BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
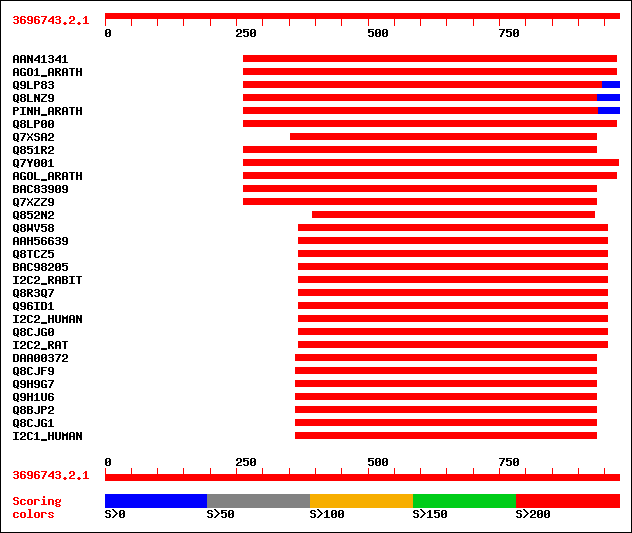
Query= 3696743.2.1
(977 letters)
Database: /db/trembl-ebi/tmp/swall
1,288,562 sequences; 411,996,502 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|AAN41341|AAN41341 Putative leaf development protein Argo... 397 e-109
sw|O04379|AGO1_ARATH Argonaute protein. 397 e-109
sptr|Q9LP83|Q9LP83 T1N15.2. 394 e-109
sptr|Q8LNZ9|Q8LNZ9 AGO1 homologous protein. 380 e-107
sw|Q9XGW1|PINH_ARATH PINHEAD protein (ZWILLE protein). 371 e-104
sptr|Q8LP00|Q8LP00 ZLL/PNH homologous protein. 362 6e-99
sptr|Q7XSA2|Q7XSA2 OSJNBa0005N02.3 protein. 361 8e-99
sptr|Q851R2|Q851R2 Putative argonaute protein. 323 2e-87
sptr|Q7Y001|Q7Y001 Putative leaf development and shoot apical me... 311 9e-84
sw|Q9SJK3|AGOL_ARATH Argonaute-like protein At2g27880. 309 4e-83
sptrnew|BAC83909|BAC83909 Putative leaf development protein Argo... 299 4e-80
sptr|Q7XZZ9|Q7XZZ9 Putative leaf development and shoot apical me... 298 6e-80
sptr|Q852N2|Q852N2 Putative argonaute protein. 272 5e-72
sptr|Q8WV58|Q8WV58 Hypothetical protein (Eukaryotic translation ... 258 1e-67
sptrnew|AAH56639|AAH56639 Hypothetical protein (Fragment). 258 1e-67
sptr|Q8TCZ5|Q8TCZ5 Eukaryotic initiation factor 2C2. 258 1e-67
sptrnew|BAC98205|BAC98205 MKIAA1567 protein (Fragment). 258 1e-67
sw|O77503|I2C2_RABIT Eukaryotic translation initiation factor 2C... 258 1e-67
sptr|Q8R3Q7|Q8R3Q7 Similar to eukaryotic translation initiation ... 258 1e-67
sptr|Q96ID1|Q96ID1 Hypothetical protein. 258 1e-67
sw|Q9UKV8|I2C2_HUMAN Eukaryotic translation initiation factor 2C... 258 1e-67
sptr|Q8CJG0|Q8CJG0 Piwi/Argonaute family protain meIF2C2. 256 3e-67
sw|Q9QZ81|I2C2_RAT Eukaryotic translation initiation factor 2C 2... 256 3e-67
sptrnew|DAA00372|DAA00372 Argonaute 3. 256 4e-67
sptr|Q8CJF9|Q8CJF9 Piwi/Argonaute family protain meIF2C3. 256 4e-67
sptr|Q9H9G7|Q9H9G7 Hypothetical protein FLJ12765. 256 4e-67
sptr|Q9H1U6|Q9H1U6 DJ665N4.6 (Novel protein) (Fragment). 256 4e-67
sptr|Q8BJP2|Q8BJP2 Eukaryotic translation initiation factor 2C 1. 244 1e-63
sptr|Q8CJG1|Q8CJG1 Piwi/Argonaute family protain meIF2C1. 244 1e-63
sw|Q9UL18|I2C1_HUMAN Eukaryotic translation initiation factor 2C... 244 1e-63
>sptrnew|AAN41341|AAN41341 Putative leaf development protein
Argonaute.
