BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
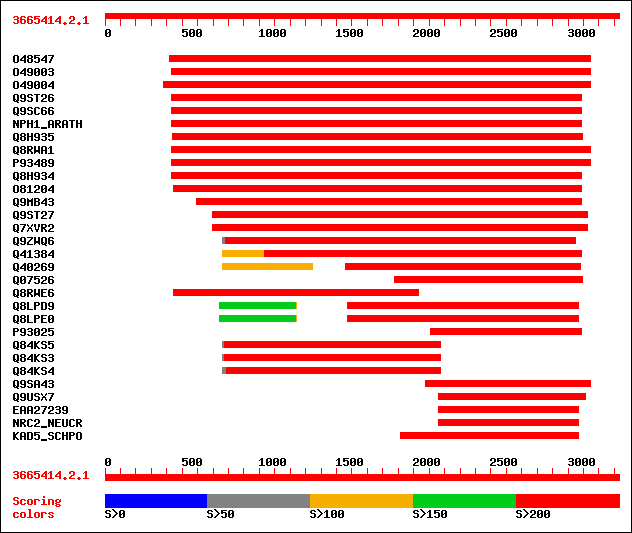
Query= 3665414.2.1
(3330 letters)
Database: /db/trembl-ebi/tmp/swall
1,302,759 sequences; 415,658,970 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O48547|O48547 Nonphototropic hypocotyl 1. 1729 0.0
sptr|O49003|O49003 NPH1-1. 1426 0.0
sptr|O49004|O49004 NPH1-2. 1423 0.0
sptr|Q9ST26|Q9ST26 Nonphototrophic hypocotyl 1a. 1420 0.0
sptr|Q9SC66|Q9SC66 Non-phototropic hypocotyl NPH1. 1408 0.0
sw|O48963|NPH1_ARATH Nonphototropic hypocotyl protein 1 (EC 2.7.... 1158 0.0
sptr|Q8H935|Q8H935 Phototropin. 1144 0.0
sptr|Q8RWA1|Q8RWA1 Phototropin 1. 1135 0.0
sptr|P93489|P93489 Phototropin-like protein PsPK4. 1135 0.0
sptr|Q8H934|Q8H934 Phototropin. 1132 0.0
sptr|O81204|O81204 NONPHOTOTROPIC HYPOCOTYL 2 (Genomic DNA, chro... 1004 0.0
sptr|Q9MB43|Q9MB43 Phototropin. 995 0.0
sptr|Q9ST27|Q9ST27 Nonphototrophic hypocotyl 1b. 975 0.0
sptr|Q7XVR2|Q7XVR2 OSJNBa0079M09.14 protein. 958 0.0
sptr|Q9ZWQ6|Q9ZWQ6 PHY3. 842 0.0
sptr|Q41384|Q41384 Protein kinase (Fragment). 834 0.0
sptr|Q40269|Q40269 Protein kinase (Fragment). 798 0.0
sptr|Q07526|Q07526 Protein kinase PSPK-5 (EC 2.7.1.-) (Probable ... 634 e-180
sptr|Q8RWE6|Q8RWE6 Hypothetical protein (At5g58140). 490 e-137
sptr|Q8LPD9|Q8LPD9 Putative blue light receptor. 479 e-133
sptr|Q8LPE0|Q8LPE0 Putative blue light receptor. 476 e-132
sptr|P93025|P93025 Putative serine/threonine protein kinase (Fra... 473 e-132
sptr|Q84KS5|Q84KS5 Phytochrome 3 (Fragment). 444 e-123
sptr|Q84KS3|Q84KS3 Phytochrome 3 (Fragment). 441 e-122
sptr|Q84KS4|Q84KS4 Phytochrome 3 (Fragment). 426 e-117
sptr|Q9SA43|Q9SA43 F3O9.24 protein. 330 9e-89
sptr|Q9USX7|Q9USX7 Putative ser/thr protein kinase. 303 1e-80
sptrnew|EAA27239|EAA27239 Hypothetical protein. 301 4e-80
sw|O42626|NRC2_NEUCR Serine/threonine protein kinase nrc-2 (EC 2... 301 4e-80
sw|Q09831|KAD5_SCHPO Probable serine/threonine-protein kinase C4... 298 3e-79
>sptr|O48547|O48547 Nonphototropic hypocotyl 1.
Length = 911
Score = 1729 bits (4477), Expect = 0.0
Identities = 867/911 (95%), Positives = 867/911 (95%)
Frame = +1
Query: 415 MAFKGLPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQ 594
MAFKGLPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLL VGRATQ
Sbjct: 1 MAFKGLPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLAAGDGGDADVGRATQ 60
Query: 595 RAAEWGLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRA 774
RAAEWGLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDE LPRVSEELRA
Sbjct: 61 RAAEWGLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDERAAAAGAGRALPRVSEELRA 120
Query: 775 ALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKI 954
ALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKI
Sbjct: 121 ALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKI 180
Query: 955 RQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDT 1134
RQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDT
Sbjct: 181 RQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDT 240
Query: 1135 ALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALV 1314
ALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALV
Sbjct: 241 ALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALV 300
Query: 1315 PGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILK 1494
PGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILK
Sbjct: 301 PGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILK 360
Query: 1495 PREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASD 1674
PREDPLL KKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASD
Sbjct: 361 PREDPLLDSDDERPDSFDDDFRKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASD 420
Query: 1675 SFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWN 1854
SFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWN
Sbjct: 421 SFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWN 480
Query: 1855 LFHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLR 2034
LFHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLR
Sbjct: 481 LFHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLR 540
Query: 2035 PEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVE 2214
PEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVE
Sbjct: 541 PEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVE 600
Query: 2215 LLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVD 2394
LLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVD
Sbjct: 601 LLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVD 660
Query: 2395 YCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGH 2574
YCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGH
Sbjct: 661 YCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGH 720
Query: 2575 ISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPE 2754
ISLTDFDLSCLTSCRPQVFLPHDID NPIFFAEPMRASNSFVGTEEYIAPE
Sbjct: 721 ISLTDFDLSCLTSCRPQVFLPHDIDKKKKRRKSRSNPIFFAEPMRASNSFVGTEEYIAPE 780
Query: 2755 IITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAAR 2934
IITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAAR
Sbjct: 781 IITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAAR 840
Query: 2935 QLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDTLEATLPP 3114
QLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDTLEATLPP
Sbjct: 841 QLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDTLEATLPP 900
Query: 3115 PAADAAHTDMF 3147
PAADAAHTDMF
Sbjct: 901 PAADAAHTDMF 911
>sptr|O49003|O49003 NPH1-1.
