RTYL RPWK+YSRT+Y+Q+++ +D
Sbjct: 94 AQGRTDPNQNTGISIQKCKVAAASDLAA-VQTSFRTYLGRPWKQYSRTVYLQSELDSVVD 152
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
PKGWLEW+G F L+TL+Y E N G GA S R KW G + ++ FTV +FI G
Sbjct: 153 PKGWLEWDGTFALDTLYYGEYQNTGAGASTSNRVKWKGYRVISSSSEASTFTVGSFIDGD 212
Query: 326 QFIPKFGVPFIPGL 285
++ +PF GL
Sbjct: 213 VWLAGTSIPFSTGL 226
>sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC
3.1.1.11) (Fragment).
Length = 277
Score = 206 bits (525), Expect = 3e-52
Identities = 94/194 (48%), Positives = 124/194 (63%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
YQC+F YQDTLY H+ RQF+R+C V GT+DFIFGN+ VLQNC + RKP NQ N+ T
Sbjct: 84 YQCSFVAYQDTLYVHSLRQFYRECDVYGTVDFIFGNAAAVLQNCNLYARKPNKNQRNLFT 143
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D G + NC + D + R YL RPWK YSRT+++ + + I+
Sbjct: 144 AQGREDPNQSTGISIINCKVAAAADLIP-VKSEFRNYLGRPWKMYSRTVFLNSLMEDLIE 202
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWLEWNG F L+TL+Y E +NRG GA+ S R W G + +T +FTV+ FIQG
Sbjct: 203 PAGWLEWNGTFALDTLYYGEYNNRGPGANTSGRVTWPGYRVITNSTEASQFTVQNFIQGN 262
Query: 326 QFIPKFGVPFIPGL 285
+++ +G+PF GL
Sbjct: 263 EWLNSYGIPFFSGL 276
>sptrnew|CAE76634|CAE76634 Pectin methylesterase (EC 3.1.1.11)
(Fragment).
Length = 254
Score = 206 bits (523), Expect = 5e-52
Identities = 94/194 (48%), Positives = 122/194 (62%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C GYQDT+Y H+ RQF+R+C + GT+DFIFGN+ VV QNC + RKPMA Q N IT
Sbjct: 60 YRCNIIGYQDTMYVHSNRQFYRECDIYGTVDFIFGNAAVVFQNCSLYARKPMAQQKNTIT 119
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQ R+D G +HNC I PD E + G +TYL RPWK YS+T+Y+ + +G I
Sbjct: 120 AQNRKDPNQNTGISIHNCRILATPDL-EASKGSFQTYLGRPWKLYSKTVYMLSYMGDHIH 178
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P+GWLEWN F L+TL+Y E N G G + +R W G + +T FTV FI G
Sbjct: 179 PRGWLEWNATFALDTLYYGEYMNYGPGGAIGQRVTWPGFRVITSTVEANRFTVAQFISGS 238
Query: 326 QFIPKFGVPFIPGL 285
++P GV F+ GL
Sbjct: 239 SWLPSTGVAFVAGL 252
>sptr|Q9FF78|Q9FF78 Pectinesterase.
Length = 564
Score = 205 bits (522), Expect = 6e-52
Identities = 94/190 (49%), Positives = 126/190 (66%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+CT D +QDT+Y HAQRQF+RDC + GT+DFIFGN+ VV Q C I PR+PM Q N IT
Sbjct: 376 YKCTMDAFQDTMYAHAQRQFYRDCVILGTVDFIFGNAAVVFQKCEILPRRPMKGQQNTIT 435
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR+D G +HNCTI+P + + I+T+L RPWK++S T+ +++ + FI+
Sbjct: 436 AQGRKDPNQNTGISIHNCTIKPLDNLTD-----IQTFLGRPWKDFSTTVIMKSFMDKFIN 490
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
PKGWL W GD +T+FYAE N G GA R KW G+KT ++ +FTV+ FI G
Sbjct: 491 PKGWLPWTGDTAPDTIFYAEYLNSGPGASTKNRVKWQGLKTSLTKKEANKFTVKPFIDGN 550
Query: 326 QFIPKFGVPF 297
++P VPF
Sbjct: 551 NWLPATKVPF 560
>sptr|Q9FK05|Q9FK05 Pectinesterase.