Length = 1048
Score = 397 bits (1020), Expect = e-109
Identities = 194/237 (81%), Positives = 208/237 (87%)
Frame = -2
Query: 973 PKGGLSLVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 794
P+ G+ + LLI+F R+TG KP RIIFYRDGVSEGQFYQVLLYELDAIRKACASLE
Sbjct: 813 PQKGVVTGGMIKELLIAFRRSTGHKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 872
Query: 793 SDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSH 614
+ YQPPVTFVVVQKRHHTRLF NHND+ + DRSGNILPGTVVDSKICHPTEFDFYLCSH
Sbjct: 873 AGYQPPVTFVVVQKRHHTRLFAQNHNDRHSVDRSGNILPGTVVDSKICHPTEFDFYLCSH 932
Query: 613 AGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 434
AGIQGTSRPAHYHVLWDEN FTADGLQ+LTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA
Sbjct: 933 AGIQGTSRPAHYHVLWDENNFTADGLQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 992
Query: 433 RFYMEPDTSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
RFYMEP+TSDSGS+ASG + +RG R+TR N AVRPLPALKENVKRVMFYC
Sbjct: 993 RFYMEPETSDSGSMASG-SMARGGGMAGRSTRGPNVNAAVRPLPALKENVKRVMFYC 1048
>sw|O04379|AGO1_ARATH Argonaute protein.
Length = 1048
Score = 397 bits (1020), Expect = e-109
Identities = 194/237 (81%), Positives = 208/237 (87%)
Frame = -2
Query: 973 PKGGLSLVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 794
P+ G+ + LLI+F R+TG KP RIIFYRDGVSEGQFYQVLLYELDAIRKACASLE
Sbjct: 813 PQKGVVTGGMIKELLIAFRRSTGHKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 872
Query: 793 SDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSH 614
+ YQPPVTFVVVQKRHHTRLF NHND+ + DRSGNILPGTVVDSKICHPTEFDFYLCSH
Sbjct: 873 AGYQPPVTFVVVQKRHHTRLFAQNHNDRHSVDRSGNILPGTVVDSKICHPTEFDFYLCSH 932
Query: 613 AGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 434
AGIQGTSRPAHYHVLWDEN FTADGLQ+LTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA
Sbjct: 933 AGIQGTSRPAHYHVLWDENNFTADGLQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 992
Query: 433 RFYMEPDTSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
RFYMEP+TSDSGS+ASG + +RG R+TR N AVRPLPALKENVKRVMFYC
Sbjct: 993 RFYMEPETSDSGSMASG-SMARGGGMAGRSTRGPNVNAAVRPLPALKENVKRVMFYC 1048
>sptr|Q9LP83|Q9LP83 T1N15.2.
Length = 1123
Score = 394 bits (1011), Expect(2) = e-109
Identities = 193/230 (83%), Positives = 205/230 (89%), Gaps = 3/230 (1%)
Frame = -2
Query: 943 LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFV 764
+G LLI+F R+TG KP RIIFYRDGVSEGQFYQVLLYELDAIRKACASLE+ YQPPVTFV
Sbjct: 895 IGELLIAFRRSTGHKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEAGYQPPVTFV 954
Query: 763 VVQKRHHTRLFVNNHNDQRAADRSGNILPG---TVVDSKICHPTEFDFYLCSHAGIQGTS 593
VVQKRHHTRLF NHND+ + DRSGNILPG TVVDSKICHPTEFDFYLCSHAGIQGTS
Sbjct: 955 VVQKRHHTRLFAQNHNDRHSVDRSGNILPGETSTVVDSKICHPTEFDFYLCSHAGIQGTS 1014
Query: 592 RPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPD 413
RPAHYHVLWDEN FTADGLQ+LTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP+
Sbjct: 1015 RPAHYHVLWDENNFTADGLQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPE 1074
Query: 412 TSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
TSDSGS+ASG + +RG R+TR N AVRPLPALKENVKRVMFYC
Sbjct: 1075 TSDSGSMASG-SMARGGGMAGRSTRGPNVNAAVRPLPALKENVKRVMFYC 1123
Score = 25.4 bits (54), Expect(2) = e-109
Identities = 9/13 (69%), Positives = 12/13 (92%)
Frame = -1
Query: 977 DPQRGTVSGGMIR 939
DPQ+G V+GGMI+
Sbjct: 872 DPQKGVVTGGMIK 884
>sptr|Q8LNZ9|Q8LNZ9 AGO1 homologous protein.