Length = 923
Score = 1426 bits (3691), Expect = 0.0
Identities = 725/910 (79%), Positives = 777/910 (85%), Gaps = 4/910 (0%)
Frame = +1
Query: 430 LPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQRAAEW 609
LPRDSRGSLEVFNP + S A P PA S + VG+ATQRAAEW
Sbjct: 23 LPRDSRGSLEVFNPSS--SSAAVEPPSAFRPAARSASPFIEEATGGIEDVGKATQRAAEW 80
Query: 610 GLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRAALSAF 789
GLVLQT+E TGRPQGV AR SG G S +PRVSEELRAALSAF
Sbjct: 81 GLVLQTNEQTGRPQGVSARSSGG------GGSARSSSDDKAVAGAIPRVSEELRAALSAF 134
Query: 790 QQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALA 969
QQTFVVSDA+RP HPI+YASAGFFNMTGY+S EVVGRNCRFLQGSGTDP EI+KIRQALA
Sbjct: 135 QQTFVVSDASRPGHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPAEIAKIRQALA 194
Query: 970 AGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPN 1149
GSNYCGR+LNYKKDGT FWNLLT+APIKDE+GRVLKFIGMQVEVSKYTEGNKDT +RPN
Sbjct: 195 NGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFIGMQVEVSKYTEGNKDTVVRPN 254
Query: 1150 GLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALV--PGK 1323
GLPESLIKYDARQKD ARSSVSELLLA+K+PRSLSES N+T KRKSQES G+ PGK
Sbjct: 255 GLPESLIKYDARQKDQARSSVSELLLAIKNPRSLSESTNSTFKRKSQESVGALTGDRPGK 314
Query: 1324 RSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILKPR- 1500
RSSE+GSRRNS SG R SLQKISEVPE G+K+RKSGL S M L+GMG GN+EK++LKPR
Sbjct: 315 RSSESGSRRNSKSGARTSLQKISEVPERGSKSRKSGLYSLMSLLGMGPGNIEKDMLKPRD 374
Query: 1501 EDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSF 1680
EDPLL +KEMR+GIDLATTLERIEKNFVITDPRLPDNPIIFASDSF
Sbjct: 375 EDPLLDSDDERPESFDDELRRKEMRRGIDLATTLERIEKNFVITDPRLPDNPIIFASDSF 434
Query: 1681 LRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLF 1860
L+LTEY REEILGRNCRFLQGPETDR TV+KIRDAIDNQTEVTVQLINYTKSGKKFWNLF
Sbjct: 435 LQLTEYSREEILGRNCRFLQGPETDRATVRKIRDAIDNQTEVTVQLINYTKSGKKFWNLF 494
Query: 1861 HLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLRPE 2040
HLQPMRDQKGDVQYFIGVQLDGTE VR+AA ++G +L+KKTA+NIDEAAKELPDANLRPE
Sbjct: 495 HLQPMRDQKGDVQYFIGVQLDGTEHVRDAAEREGVMLIKKTAENIDEAAKELPDANLRPE 554
Query: 2041 DLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELL 2220
DLWANHSK VLPKPHMKD+ASWRAIQKVLE GE+IDLKHFRPV+PLGSGDTGSVHLVELL
Sbjct: 555 DLWANHSKVVLPKPHMKDSASWRAIQKVLEGGENIDLKHFRPVKPLGSGDTGSVHLVELL 614
Query: 2221 GTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYC 2400
TGEYFAMKAMDK+VMLNRNKVHRA AER+ILDMLDHPFLPTLYASFQTKTHICLI DY
Sbjct: 615 NTGEYFAMKAMDKNVMLNRNKVHRANAEREILDMLDHPFLPTLYASFQTKTHICLITDYY 674
Query: 2401 AGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHIS 2580
GGELF+LLDRQP+KVL+EDAVRFYAAEVV ALEYLHCQGIIYRDLKPENILLHRDGHIS
Sbjct: 675 PGGELFLLLDRQPLKVLREDAVRFYAAEVVIALEYLHCQGIIYRDLKPENILLHRDGHIS 734
Query: 2581 LTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEII 2760
LTDFDLSCLTSCRPQVFLP + + +PIFFAEPMRASNSFVGTEEYIAPEII
Sbjct: 735 LTDFDLSCLTSCRPQVFLPEEAN-KKSRRKSRSSPIFFAEPMRASNSFVGTEEYIAPEII 793
Query: 2761 TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQL 2940
TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASI VSL ARQL
Sbjct: 794 TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASISVSLPARQL 853
Query: 2941 IYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL-QDTLEATLPPP 3117
IYRLLHRDP+NRLGSYEG+ EIK+HPFFRGINWALVR PP+L+APL D + +
Sbjct: 854 IYRLLHRDPSNRLGSYEGSNEIKEHPFFRGINWALVRGTAPPKLDAPLFPDDTDKGMGDA 913
Query: 3118 AADAAHTDMF 3147
AA HTDMF
Sbjct: 914 AAADTHTDMF 923
>sptr|O49004|O49004 NPH1-2.