Length = 587
Score = 205 bits (522), Expect = 6e-52
Identities = 95/194 (48%), Positives = 123/194 (63%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C GYQD LY H+ RQFFR+C + GT+DFIFGN+ V+LQ+C I RKPMA Q IT
Sbjct: 393 YRCNIIGYQDALYVHSNRQFFRECEIYGTVDFIFGNAAVILQSCNIYARKPMAQQKITIT 452
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQ R+D G +H C + PD E + G TYL RPWK YSR +Y+ +D+G ID
Sbjct: 453 AQNRKDPNQNTGISIHACKLLATPDL-EASKGSYPTYLGRPWKLYSRVVYMMSDMGDHID 511
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P+GWLEWNG F L++L+Y E N+G G+ + +R KW G +T +FTV FI G
Sbjct: 512 PRGWLEWNGPFALDSLYYGEYMNKGLGSGIGQRVKWPGYHVITSTVEASKFTVAQFISGS 571
Query: 326 QFIPKFGVPFIPGL 285
++P GV F GL
Sbjct: 572 SWLPSTGVSFFSGL 585
>sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 205 bits (521), Expect = 8e-52
Identities = 99/193 (51%), Positives = 127/193 (65%)
Frame = -2
Query: 863 QCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITA 684
+C D +QDTLYTH+ RQF+RDC +TGT+DFIFGN+ VV QN I RKP + Q N++TA
Sbjct: 125 RCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGNAAVVFQNSKIAARKPGSGQKNMVTA 184
Query: 683 QGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDP 504
QGR D GT + NC I P D G ++TYL RPWK YSRT+++Q++IG IDP
Sbjct: 185 QGREDPNQNTGTSIQNCDIIPSSDL-APVKGSVKTYLGRPWKAYSRTVFMQSNIGDHIDP 243
Query: 503 KGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQ 324
+GW W+GDF L+TL+Y E N+G GA SKR KW G ++ EA K FTV IQG
Sbjct: 244 EGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWPGYHILSAAEATK-FTVGQLIQGGV 302
Query: 323 FIPKFGVPFIPGL 285
++ GV + GL
Sbjct: 303 WLKSTGVAYTEGL 315
>sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 205 bits (521), Expect = 8e-52
Identities = 99/193 (51%), Positives = 127/193 (65%)
Frame = -2
Query: 863 QCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITA 684
+C D +QDTLYTH+ RQF+RDC +TGT+DFIFGN+ VV QN I RKP + Q N++TA
Sbjct: 125 RCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGNAAVVFQNSKIAARKPGSGQKNMVTA 184
Query: 683 QGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDP 504
QGR D GT + NC I P D G ++TYL RPWK YSRT+++Q++IG IDP
Sbjct: 185 QGREDPNQNTGTSIQNCDIIPSSDL-APVKGSVKTYLGRPWKAYSRTVFMQSNIGDHIDP 243
Query: 503 KGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQ 324
+GW W+GDF L+TL+Y E N+G GA SKR KW G ++ EA K FTV IQG
Sbjct: 244 EGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWPGYHILSAAEATK-FTVGQLIQGGV 302
Query: 323 FIPKFGVPFIPGL 285
++ GV + GL
Sbjct: 303 WLKSTGVAYTEGL 315
>sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11)
(Pectinesterase) (Pectin methylesterase) (Fragment).
Length = 530
Score = 205 bits (521), Expect = 8e-52
Identities = 98/193 (50%), Positives = 122/193 (63%)
Frame = -2
Query: 863 QCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITA 684
+C D YQDTLY H+QRQF+RD V+GTIDFIFGN+ VV Q C + RKP NQ N++TA
Sbjct: 337 RCPIDAYQDTLYAHSQRQFYRDSYVSGTIDFIFGNAAVVFQKCQLVARKPSKNQKNMVTA 396
Query: 683 QGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDP 504
QGR D GT + C I PD E + +TYL RPWKEYSRT+ +Q+ +GG IDP
Sbjct: 397 QGRTDPNQATGTSIQFCDIIASPDL-EPVVKEFKTYLGRPWKEYSRTVVMQSYLGGLIDP 455
Query: 503 KGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQ 324
GW EW+G+F L+TL+Y E N G GA SKR KW G +T FTV IQG
Sbjct: 456 AGWAEWSGEFALKTLYYGEYMNNGPGAGTSKRVKWPGYHVITDPAEAMPFTVAELIQGGS 515
Query: 323 FIPKFGVPFIPGL 285
++ GV ++ GL
Sbjct: 516 WLSSTGVAYVDGL 528
>sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11)
(AT5g49180/K21P3_5) (Pectinesterase).
Length = 571
Score = 205 bits (521), Expect = 8e-52
Identities = 101/191 (52%), Positives = 128/191 (67%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y C DGYQDTLY H+ RQFFRDC V+GT+DFIFG+ VVLQNC I RKPM +Q+ +IT
Sbjct: 379 YNCQIDGYQDTLYVHSHRQFFRDCTVSGTVDFIFGDGIVVLQNCNIVVRKPMKSQSCMIT 438
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR DKR G VL NC I P + + YL RPWKE+SRTI + I ID
Sbjct: 439 AQGRSDKRESTGLVLQNCHITGEPAYIPVKSIN-KAYLGRPWKEFSRTIIMGTTIDDVID 497
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWL WNGDF L TL+YAE +N G G++ ++R KW GIK ++ ++A + FT F++G
Sbjct: 498 PAGWLPWNGDFALNTLYYAEYENNGPGSNQAQRVKWPGIKKLSPKQALR-FTPARFLRGN 556
Query: 326 QFIPKFGVPFI 294
+IP VP++
Sbjct: 557 LWIPPNRVPYM 567
>sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment).