Length = 909
Score = 380 bits (976), Expect(2) = e-107
Identities = 185/225 (82%), Positives = 201/225 (89%), Gaps = 1/225 (0%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLISF R+TG+KP+RIIFYRDGVSEGQFYQVLLYEL+AIRKACASLE++YQP VTF+VVQ
Sbjct: 686 LLISFKRSTGEKPQRIIFYRDGVSEGQFYQVLLYELNAIRKACASLETNYQPKVTFIVVQ 745
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF +NHNDQ + DRSGNILPGTVVDSKICHPTEFDFYLCSHAGI+GTSRPAHYH
Sbjct: 746 KRHHTRLFAHNHNDQNSVDRSGNILPGTVVDSKICHPTEFDFYLCSHAGIKGTSRPAHYH 805
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSGS 395
VLW+EN FTAD LQ LTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS S
Sbjct: 806 VLWNENNFTADALQILTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSSS 865
Query: 394 VASGATTSRGPPPGARNTRAGA-ANVAVRPLPALKENVKRVMFYC 263
V SG RGP G+ +R A AV+PLPALK++VKRVMFYC
Sbjct: 866 VVSGPGV-RGPLSGSSTSRTRAPGGAAVKPLPALKDSVKRVMFYC 909
Score = 32.7 bits (73), Expect(2) = e-107
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = -1
Query: 977 DPQRGTVSGGMIRE 936
DPQRGTVSGGMIRE
Sbjct: 672 DPQRGTVSGGMIRE 685
>sw|Q9XGW1|PINH_ARATH PINHEAD protein (ZWILLE protein).
Length = 988
Score = 371 bits (953), Expect(2) = e-104
Identities = 181/227 (79%), Positives = 197/227 (86%), Gaps = 2/227 (0%)
Frame = -2
Query: 937 NLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVV 758
+LLISF +ATGQKP RIIFYRDGVSEGQFYQVLLYELDAIRKACASLE +YQPPVTF+VV
Sbjct: 774 DLLISFRKATGQKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPNYQPPVTFIVV 833
Query: 757 QKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHY 578
QKRHHTRLF NNH D+ + DRSGNILPGTVVD+KICHPTEFDFYLCSHAGIQGTSRPAHY
Sbjct: 834 QKRHHTRLFANNHRDKNSTDRSGNILPGTVVDTKICHPTEFDFYLCSHAGIQGTSRPAHY 893
Query: 577 HVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPD-TSDS 401
HVLWDEN FTADG+Q+LTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFY+EP+ D+
Sbjct: 894 HVLWDENNFTADGIQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYLEPEIMQDN 953
Query: 400 GSVASGATTSRGPPPGARNTR-AGAANVAVRPLPALKENVKRVMFYC 263
GS PG +NT+ +V V+PLPALKENVKRVMFYC
Sbjct: 954 GS------------PGKKNTKTTTVGDVGVKPLPALKENVKRVMFYC 988
Score = 28.9 bits (63), Expect(2) = e-104
Identities = 12/14 (85%), Positives = 13/14 (92%)
Frame = -1
Query: 977 DPQRGTVSGGMIRE 936
DP RGTVSGGMIR+
Sbjct: 761 DPVRGTVSGGMIRD 774
>sptr|Q8LP00|Q8LP00 ZLL/PNH homologous protein.