Length = 927
Score = 1423 bits (3684), Expect = 0.0
Identities = 729/927 (78%), Positives = 779/927 (84%), Gaps = 5/927 (0%)
Frame = +1
Query: 382 GAGGL*LSHRSMAFKGLPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXX 561
G GG + LPRDSRGSLEVFNP S A P PA S +
Sbjct: 12 GGGGDGYDEPQRPKQQLPRDSRGSLEVFNP----SSSAVEPPSAFRPAARSASPFIEEVA 67
Query: 562 XXXXXVGRATQRAAEWGLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXX 741
VG+ATQRAAEWGLVLQT+E TGRPQGV AR SG G S
Sbjct: 68 GGIEDVGKATQRAAEWGLVLQTNEQTGRPQGVSARSSGG------GGSARSSSDDKAVAG 121
Query: 742 XLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQG 921
+PRVSEELRAALSAFQQTFVVSDA+RP HPI+YASAGFFNMTGY+S EVVGRNCRFLQG
Sbjct: 122 AIPRVSEELRAALSAFQQTFVVSDASRPGHPIMYASAGFFNMTGYTSKEVVGRNCRFLQG 181
Query: 922 SGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVE 1101
SGTDP EI+KIRQALA GSNYCGR+LNYKKDGT FWNLLT+APIKDE+GRVLKFIGMQVE
Sbjct: 182 SGTDPAEIAKIRQALADGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFIGMQVE 241
Query: 1102 VSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKR 1281
VSKYTEGNKDTA+RPNGLPESLIKYDARQKD ARSSVSELLLA+K+PRSLSES N+T KR
Sbjct: 242 VSKYTEGNKDTAVRPNGLPESLIKYDARQKDQARSSVSELLLAIKNPRSLSESTNSTFKR 301
Query: 1282 KSQESAGSALV--PGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLI 1455
KSQES G PGKRSSE+GSRRNS SG R SLQKISEVPE GNK+RKSGL S M L+
Sbjct: 302 KSQESVGPLTGDRPGKRSSESGSRRNSKSGARTSLQKISEVPERGNKSRKSGLYSLMSLL 361
Query: 1456 GMGHGNVEKNILKPR-EDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVIT 1632
GMG GN+EK++LKPR EDPLL +KEMR+GIDLATTLERIEKNFVIT
Sbjct: 362 GMGPGNIEKDMLKPRDEDPLLDSDDERPESFDDELRRKEMRRGIDLATTLERIEKNFVIT 421
Query: 1633 DPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTV 1812
DPRLPDNPIIFASDSFL+LTEY REEILGRNCRFLQGPETDR TV+KIRDAIDNQTEVTV
Sbjct: 422 DPRLPDNPIIFASDSFLQLTEYSREEILGRNCRFLQGPETDRATVRKIRDAIDNQTEVTV 481
Query: 1813 QLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADN 1992
QLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE VR+AA ++G +L+KKTA+N
Sbjct: 482 QLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTEHVRDAAEREGVMLIKKTAEN 541
Query: 1993 IDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVR 2172
IDEAAKELPDANLRPEDLWANHSK VLPKPHMKD+ASWRAIQKVLE GE+IDLKHFRPV+
Sbjct: 542 IDEAAKELPDANLRPEDLWANHSKVVLPKPHMKDSASWRAIQKVLEGGENIDLKHFRPVK 601
Query: 2173 PLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLY 2352
PLGSGDTGSVHLVELL TGEYFAMKAMDK+VMLNRNKVHRA AER+ILDMLDHPFLPTLY
Sbjct: 602 PLGSGDTGSVHLVELLNTGEYFAMKAMDKNVMLNRNKVHRANAEREILDMLDHPFLPTLY 661
Query: 2353 ASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYR 2532
ASFQTKTHICLI DY GGELF+LLDRQP+KVL+EDAVRFYAAEVV ALEYLHCQGIIYR
Sbjct: 662 ASFQTKTHICLITDYYPGGELFLLLDRQPLKVLREDAVRFYAAEVVIALEYLHCQGIIYR 721
Query: 2533 DLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRA 2712
DLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLP + + +P+FFAEPMRA
Sbjct: 722 DLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPEEAN-KKSRRKSRSSPVFFAEPMRA 780
Query: 2713 SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKD 2892
SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKD
Sbjct: 781 SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKD 840
Query: 2893 IRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPEL 3072
IRFPASI VSL ARQLIYRLLHRDP+NRLGSYEG+ EIK+HPFFRGINWALVR PP+L
Sbjct: 841 IRFPASISVSLPARQLIYRLLHRDPSNRLGSYEGSNEIKEHPFFRGINWALVRGTAPPKL 900
Query: 3073 EAPL--QDTLEATLPPPAADAAHTDMF 3147
+APL T + AA HTDMF
Sbjct: 901 DAPLFPDGTDKGLGDATAAADNHTDMF 927
>sptr|Q9ST26|Q9ST26 Nonphototrophic hypocotyl 1a.