Length = 274
Score = 205 bits (521), Expect = 8e-52
Identities = 99/193 (51%), Positives = 127/193 (65%)
Frame = -2
Query: 863 QCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITA 684
+C D +QDTLYTH+ RQF+RDC +TGT+DFIFGN+ VV QN I RKP + Q N++TA
Sbjct: 84 RCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGNAAVVFQNSKIAARKPGSGQKNMVTA 143
Query: 683 QGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDP 504
QGR D GT + NC I P D G ++TYL RPWK YSRT+++Q++IG IDP
Sbjct: 144 QGREDPNQNTGTSIQNCDIIPSSDL-APVKGSVKTYLGRPWKAYSRTVFMQSNIGDHIDP 202
Query: 503 KGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQQ 324
+GW W+GDF L+TL+Y E N+G GA SKR KW G ++ EA K FTV IQG
Sbjct: 203 EGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWPGYHILSAAEATK-FTVGQLIQGGV 261
Query: 323 FIPKFGVPFIPGL 285
++ GV + GL
Sbjct: 262 WLKSTGVAYTEGL 274
>sptr|Q94CB1|Q94CB1 Putative pectinesterase.
Length = 619
Score = 203 bits (517), Expect = 2e-51
Identities = 95/198 (47%), Positives = 127/198 (64%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F+GYQDTLY H+ RQF+R+C + GT+DFIFGN+ + QNC I RKPMA Q N IT
Sbjct: 424 YRCSFEGYQDTLYVHSLRQFYRECDIYGTVDFIFGNAAAIFQNCNIYARKPMAKQKNAIT 483
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
A GR D G + NCTI+ PD E + T+L RPWK YSRT+++Q+ I +
Sbjct: 484 AHGRLDPNQNTGISIINCTIKAAPDLAAEPKSAM-TFLGRPWKPYSRTVFMQSYISDIVQ 542
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWLEWNG GL+T++Y E N G GA+ ++R +W G + EA FTV F G
Sbjct: 543 PVGWLEWNGTIGLDTIYYGEYSNFGPGANTNQRVQWLGYNLLNLAEAM-NFTVYNFTMGD 601
Query: 326 QFIPKFGVPFIPGLLPQE 273
++P+ +PF GLL +E
Sbjct: 602 TWLPQTDIPFYGGLLSKE 619
>sptr|Q8LQA0|Q8LQA0 Putative pectinesterase.
Length = 557
Score = 203 bits (517), Expect = 2e-51
Identities = 92/194 (47%), Positives = 120/194 (61%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
YQC+F+ YQDTLYTH+ RQF+R C V GT+D++FGN+ VV Q+C + R PM Q+N +T
Sbjct: 363 YQCSFEAYQDTLYTHSLRQFYRACDVYGTVDYVFGNAAVVFQDCTLYNRLPMQGQSNTVT 422
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D GT + C I PD YL RPWK YSRT+ +Q+ +GG ID
Sbjct: 423 AQGRTDPNQNTGTTIQGCAIVAAPDLAANTAFATTNYLGRPWKLYSRTVIMQSVVGGLID 482
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GW+ W+GD+ L TL+YAE +N G GAD S+R W G + FTV + G
Sbjct: 483 PAGWMPWDGDYALSTLYYAEYNNSGAGADTSRRVTWPGYHVLNSTADAGNFTVGNMVLGD 542
Query: 326 QFIPKFGVPFIPGL 285
++P+ GVPF GL
Sbjct: 543 FWLPQTGVPFTSGL 556
>sptr|Q9SG78|Q9SG78 Putative pectinesterase.
Length = 617
Score = 203 bits (517), Expect = 2e-51
Identities = 95/198 (47%), Positives = 127/198 (64%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F+GYQDTLY H+ RQF+R+C + GT+DFIFGN+ + QNC I RKPMA Q N IT
Sbjct: 422 YRCSFEGYQDTLYVHSLRQFYRECDIYGTVDFIFGNAAAIFQNCNIYARKPMAKQKNAIT 481
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
A GR D G + NCTI+ PD E + T+L RPWK YSRT+++Q+ I +
Sbjct: 482 AHGRLDPNQNTGISIINCTIKAAPDLAAEPKSAM-TFLGRPWKPYSRTVFMQSYISDIVQ 540
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P GWLEWNG GL+T++Y E N G GA+ ++R +W G + EA FTV F G
Sbjct: 541 PVGWLEWNGTIGLDTIYYGEYSNFGPGANTNQRVQWLGYNLLNLAEAM-NFTVYNFTMGD 599
Query: 326 QFIPKFGVPFIPGLLPQE 273
++P+ +PF GLL +E
Sbjct: 600 TWLPQTDIPFYGGLLSKE 617
>sptr|Q84W31|Q84W31 Hypothetical protein.