Length = 978
Score = 362 bits (928), Expect = 6e-99
Identities = 176/237 (74%), Positives = 195/237 (82%)
Frame = -2
Query: 973 PKGGLSLVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 794
P+ G + LLISF +ATGQKP RIIFYRDGVSEGQFYQVLLYELDAIRKACASLE
Sbjct: 756 PQRGTVTGGMIRELLISFRKATGQKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 815
Query: 793 SDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSH 614
YQPPVTFVVVQKRHHTRLF NNH D+ + D+SGNILPGTVVDSKICHP+EFDFYLCSH
Sbjct: 816 PIYQPPVTFVVVQKRHHTRLFANNHKDRSSTDKSGNILPGTVVDSKICHPSEFDFYLCSH 875
Query: 613 AGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 434
AGIQGTSRPAHYHVLWDEN FTAD +QTLTNNLCYTYARCTRSVS+VPPAYYAHLAAFRA
Sbjct: 876 AGIQGTSRPAHYHVLWDENNFTADEMQTLTNNLCYTYARCTRSVSVVPPAYYAHLAAFRA 935
Query: 433 RFYMEPDTSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
RFYMEP+ S++ + + +T + G +V+PLPA+KE VKRVMFYC
Sbjct: 936 RFYMEPEMSENQTTSKSSTGTNG--------------TSVKPLPAVKEKVKRVMFYC 978
>sptr|Q7XSA2|Q7XSA2 OSJNBa0005N02.3 protein.
Length = 1254
Score = 361 bits (927), Expect = 8e-99
Identities = 182/231 (78%), Positives = 186/231 (80%), Gaps = 37/231 (16%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYR-------------------------------------DGV 866
LLISF RATGQKP+RIIFYR DGV
Sbjct: 945 LLISFKRATGQKPQRIIFYRFLSLYRNLRGQHFAGFYPCMDIFLTIIVCDFNTCHFRDGV 1004
Query: 865 SEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGN 686
SEGQFYQVLLYELDAIRKACASLE +YQPPVTFVVVQKRHHTRLF NNHNDQR DRSGN
Sbjct: 1005 SEGQFYQVLLYELDAIRKACASLEPNYQPPVTFVVVQKRHHTRLFANNHNDQRTVDRSGN 1064
Query: 685 ILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYT 506
ILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENKFTAD LQTLTNNLCYT
Sbjct: 1065 ILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENKFTADELQTLTNNLCYT 1124
Query: 505 YARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSGSVASGATTSRGPPPG 353
YARCTRSVSIVPPAYYAHLAAFRARFYMEP+TSDSGS+ASGA TSRG PPG
Sbjct: 1125 YARCTRSVSIVPPAYYAHLAAFRARFYMEPETSDSGSMASGAATSRGLPPG 1175
>sptr|Q851R2|Q851R2 Putative argonaute protein.
Length = 1058
Score = 323 bits (828), Expect = 2e-87
Identities = 160/226 (70%), Positives = 181/226 (80%), Gaps = 2/226 (0%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI+F + TG++P+RIIFYRDGVSEGQF VLL+E+DAIRKACASLE Y PPVTFVVVQ
Sbjct: 845 LLIAFRKKTGRRPERIIFYRDGVSEGQFSHVLLHEMDAIRKACASLEEGYLPPVTFVVVQ 904
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF H + D+SGNILPGTVVD +ICHPTEFDFYLCSHAGIQGTSRP HYH
Sbjct: 905 KRHHTRLFPEVHGRRDMTDKSGNILPGTVVDRQICHPTEFDFYLCSHAGIQGTSRPTHYH 964
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSGS 395
VL+DEN FTAD LQ+LTNNLCYTYARCTR+VS+VPPAYYAHLAAFRAR+Y+E ++SD GS
Sbjct: 965 VLYDENHFTADALQSLTNNLCYTYARCTRAVSVVPPAYYAHLAAFRARYYVEGESSDGGS 1024
Query: 394 V--ASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
+SG +R P V VR LP +KENVK VMFYC
Sbjct: 1025 TPGSSGQAVAREGP------------VEVRQLPKIKENVKDVMFYC 1058
>sptr|Q7Y001|Q7Y001 Putative leaf development and shoot apical
meristem regulating protein.
Length = 1055
Score = 311 bits (797), Expect = 9e-84
Identities = 160/238 (67%), Positives = 181/238 (76%)
Frame = -2
Query: 976 IPKGGLSLVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASL 797
I +GG+ + LL SF++ TGQKP RIIFYRDG+SEGQF QVLLYE+DAIRKACASL
Sbjct: 840 IIRGGM-----IRELLRSFYQETGQKPSRIIFYRDGISEGQFSQVLLYEMDAIRKACASL 894
Query: 796 ESDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCS 617
+ Y PPVTFVVVQKRHHTRLF N D DRSGNILPGTVVD+ ICHP+EFDFYLCS
Sbjct: 895 QEGYLPPVTFVVVQKRHHTRLFPENRRDMM--DRSGNILPGTVVDTMICHPSEFDFYLCS 952
Query: 616 HAGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFR 437
H+GI+GTSRP HYHVL DEN F AD LQTLT NL YTYARCTR+VSIVPPAYYAHL AFR
Sbjct: 953 HSGIKGTSRPTHYHVLLDENGFKADTLQTLTYNLSYTYARCTRAVSIVPPAYYAHLGAFR 1012
Query: 436 ARFYMEPDTSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
AR+YME + SD GS S + T+R + + +PLP +KENVKR MFYC
Sbjct: 1013 ARYYMEDEHSDQGS--SSSVTTR-------------TDRSTKPLPEIKENVKRFMFYC 1055
>sw|Q9SJK3|AGOL_ARATH Argonaute-like protein At2g27880.