Length = 921
Score = 1420 bits (3677), Expect = 0.0
Identities = 727/889 (81%), Positives = 768/889 (86%), Gaps = 4/889 (0%)
Frame = +1
Query: 430 LPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQRAAEW 609
LPRDSRGSLEVFNP + S R + P AS P L V A QRAAEW
Sbjct: 26 LPRDSRGSLEVFNPSSASSFRTAAAA----PKSAS-PFLAIPDREEDNVV--AQQRAAEW 78
Query: 610 GLVLQTDEHTGRPQGVVARPS-GSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRAALSA 786
GLVLQTD HTG PQGV ARPS GS RTS N + +PRVSEELRAALS
Sbjct: 79 GLVLQTDHHTGLPQGVSARPSSGSARTSSEDNPQQQQSAAA-----IPRVSEELRAALSV 133
Query: 787 FQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQAL 966
FQQTFVVSDAT P+HPI+YASAGFFNMTGY+S EVVGRNCRFLQGSGTDP EI KIRQ+L
Sbjct: 134 FQQTFVVSDATHPNHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPHEIDKIRQSL 193
Query: 967 AAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRP 1146
A GSNYCGRILNYKKDGTPFWNLLT+APIKDEDGR+LKFIGMQVEVSKYTEG KDT +RP
Sbjct: 194 ANGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFIGMQVEVSKYTEGKKDTVVRP 253
Query: 1147 NGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSAL--VPG 1320
NGL ESLIKYDARQKDHARSSVSELLLALK+PRSLSES NNTLKRKSQES ++ VP
Sbjct: 254 NGLSESLIKYDARQKDHARSSVSELLLALKNPRSLSESSNNTLKRKSQESLSMSMTEVPS 313
Query: 1321 KRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILKPR 1500
KRSSE+GSRRNS SG R+SLQKI+EVP+ GN+TRKSGLR+FMG +GMGHG+VEKN+LKPR
Sbjct: 314 KRSSESGSRRNSRSGTRSSLQKINEVPDQGNRTRKSGLRAFMGFLGMGHGSVEKNMLKPR 373
Query: 1501 -EDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDS 1677
EDPL+ +KEMR+GIDLATTLERIEKNFVITDPRLPDNPIIFASDS
Sbjct: 374 DEDPLIDSDDERPESFEDEFRRKEMRRGIDLATTLERIEKNFVITDPRLPDNPIIFASDS 433
Query: 1678 FLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNL 1857
FL+LTEY REEILGRNCRFLQGPETDR TV+KIRDAIDNQ EVTVQLINYTKSGKKFWNL
Sbjct: 434 FLQLTEYNREEILGRNCRFLQGPETDRATVRKIRDAIDNQAEVTVQLINYTKSGKKFWNL 493
Query: 1858 FHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLRP 2037
FHLQPMRDQKGDVQYFIGVQLDGTE V++ AAK+G +LVKKTADNIDEAAKELPDANLRP
Sbjct: 494 FHLQPMRDQKGDVQYFIGVQLDGTEHVQDDAAKEGVVLVKKTADNIDEAAKELPDANLRP 553
Query: 2038 EDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVEL 2217
EDLWANHSK VLP PHMKDTASWRAIQKVLE+GESI LKHFRPV+PLGSGDTGSVHLVEL
Sbjct: 554 EDLWANHSKVVLPNPHMKDTASWRAIQKVLESGESIGLKHFRPVKPLGSGDTGSVHLVEL 613
Query: 2218 LGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDY 2397
L TGEYFAMKAMDKS+MLNRNKVHRATAERQILD+LDHPFLPTLYASFQTKTHICLI DY
Sbjct: 614 LNTGEYFAMKAMDKSIMLNRNKVHRATAERQILDLLDHPFLPTLYASFQTKTHICLITDY 673
Query: 2398 CAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHI 2577
C GGELF+LLD QP+KVL EDAVRFYAAEVV ALEYLHCQGIIYRDLKPENILLHRDGHI
Sbjct: 674 CPGGELFVLLDNQPLKVLHEDAVRFYAAEVVVALEYLHCQGIIYRDLKPENILLHRDGHI 733
Query: 2578 SLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEI 2757
SLTDFDLSCLTSCRPQVFLP D D PIFFAEPMRASNSFVGTEEYIAPEI
Sbjct: 734 SLTDFDLSCLTSCRPQVFLPEDADEKKGRKNGSY-PIFFAEPMRASNSFVGTEEYIAPEI 792
Query: 2758 ITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQ 2937
ITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASI VSLAARQ
Sbjct: 793 ITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASISVSLAARQ 852
Query: 2938 LIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL 3084
L+YRLLHRDPANRLGSYEGA EIK HPFFRGINW L+RA PP+LE PL
Sbjct: 853 LMYRLLHRDPANRLGSYEGANEIKGHPFFRGINWPLIRATAPPKLEIPL 901
>sptr|Q9SC66|Q9SC66 Non-phototropic hypocotyl NPH1.
Length = 921
Score = 1408 bits (3644), Expect = 0.0
Identities = 716/888 (80%), Positives = 764/888 (86%), Gaps = 3/888 (0%)
Frame = +1
Query: 430 LPRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQRAAEW 609
LPRDSRGSLEVFNP + S R + A ++ P L V A QRAAEW
Sbjct: 26 LPRDSRGSLEVFNPSSASSFRTAAAA-----AKSASPFLAIPDREEDNVV--AQQRAAEW 78
Query: 610 GLVLQTDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRAALSAF 789
GLVLQTD HTG PQGV RPS + + S ++ +PRVSEELRAALSAF
Sbjct: 79 GLVLQTDYHTGLPQGVSTRPSSCSARTSS----EDTPQQQQSAAAIPRVSEELRAALSAF 134
Query: 790 QQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALA 969
QQTFVVSDATRP+HPI+YASAGFFNMTGY+S EVVGRNCRFLQGSGTDP EI KIRQALA
Sbjct: 135 QQTFVVSDATRPNHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPHEIDKIRQALA 194
Query: 970 AGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPN 1149
GSNYCGRILNYKKDGTPFWNLLT+APIKDEDGR+LKFIGMQVEVSKYTEG K+T +RPN
Sbjct: 195 NGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFIGMQVEVSKYTEGKKETVVRPN 254