Length = 564
Score = 202 bits (515), Expect = 4e-51
Identities = 93/190 (48%), Positives = 125/190 (65%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+CT D +QDT+Y HAQRQF+RDC + GT+DFIFGN+ VV Q C I PR+PM Q N IT
Sbjct: 376 YKCTMDAFQDTMYAHAQRQFYRDCVILGTVDFIFGNAAVVFQKCEILPRRPMKGQQNTIT 435
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR+D G +HNCTI+P + + +T+L RPWK++S T+ +++ + FI+
Sbjct: 436 AQGRKDPNQNTGISIHNCTIKPLDNLTD-----TQTFLDRPWKDFSTTVIMKSFMDKFIN 490
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
PKGWL W GD +T+FYAE N G GA R KW G+KT ++ +FTV+ FI G
Sbjct: 491 PKGWLPWTGDTAPDTIFYAEYLNSGPGASTKNRVKWQGLKTSLTKKEANKFTVKPFIDGN 550
Query: 326 QFIPKFGVPF 297
++P VPF
Sbjct: 551 NWLPATKVPF 560
>sptr|Q84R10|Q84R10 Putative pectinesterase.
Length = 519
Score = 202 bits (515), Expect = 4e-51
Identities = 95/194 (48%), Positives = 126/194 (64%)
Frame = -2
Query: 866 YQCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIIT 687
Y+C+F GYQDTL+TH+ RQF+RD + GTIDFIFG++ V QNC I R+PM +Q N+IT
Sbjct: 327 YRCSFKGYQDTLFTHSLRQFYRDRHIYGTIDFIFGDAAAVFQNCDIFVRRPMDHQGNMIT 386
Query: 686 AQGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFID 507
AQGR D + G + + I P+F E G+ ++YL RPWK+YSRT++++ DI ID
Sbjct: 387 AQGRDDPHTNSGISIQHSRIRAAPEF-EAVKGRFKSYLGRPWKKYSRTVFLKTDIDELID 445
Query: 506 PKGWLEWNGDFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
P+GW EW+G + L TL+Y E N G GA +R W G + EE FTV FIQG
Sbjct: 446 PRGWREWSGSYALSTLYYGEFMNTGAGAGTGRRVNWPGFHVLRGEEEASPFTVSRFIQGD 505
Query: 326 QFIPKFGVPFIPGL 285
+IP GVPF G+
Sbjct: 506 SWIPITGVPFSAGV 519
>sptr|Q8S122|Q8S122 Putative pectin esterase.
Length = 563
Score = 202 bits (515), Expect = 4e-51
Identities = 94/194 (48%), Positives = 123/194 (63%), Gaps = 1/194 (0%)
Frame = -2
Query: 863 QCTFDGYQDTLYTHAQRQFFRDCRVTGTIDFIFGNSQVVLQNCLIQPRKPMANQANIITA 684
+C DGYQDTLY H RQF+RDC V+GT+DF+FGN+ VLQ C++ R+P Q N +TA
Sbjct: 371 RCRLDGYQDTLYAHQLRQFYRDCAVSGTVDFVFGNAAAVLQGCVLTARRPAQAQKNAVTA 430
Query: 683 QGRRDKRSVGGTVLHNCTIEPHPDFKEEAGGKIRTYLARPWKEYSRTIYIQNDIGGFIDP 504
QGR D GT +H C + P PD A + T+L RPWKEYSRT+Y+ + + +DP
Sbjct: 431 QGRTDPNQNTGTSIHRCRVVPAPDL-APAAKQFPTFLGRPWKEYSRTVYMLSYLDSHVDP 489
Query: 503 KGWLEWNG-DFGLETLFYAEVDNRGDGADMSKRAKWGGIKTVTYEEAQKEFTVETFIQGQ 327
+GWLEWNG DF L+TLFY E N+G GA + R W G +T + +FTV FIQG
Sbjct: 490 RGWLEWNGADFALKTLFYGEYQNQGPGASTAGRVNWPGYHVITDQSVAMQFTVGQFIQGG 549
Query: 326 QFIPKFGVPFIPGL 285
++ GV + GL
Sbjct: 550 NWLKATGVNYNEGL 563
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 564,832,845
Number of Sequences: 1395590
Number of extensions: 12290168
Number of successful extensions: 41989
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 40318
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 41680
length of database: 442,889,342
effective HSP length: 122
effective length of database: 272,627,362
effective search space used: 45256142092
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