Length = 997
Score = 309 bits (791), Expect = 4e-83
Identities = 158/237 (66%), Positives = 179/237 (75%)
Frame = -2
Query: 973 PKGGLSLVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLE 794
P+ GL + I+F RATGQ P+RIIFYRDGVSEGQF QVLL+E+ AIRKAC SL+
Sbjct: 774 PQRGLVHSGLIREHFIAFRRATGQIPQRIIFYRDGVSEGQFSQVLLHEMTAIRKACNSLQ 833
Query: 793 SDYQPPVTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSH 614
+Y P VTFV+VQKRHHTRLF H ++ D+SGNI PGTVVD+KICHP EFDFYL SH
Sbjct: 834 ENYVPRVTFVIVQKRHHTRLFPEQHGNRDMTDKSGNIQPGTVVDTKICHPNEFDFYLNSH 893
Query: 613 AGIQGTSRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA 434
AGIQGTSRPAHYHVL DEN FTAD LQ LTNNLCYTYARCT+SVSIVPPAYYAHLAAFRA
Sbjct: 894 AGIQGTSRPAHYHVLLDENGFTADQLQMLTNNLCYTYARCTKSVSIVPPAYYAHLAAFRA 953
Query: 433 RFYMEPDTSDSGSVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
R+YME + SD GS S R++ G V + LPA+K+NVK VMFYC
Sbjct: 954 RYYMESEMSDGGSSRS------------RSSTTGVGQV-ISQLPAIKDNVKEVMFYC 997
>sptrnew|BAC83909|BAC83909 Putative leaf development protein
Argonaute.
Length = 1052
Score = 299 bits (766), Expect = 4e-80
Identities = 151/225 (67%), Positives = 168/225 (74%), Gaps = 1/225 (0%)
Frame = -2
Query: 934 LLISFWRATGQ-KPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVV 758
LL+SF+ + KP+RIIFYRDGVS+GQF VLLYE+DAI+KA ASL+ Y+P VTFVVV
Sbjct: 837 LLMSFYSKNAKRKPQRIIFYRDGVSDGQFLHVLLYEMDAIKKAIASLDPAYRPLVTFVVV 896
Query: 757 QKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHY 578
QKRHHTRLF H Q DRSGN+ PGTVVD+ ICHP+EFDFYLCSHAGIQGTSRP HY
Sbjct: 897 QKRHHTRLFPEVHGRQDLTDRSGNVRPGTVVDTNICHPSEFDFYLCSHAGIQGTSRPTHY 956
Query: 577 HVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSG 398
HVL DEN+F+AD LQ LT NLCYTYARCTRSVS+VPPAYYAHLAAFRAR+Y EP D
Sbjct: 957 HVLHDENRFSADQLQMLTYNLCYTYARCTRSVSVVPPAYYAHLAAFRARYYDEPPAMDGA 1016
Query: 397 SVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
S G AG AVR LP +KENVK VMFYC
Sbjct: 1017 SSVGS---------GGNQAAAGGQPPAVRRLPQIKENVKDVMFYC 1052
>sptr|Q7XZZ9|Q7XZZ9 Putative leaf development and shoot apical
meristem regulating protein.