Query: 1150 GLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSAL--VPGK 1323
GL ESLIKYDARQKDHARSSVSELLLALK+PRSLSES NNTLKRKSQES ++ VP K
Sbjct: 255 GLSESLIKYDARQKDHARSSVSELLLALKNPRSLSESSNNTLKRKSQESLSMSMSEVPSK 314
Query: 1324 RSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKNILKPR- 1500
RSSE+GSRRNS SG R+SLQKI+EVP+ N+TRKSGLR+FMG +GMGHG+VEKN+LKPR
Sbjct: 315 RSSESGSRRNSRSGTRSSLQKINEVPDQVNRTRKSGLRAFMGFLGMGHGSVEKNMLKPRD 374
Query: 1501 EDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSF 1680
EDPL+ +KEMR+GIDLATTLERIEKNFVITDPRLPDNPIIFASDSF
Sbjct: 375 EDPLIDSDDERPESFEDEFRRKEMRRGIDLATTLERIEKNFVITDPRLPDNPIIFASDSF 434
Query: 1681 LRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLF 1860
L+LTEY REEILGRNCRFLQGPETDR V+KIRDAIDNQ EVTVQLINYTKSGKKFWNLF
Sbjct: 435 LQLTEYNREEILGRNCRFLQGPETDRAIVRKIRDAIDNQAEVTVQLINYTKSGKKFWNLF 494
Query: 1861 HLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLRPE 2040
HLQPMRDQKGDVQYFIGVQLDGTE V++ AA++G +LVKKTADNIDEAAKELPDANLRPE
Sbjct: 495 HLQPMRDQKGDVQYFIGVQLDGTEHVQDDAAEEGVVLVKKTADNIDEAAKELPDANLRPE 554
Query: 2041 DLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELL 2220
DLWANHSK VLP PHMKDTASWRAIQKVLE+GESI LKHFRP++PLGSGDTGSVHLVELL
Sbjct: 555 DLWANHSKVVLPNPHMKDTASWRAIQKVLESGESIGLKHFRPIKPLGSGDTGSVHLVELL 614
Query: 2221 GTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYC 2400
TGEYFAMKAMDKS+MLNRNKVHRATAERQILD+LDHPFLPTLYASFQTKTHICLI DYC
Sbjct: 615 NTGEYFAMKAMDKSIMLNRNKVHRATAERQILDLLDHPFLPTLYASFQTKTHICLITDYC 674
Query: 2401 AGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHIS 2580
GGELF+LLD QP+KVL EDAVRFY AEVV ALEYLHCQGIIYRDLKPENILLHRDGHIS
Sbjct: 675 PGGELFVLLDNQPLKVLHEDAVRFYVAEVVIALEYLHCQGIIYRDLKPENILLHRDGHIS 734
Query: 2581 LTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEII 2760
LTDFDLSCLTSCRPQVFLP D D PIFFAEPMRASNSFVGTEEYIAPEII
Sbjct: 735 LTDFDLSCLTSCRPQVFLPEDADEKKGRKNRSY-PIFFAEPMRASNSFVGTEEYIAPEII 793
Query: 2761 TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQL 2940
TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASI VSLAARQL
Sbjct: 794 TGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASILVSLAARQL 853
Query: 2941 IYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL 3084
+YRLLHRDPANRLGSYEGA EIK HPFFRGINW L+RA PP+LE PL
Sbjct: 854 MYRLLHRDPANRLGSYEGANEIKGHPFFRGINWPLIRATAPPKLEIPL 901
>sw|O48963|NPH1_ARATH Nonphototropic hypocotyl protein 1 (EC 2.7.1.37)
(Phototropin).
Length = 996
Score = 1158 bits (2996), Expect = 0.0
Identities = 620/960 (64%), Positives = 701/960 (73%), Gaps = 75/960 (7%)
Frame = +1
Query: 430 LPRDSRGSLEVFNP-------DAPV-----------SDRATTSPFLLP---PA------- 525
LPRD+RGSLEVFNP D PV SD TSP P PA
Sbjct: 16 LPRDTRGSLEVFNPSTQLTRPDNPVFRPEPPAWQNLSDPRGTSPQPRPQQEPAPSNPVRS 75
Query: 526 ---VASHPSLLXXXXXXXXXVGRAT--------------QRAAEWGLVLQTDEHTGRPQG 654
+A S + + + T QRAAEWGLVL+TD TG+PQG
Sbjct: 76 DQEIAVTTSWMALKDPSPETISKKTITAEKPQKSAVAAEQRAAEWGLVLKTDTKTGKPQG 135
Query: 655 VVARPSGSNRTSESGNSIDEXXXXXXX---------------XXXLPRVSEELRAALSAF 789
V R SG +G +PRVSE+L+ ALS F
Sbjct: 136 VGVRNSGGTENDPNGKKTTSQRNSQNSCRSSGEMSDGDVPGGRSGIPRVSEDLKDALSTF 195
Query: 790 QQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALA 969
QQTFVVSDAT+PD+PI+YASAGFFNMTGY+S EVVGRNCRFLQGSGTD E++KIR+ LA
Sbjct: 196 QQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDADELAKIRETLA 255
Query: 970 AGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPN 1149
AG+NYCGRILNYKKDGT FWNLLT+APIKDE G+VLKFIGMQVEVSK+TEG K+ ALRPN
Sbjct: 256 AGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFIGMQVEVSKHTEGAKEKALRPN 315
Query: 1150 GLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALVPGKRS 1329
GLPESLI+YDARQKD A +SV+EL+ A+K PR+LSES N ES P +R
Sbjct: 316 GLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTNLHPFMTKSESDELPKKPARRM 375
Query: 1330 SET---GSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHG---NVEKNIL 1491
SE RRNS G RNS+Q+I+E+PE K+RKS L SFMG+ +++ +
Sbjct: 376 SENVVPSGRRNSGGGRRNSMQRINEIPE--KKSRKSSL-SFMGIKKKSESLDESIDDGFI 432
Query: 1492 K-PREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFA 1668
+ ED + +KEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFA
Sbjct: 433 EYGEEDDEISDRDERPESVDDKVRQKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFA 492
Query: 1669 SDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKF 1848
SDSFL LTEY REEILGRNCRFLQGPETD TVKKIR+AIDNQTEVTVQLINYTKSGKKF