Length = 895
Score = 298 bits (764), Expect = 6e-80
Identities = 152/225 (67%), Positives = 174/225 (77%), Gaps = 1/225 (0%)
Frame = -2
Query: 934 LLISFWRATGQ-KPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVV 758
L+ SF +A G KP RIIFYRDGVSEGQF QVLL E+DAIRKACAS+E Y PPVTFVVV
Sbjct: 679 LIESFRKANGSYKPGRIIFYRDGVSEGQFSQVLLSEMDAIRKACASIEEGYLPPVTFVVV 738
Query: 757 QKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHY 578
QKRHHTRLF +H+ + DRS NILPGTVVD+KICHP+EFDFYLCSH+GIQGTS P HY
Sbjct: 739 QKRHHTRLFPEDHHARDQMDRSRNILPGTVVDTKICHPSEFDFYLCSHSGIQGTSHPTHY 798
Query: 577 HVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSG 398
+VL+DEN F+AD LQTLT +LCYTYARCTRSVSIVPP YYAHLAA RAR Y+E G
Sbjct: 799 YVLFDENNFSADALQTLTYHLCYTYARCTRSVSIVPPVYYAHLAASRARHYLE-----EG 853
Query: 397 SVASGATTSRGPPPGARNTRAGAANVAVRPLPALKENVKRVMFYC 263
S+ ++S G+R G V V+PLP +KENVK+ MFYC
Sbjct: 854 SLPDHGSSSASAAGGSRRNDRG---VPVKPLPEIKENVKQFMFYC 895
>sptr|Q852N2|Q852N2 Putative argonaute protein.
Length = 1192
Score = 272 bits (696), Expect = 5e-72
Identities = 131/179 (73%), Positives = 149/179 (83%)
Frame = -2
Query: 931 LISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQK 752
LI+F + TG++P+RIIFYRDGVSEGQF +VLL+E+DAIRKACASLE Y PPVTFVVVQK
Sbjct: 600 LIAFRKKTGRRPERIIFYRDGVSEGQFSRVLLHEMDAIRKACASLEEGYLPPVTFVVVQK 659
Query: 751 RHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHV 572
RHHTRLF H + D+SGNILPGTV D +ICHPTEF FYLCSHAGIQGTSRP HYHV
Sbjct: 660 RHHTRLFPEVHGRRDMTDKSGNILPGTVKDRQICHPTEFYFYLCSHAGIQGTSRPTHYHV 719
Query: 571 LWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDSGS 395
L+DEN FTAD LQTLTNNLCY YARCT +VS+VPPAYY+HLAA A ++ +S SGS
Sbjct: 720 LYDENHFTADELQTLTNNLCYIYARCTHAVSVVPPAYYSHLAASHAHCCIKGHSSGSGS 778
>sptr|Q8WV58|Q8WV58 Hypothetical protein (Eukaryotic translation
initiation factor 2C, 2).
Length = 585
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 368 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 427
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 428 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 485
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 486 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 545
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 546 KEHDSAEGSHTSGQSNGR 563
>sptrnew|AAH56639|AAH56639 Hypothetical protein (Fragment).
Length = 437
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 220 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 279
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 280 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 337
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 338 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 397
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 398 KEHDSAEGSHTSGQSNGR 415
>sptr|Q8TCZ5|Q8TCZ5 Eukaryotic initiation factor 2C2.
Length = 851
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 634 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 693
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 694 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 751
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 752 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 811
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 812 KEHDSAEGSHTSGQSNGR 829
>sptrnew|BAC98205|BAC98205 MKIAA1567 protein (Fragment).
Length = 703
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 486 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 545
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 546 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 603
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 604 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 663
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 664 KEHDSAEGSHTSGQSNGR 681
>sw|O77503|I2C2_RABIT Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2).
Length = 813
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 596 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 655
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 656 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 713
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 714 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 773
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 774 KEHDSAEGSHTSGQSNGR 791
>sptr|Q8R3Q7|Q8R3Q7 Similar to eukaryotic translation initiation
factor 2C, 2 (Fragment).
Length = 530
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 313 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 372
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 373 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 430
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 431 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 490
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 491 KEHDSAEGSHTSGQSNGR 508
>sptr|Q96ID1|Q96ID1 Hypothetical protein.
Length = 377
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 160 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 219
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 220 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 277
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 278 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 337
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 338 KEHDSAEGSHTSGQSNGR 355
>sw|Q9UKV8|I2C2_HUMAN Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2) (Fragment).
Length = 377
Score = 258 bits (658), Expect = 1e-67
Identities = 129/198 (65%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 160 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 219
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 220 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 277
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 278 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 337
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 338 KEHDSAEGSHTSGQSNGR 355
>sptr|Q8CJG0|Q8CJG0 Piwi/Argonaute family protain meIF2C2.