Sbjct: 493 SDSFLELTEYSREEILGRNCRFLQGPETDLTTVKKIRNAIDNQTEVTVQLINYTKSGKKF 552
Query: 1849 WNLFHLQPMRDQKGDVQYFIGVQLDGTERVR-------EAAAKDGAILVKKTADNIDEAA 2007
WN+FHLQPMRDQKG+VQYFIGVQLDG++ V E A K+G LVKKTA NIDEA
Sbjct: 553 WNIFHLQPMRDQKGEVQYFIGVQLDGSKHVEPVRNVIEETAVKEGEDLVKKTAVNIDEAV 612
Query: 2008 KELPDANLRPEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSG 2187
+ELPDAN+ PEDLWANHSK V KPH KD+ W AIQKVLE+GE I LKHF+PV+PLGSG
Sbjct: 613 RELPDANMTPEDLWANHSKVVHCKPHRKDSPPWIAIQKVLESGEPIGLKHFKPVKPLGSG 672
Query: 2188 DTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQT 2367
DTGSVHLVEL+GT + FAMKAMDK+VMLNRNKVHRA AER+ILD+LDHPFLP LYASFQT
Sbjct: 673 DTGSVHLVELVGTDQLFAMKAMDKAVMLNRNKVHRARAEREILDLLDHPFLPALYASFQT 732
Query: 2368 KTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPE 2547
KTHICLI DY GGELFMLLDRQP KVLKEDAVRFYAA+VV ALEYLHCQGIIYRDLKPE
Sbjct: 733 KTHICLITDYYPGGELFMLLDRQPRKVLKEDAVRFYAAQVVVALEYLHCQGIIYRDLKPE 792
Query: 2548 NILLHRDGHISLTDFDLSCLTSCRPQVFLPH-DIDXXXXXXXXXXNPIFFAEPMRASNSF 2724
N+L+ +G ISL+DFDLSCLTSC+PQ+ +P D PIF AEPMRASNSF
Sbjct: 793 NVLIQGNGDISLSDFDLSCLTSCKPQLLIPSIDEKKKKKQQKSQQTPIFMAEPMRASNSF 852
Query: 2725 VGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFP 2904
VGTEEYIAPEII+GAGHTSAVDWWALGIL+YEMLYGYTPFRGKTRQ+TF N+L KD++FP
Sbjct: 853 VGTEEYIAPEIISGAGHTSAVDWWALGILMYEMLYGYTPFRGKTRQKTFTNVLQKDLKFP 912
Query: 2905 ASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL 3084
ASI SL +QLI+RLL RDP RLG +EGA E+KQH FF+GINWAL+R PPELE P+
Sbjct: 913 ASIPASLQVKQLIFRLLQRDPKKRLGCFEGANEVKQHSFFKGINWALIRCTNPPELETPI 972
>sptr|Q8H935|Q8H935 Phototropin.
Length = 963
Score = 1144 bits (2959), Expect = 0.0
Identities = 607/949 (63%), Positives = 691/949 (72%), Gaps = 63/949 (6%)
Frame = +1
Query: 436 RDSRGSLEVFNPDAP---------------------VSDRATTSPFLLPPAVASHPSLLX 552
RD RGSLEVFNP + S R T +P L ++
Sbjct: 6 RDHRGSLEVFNPSSSETNGTPNPNPNPNPSNSWNTGTSSRGTEAPPLRDSIISDEVPTAT 65
Query: 553 XXXXXXXXVGR---------ATQRAAEWGLVLQTDEHTGRPQGVVARPSGSNRTS--ESG 699
A QRAAEWGLVL+TD TG+PQGV R SG S +S
Sbjct: 66 SWMALKETTPSPKSGESGSVAEQRAAEWGLVLKTDSETGKPQGVGVRGSGGGGGSRRDSN 125
Query: 700 NSI----DEXXXXXXXXXXLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNM 867
NS+ + +PRVSE+LR ALSAFQQTFVVSDAT+PD+PI+YASAGFF+M
Sbjct: 126 NSVRSSGESSDDGREGGRGIPRVSEDLRDALSAFQQTFVVSDATKPDYPIMYASAGFFSM 185
Query: 868 TGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVA 1047
TGY+S EV+GRNCRF+QG+ TDP +++KIR+ALAAG++YCGR+LNYKKDGT FWNLLT+A
Sbjct: 186 TGYTSKEVIGRNCRFMQGADTDPNDVAKIREALAAGTSYCGRLLNYKKDGTTFWNLLTIA 245
Query: 1048 PIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLL 1227
PIKDE G++LK IGMQVEVSK+TEG K+ LRPNGLPESLI+YDARQK+ A SSV+EL+
Sbjct: 246 PIKDEHGKILKLIGMQVEVSKHTEGTKEKMLRPNGLPESLIRYDARQKEKANSSVTELVE 305
Query: 1228 AL-KDPRSLSESRNNTL--KRKSQESAGSALVPGKRSSETGS-------RRNSHSGM--R 1371
A+ K PRSLSES N K+ + S A P SS S RR SHSG
Sbjct: 306 AVSKRPRSLSESANRLPFNKKPTNGSNDHATPPNSESSSRKSGSTLRSFRRKSHSGAGNS 365
Query: 1372 NSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEK--------NILKPREDPLLXXXX 1527
NS+ I+E+PE NK+R+ RSFMG + N E+ ED L
Sbjct: 366 NSMHPITELPENNNKSRR---RSFMGFMRKSLSNNERFNHEQVIDRNSSEDEDRL----- 417
Query: 1528 XXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCRE 1707
K+E RKG DLATTLERIEKNFVITDPRLPDNPIIFASDSFL LTEY RE
Sbjct: 418 -DSFDEQNIAQKREKRKGFDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSRE 476
Query: 1708 EILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQK 1887
EILGRNCRFLQGPETD TVKKIR AIDNQTEVTVQLINYTK+GKKFWNLFHLQPMRDQK
Sbjct: 477 EILGRNCRFLQGPETDPATVKKIRYAIDNQTEVTVQLINYTKTGKKFWNLFHLQPMRDQK 536
Query: 1888 GDVQYFIGVQLDGTERVR-------EAAAKDGAILVKKTADNIDEAAKELPDANLRPEDL 2046
G+VQYFIGVQLDG++ V E AK+G LVKKTA+N+D+A +ELPDAN++PEDL
Sbjct: 537 GEVQYFIGVQLDGSQHVEPLHNGIAEDTAKEGENLVKKTAENVDDALRELPDANMKPEDL 596
Query: 2047 WANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGT 2226
W NHSK V PKPH ++ ++WRAIQK++E+GE I LKHF+P++PLGSGDTGSVHLVEL GT
Sbjct: 597 WMNHSKVVHPKPHRREDSAWRAIQKIMESGEQIGLKHFKPIKPLGSGDTGSVHLVELCGT 656
Query: 2227 GEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAG 2406
+FAMKAMDK VM NRNKVHRA ER+ILDMLDHPFLP LYASFQTKTHICLI DYC G
Sbjct: 657 DHHFAMKAMDKGVMPNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHICLITDYCPG 716
Query: 2407 GELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLT 2586
GELFMLLDRQP KVLKEDAVRFYA EVV ALEYLHCQGIIYRDLKPEN+LL GH+SLT
Sbjct: 717 GELFMLLDRQPAKVLKEDAVRFYATEVVVALEYLHCQGIIYRDLKPENVLLQSTGHVSLT 776
Query: 2587 DFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITG 2766
DFDLSCLTSC+P++ +P D PIF AEPMRASNSFVGTEEYIAPEIITG
Sbjct: 777 DFDLSCLTSCKPELIVPSTND----KKKGQHGPIFMAEPMRASNSFVGTEEYIAPEIITG 832
Query: 2767 AGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIY 2946
+GHT AVDWWALGILLYEM YGYTPFRGK RQRTFANILHKD++ P S QVSL+A+QLIY
Sbjct: 833 SGHTCAVDWWALGILLYEMFYGYTPFRGKNRQRTFANILHKDLKLPKSKQVSLSAKQLIY 892
Query: 2947 RLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDT 3093
LL RDP +RLGS GA +IK H FF+GINWALVR PPEL+APL DT
Sbjct: 893 HLLQRDPTSRLGSKGGANDIKHHSFFKGINWALVRCTKPPELDAPLFDT 941
>sptr|Q8RWA1|Q8RWA1 Phototropin 1.
Length = 976
Score = 1135 bits (2937), Expect = 0.0
Identities = 597/960 (62%), Positives = 691/960 (71%), Gaps = 55/960 (5%)
Frame = +1
Query: 433 PRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHP---------------SLLXXXXXX 567
PRD RGSLEVFNP + S + L P +
Sbjct: 21 PRDPRGSLEVFNPTSNSSSPVRSPSNLKNWTEIEEPRNELSEQHNEFSDEVTNTSWMAIK 80
Query: 568 XXXVGRATQRAAEWGLVLQTDEHTGRPQGVVARPSGSNRTS-------ESGNSIDEXXXX 726
G A QRAAEWGL+L TD TG+PQGV R SG + S S N++
Sbjct: 81 EGETGAAVQRAAEWGLMLTTDAETGKPQGVAVRNSGGDEPSVKLETKRNSNNTVRTSGES 140
Query: 727 XXXXX--XLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGR 900
PRVSE+L+ ALSAFQQTFVVSDAT+PD+PILYASAGFF MTGY+S EV+GR
Sbjct: 141 SDGDDPRGFPRVSEDLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGR 200
Query: 901 NCRFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLK 1080
NCRFLQG+ TDP ++++IR+AL G ++CGR+LNYKKDGTPFWNLLT++PIKD+DG VLK
Sbjct: 201 NCRFLQGADTDPDDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLK 260
Query: 1081 FIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSES 1260
IGM VEV+K+TEG+K+ LRPNGLPESLI+YDARQK+ A SSVSELL A+K PR+LSES
Sbjct: 261 LIGMLVEVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRALSES 320
Query: 1261 RNNTLKRKSQESAGSALVPGKRSSETGSRRNSHS----------GMRNSLQKISEVPEGG 1410
RKS GS + + E SRR S S +R+S+++ISE+PE
Sbjct: 321 GQRPFIRKSGGGGGSE--EDEEAVENKSRRKSDSVASFRPKPQGKIRHSMERISELPE-- 376
Query: 1411 NKTRKSGLRSFMGLIGMGHG---NVEKNILKPREDPLLXXXXXXXXXXXXXXXKKEMRKG 1581
NK + S SFMG + +++ ++ +E RKG
Sbjct: 377 NKQKNSRRGSFMGFMRKSDSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLREKRKG 436
Query: 1582 IDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRG 1761
+DLATTLERIEKNFVITDPRLPDNPIIFASDSFL LTEY REEILG+NCRFLQGPETD
Sbjct: 437 LDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGPETDPA 496
Query: 1762 TVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVR 1941
TV+KIR+AIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRD KG+VQYFIGVQLDG++ V
Sbjct: 497 TVRKIREAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVE 556
Query: 1942 -------EAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTA 2100
E AK+G +LVK+TA+N+ EA KELPDAN +P+DLW NHSK V PKPH KD
Sbjct: 557 PLHNCIAEDTAKEGELLVKETAENVGEAVKELPDANQKPDDLWMNHSKVVRPKPHRKDDD 616
Query: 2101 SWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRN 2280
+WRAIQKVLENGE + LKHFRP++PLGSGDTGSVHLVEL GTG+YFAMKAMDK VMLNRN
Sbjct: 617 AWRAIQKVLENGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRN 676
Query: 2281 KVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKED 2460
KVHRA ER+ILDMLDHPFLP LYASFQTKTH+CLI DY +GGELF+LLD+QP KVLKED
Sbjct: 677 KVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYSGGELFLLLDQQPTKVLKED 736
Query: 2461 AVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPH 2640
+VRFYAAEVV ALEYLHC GIIYRDLKPEN+L+ +GH+SLTDFDLSCLTSC+PQ+ LP
Sbjct: 737 SVRFYAAEVVIALEYLHCLGIIYRDLKPENVLIQSNGHVSLTDFDLSCLTSCKPQLILPA 796
Query: 2641 DIDXXXXXXXXXXN-------PIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWA 2799
+ P+F AEPMRASNSFVGTEEYIAPEIITG+GHTSAVDWWA
Sbjct: 797 IEEKKKRKKKKNKGQQKNQQVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWA 856
Query: 2800 LGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRL 2979
LGILLYEMLYGYTPFRGKTRQ+TFANILHKD++FP S VS +QLIY LLHRDP NRL
Sbjct: 857 LGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPHGKQLIYWLLHRDPKNRL 916
Query: 2980 GSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL----QDTLEATLPPPAADAAHTDMF 3147
GS EGA EIK HPFF+ INWALVR PPEL+ P+ + EA P D ++F
Sbjct: 917 GSLEGANEIKNHPFFKNINWALVRCTKPPELDGPILLDNDEKKEAKEIDPGLDDLQKNIF 976
>sptr|P93489|P93489 Phototropin-like protein PsPK4.