Length = 860
Score = 256 bits (654), Expect = 3e-67
Identities = 128/198 (64%), Positives = 151/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE DYQP
Sbjct: 643 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKDYQPG 702
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 703 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 760
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
RP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 761 GRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 820
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 821 KEHDSAEGSHTSGQSNGR 838
>sw|Q9QZ81|I2C2_RAT Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2) (Golgi ER protein 95 kDa) (GERp95).
Length = 863
Score = 256 bits (654), Expect = 3e-67
Identities = 128/198 (64%), Positives = 152/198 (76%), Gaps = 2/198 (1%)
Frame = -2
Query: 955 LVA*LGNLLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPP 776
L A + LLI F+++T KP RIIFYRDGVSEGQF QVL +EL AIR+AC LE +YQP
Sbjct: 646 LAAMVRELLIQFYKSTRFKPTRIIFYRDGVSEGQFQQVLHHELLAIREACIKLEKEYQPG 705
Query: 775 VTFVVVQKRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGT 596
+TF+VVQKRHHTRLF + N++ +SGNI GT VD+KI HPTEFDFYLCSHAGIQGT
Sbjct: 706 ITFIVVQKRHHTRLFCTDKNER--VGKSGNIPAGTTVDTKITHPTEFDFYLCSHAGIQGT 763
Query: 595 SRPAHYHVLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEP 416
SRP+HYHVLWD+N+F++D LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++
Sbjct: 764 SRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVD 823
Query: 415 DTSDS--GSVASGATTSR 368
DS GS SG + R
Sbjct: 824 KEHDSAEGSHTSGQSNGR 841
>sptrnew|DAA00372|DAA00372 Argonaute 3.
Length = 748
Score = 256 bits (653), Expect = 4e-67
Identities = 129/193 (66%), Positives = 147/193 (76%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGVSEGQF QVL YEL AIR+AC SLE DYQP +T++VVQ
Sbjct: 538 LLIQFYKSTRFKPTRIIFYRDGVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQ 597
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + ++ RSGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HYH
Sbjct: 598 KRHHTRLFCADRTER--VGRSGNIPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYH 655
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N FTAD LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++ DS
Sbjct: 656 VLWDDNFFTADELQLLTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAE 715
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 716 GSHVSGQSNGRDP 728
>sptr|Q8CJF9|Q8CJF9 Piwi/Argonaute family protain meIF2C3.
Length = 860
Score = 256 bits (653), Expect = 4e-67
Identities = 129/193 (66%), Positives = 147/193 (76%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGVSEGQF QVL YEL AIR+AC SLE DYQP +T++VVQ
Sbjct: 650 LLIQFYKSTRFKPTRIIFYRDGVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQ 709
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + ++ RSGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HYH
Sbjct: 710 KRHHTRLFCADRTER--VGRSGNIPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYH 767
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N FTAD LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++ DS
Sbjct: 768 VLWDDNFFTADELQLLTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAE 827
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 828 GSHVSGQSNGRDP 840
>sptr|Q9H9G7|Q9H9G7 Hypothetical protein FLJ12765.
Length = 860
Score = 256 bits (653), Expect = 4e-67
Identities = 129/193 (66%), Positives = 147/193 (76%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGVSEGQF QVL YEL AIR+AC SLE DYQP +T++VVQ
Sbjct: 650 LLIQFYKSTRFKPTRIIFYRDGVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQ 709
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + ++ RSGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HYH
Sbjct: 710 KRHHTRLFCADRTER--VGRSGNIPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYH 767
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N FTAD LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++ DS
Sbjct: 768 VLWDDNCFTADELQLLTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAE 827
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 828 GSHVSGQSNGRDP 840
>sptr|Q9H1U6|Q9H1U6 DJ665N4.6 (Novel protein) (Fragment).
Length = 640
Score = 256 bits (653), Expect = 4e-67
Identities = 129/193 (66%), Positives = 147/193 (76%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGVSEGQF QVL YEL AIR+AC SLE DYQP +T++VVQ
Sbjct: 430 LLIQFYKSTRFKPTRIIFYRDGVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQ 489
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + ++ RSGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HYH
Sbjct: 490 KRHHTRLFCADRTER--VGRSGNIPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYH 547
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N FTAD LQ LT LC+TY RCTRSVSI PAYYAHL AFRAR+++ DS
Sbjct: 548 VLWDDNCFTADELQLLTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAE 607
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 608 GSHVSGQSNGRDP 620
>sptr|Q8BJP2|Q8BJP2 Eukaryotic translation initiation factor 2C 1.
Length = 706
Score = 244 bits (624), Expect = 1e-63
Identities = 123/193 (63%), Positives = 144/193 (74%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGV EGQ Q+L YEL AIR AC LE DYQP +T++VVQ
Sbjct: 496 LLIQFYKSTRFKPTRIIFYRDGVPEGQLPQILHYELLAIRDACIKLEKDYQPGITYIVVQ 555
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + N++ +SGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HY+
Sbjct: 556 KRHHTRLFCADKNER--IGKSGNIPAGTTVDTNITHPFEFDFYLCSHAGIQGTSRPSHYY 613
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N+FTAD LQ LT LC+TY RCTRSVSI PAYYA L AFRAR+++ DS
Sbjct: 614 VLWDDNRFTADELQILTYQLCHTYVRCTRSVSIPAPAYYARLVAFRARYHLVDKEHDSGE 673
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 674 GSHISGQSNGRDP 686
>sptr|Q8CJG1|Q8CJG1 Piwi/Argonaute family protain meIF2C1.
Length = 857
Score = 244 bits (624), Expect = 1e-63
Identities = 123/193 (63%), Positives = 144/193 (74%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGV EGQ Q+L YEL AIR AC LE DYQP +T++VVQ
Sbjct: 647 LLIQFYKSTRFKPTRIIFYRDGVPEGQLPQILHYELLAIRDACIKLEKDYQPGITYIVVQ 706
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + N++ +SGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HY+
Sbjct: 707 KRHHTRLFCADKNER--IGKSGNIPAGTTVDTNITHPFEFDFYLCSHAGIQGTSRPSHYY 764
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N+FTAD LQ LT LC+TY RCTRSVSI PAYYA L AFRAR+++ DS
Sbjct: 765 VLWDDNRFTADELQILTYQLCHTYVRCTRSVSIPAPAYYARLVAFRARYHLVDKEHDSGE 824
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 825 GSHISGQSNGRDP 837
>sw|Q9UL18|I2C1_HUMAN Eukaryotic translation initiation factor 2C 1
(eIF2C 1) (eIF-2C 1) (Putative RNA-binding protein Q99).
Length = 857
Score = 244 bits (624), Expect = 1e-63
Identities = 123/193 (63%), Positives = 144/193 (74%), Gaps = 2/193 (1%)
Frame = -2
Query: 934 LLISFWRATGQKPKRIIFYRDGVSEGQFYQVLLYELDAIRKACASLESDYQPPVTFVVVQ 755
LLI F+++T KP RIIFYRDGV EGQ Q+L YEL AIR AC LE DYQP +T++VVQ
Sbjct: 647 LLIQFYKSTRFKPTRIIFYRDGVPEGQLPQILHYELLAIRDACIKLEKDYQPGITYIVVQ 706
Query: 754 KRHHTRLFVNNHNDQRAADRSGNILPGTVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYH 575
KRHHTRLF + N++ +SGNI GT VD+ I HP EFDFYLCSHAGIQGTSRP+HY+
Sbjct: 707 KRHHTRLFCADKNER--IGKSGNIPAGTTVDTNITHPFEFDFYLCSHAGIQGTSRPSHYY 764
Query: 574 VLWDENKFTADGLQTLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARFYMEPDTSDS-- 401
VLWD+N+FTAD LQ LT LC+TY RCTRSVSI PAYYA L AFRAR+++ DS
Sbjct: 765 VLWDDNRFTADELQILTYQLCHTYVRCTRSVSIPAPAYYARLVAFRARYHLVDKEHDSGE 824
Query: 400 GSVASGATTSRGP 362
GS SG + R P
Sbjct: 825 GSHISGQSNGRDP 837
Database: /db/trembl-ebi/tmp/swall
Posted date: Oct 24, 2003 8:22 PM
Number of letters in database: 411,996,502
Number of sequences in database: 1,288,562
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 800,580,766
Number of Sequences: 1288562
Number of extensions: 18409062
Number of successful extensions: 51251
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 48061
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 51004
length of database: 411,996,502
effective HSP length: 123
effective length of database: 253,503,376
effective search space used: 51207681952
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