Length = 976
Score = 1135 bits (2935), Expect = 0.0
Identities = 597/960 (62%), Positives = 690/960 (71%), Gaps = 55/960 (5%)
Frame = +1
Query: 433 PRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHP---------------SLLXXXXXX 567
PRD RGSLEVFNP + S + L P +
Sbjct: 21 PRDPRGSLEVFNPTSNSSSPVRSPSNLKNWTEIEEPRNELSEQHNEFSDEVTNTSWMAIK 80
Query: 568 XXXVGRATQRAAEWGLVLQTDEHTGRPQGVVARPSGSNRTS-------ESGNSIDEXXXX 726
G A QRAAEWGL+L TD TG+PQGV R SG + S S N++
Sbjct: 81 EGETGAAVQRAAEWGLMLTTDAETGKPQGVAVRNSGGDEPSVKLETKRNSNNTVRTSGES 140
Query: 727 XXXXX--XLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGR 900
PRVSE+L+ ALSAFQQTFVVSDAT+PD+PILYASAGFF MTGY+S EV+GR
Sbjct: 141 SDGDDPRGFPRVSEDLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGR 200
Query: 901 NCRFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLK 1080
NCRFLQG+ TDP ++++IR+AL G ++CGR+LNYKKDGTPFWNLLT++PIKD+DG VLK
Sbjct: 201 NCRFLQGADTDPDDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLK 260
Query: 1081 FIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSES 1260
IGM VEV+K+TEG+K+ LRPNGLPESLI+YDARQK+ A SSVSELL A+K PR+LSES
Sbjct: 261 LIGMLVEVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRALSES 320
Query: 1261 RNNTLKRKSQESAGSALVPGKRSSETGSRRNSHS----------GMRNSLQKISEVPEGG 1410
RKS GS + + E SRR S S +R+S+++ISE+PE
Sbjct: 321 GQRPFIRKSGGGGGSE--EDEEAVENKSRRKSDSVASFRPKPQGKIRHSMERISELPE-- 376
Query: 1411 NKTRKSGLRSFMGLIGMGHG---NVEKNILKPREDPLLXXXXXXXXXXXXXXXKKEMRKG 1581
NK + S SFMG + +++ ++ +E RKG
Sbjct: 377 NKQKNSRRGSFMGFMRKSDSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLREKRKG 436
Query: 1582 IDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRG 1761
+DLATTLERIEKNFVITDPRLPDNPIIFASDSFL LTEY REEILG+NCRFLQGPETD
Sbjct: 437 LDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGPETDPA 496
Query: 1762 TVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVR 1941
TV+KIR+AIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRD KG+VQYFIGVQLDG++ V
Sbjct: 497 TVRKIREAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVE 556
Query: 1942 -------EAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTA 2100
E AK+G +LVK+TA+N+ EA KELPDAN +P+DLW NHSK V PKPH KD
Sbjct: 557 PLHNCIAEDTAKEGELLVKETAENVGEAVKELPDANQKPDDLWMNHSKVVRPKPHRKDDD 616
Query: 2101 SWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRN 2280
+WRAIQKVLENGE + LKHFRP++PLGSGDTGSVHLVEL GTG+YFAMKAMDK VMLNRN
Sbjct: 617 AWRAIQKVLENGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRN 676
Query: 2281 KVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKED 2460
KVHRA ER+ILDMLDHPFLP LYASFQTKTH+CLI DY GGELF+LLD+QP KVLKED
Sbjct: 677 KVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKED 736
Query: 2461 AVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPH 2640
+VRFYAAEVV ALEYLHC GIIYRDLKPEN+L+ +GH+SLTDFDLSCLTSC+PQ+ LP
Sbjct: 737 SVRFYAAEVVIALEYLHCLGIIYRDLKPENVLIQSNGHVSLTDFDLSCLTSCKPQLILPA 796
Query: 2641 DIDXXXXXXXXXXN-------PIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWA 2799
+ P+F AEPMRASNSFVGTEEYIAPEIITG+GHTSAVDWWA
Sbjct: 797 IEEKKKRKKKKNKGQQKNQQVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWA 856
Query: 2800 LGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRL 2979
LGILLYEMLYGYTPFRGKTRQ+TFANILHKD++FP S VS +QLIY LLHRDP NRL
Sbjct: 857 LGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPHGKQLIYWLLHRDPKNRL 916
Query: 2980 GSYEGAMEIKQHPFFRGINWALVRAATPPELEAPL----QDTLEATLPPPAADAAHTDMF 3147
GS EGA EIK HPFF+ INWALVR PPEL+ P+ + EA P D ++F
Sbjct: 917 GSLEGANEIKNHPFFKNINWALVRCTKPPELDGPILLDNDEKKEAKEIDPGLDDLQKNIF 976
>sptr|Q8H934|Q8H934 Phototropin.
Length = 970
Score = 1132 bits (2927), Expect = 0.0
Identities = 588/929 (63%), Positives = 677/929 (72%), Gaps = 45/929 (4%)
Frame = +1
Query: 433 PRDSRGSLEVFNPDAPVSDRATTSPFLLPPAVASHP--------SLLXXXXXXXXXVGRA 588
PRD RGSLEVFNP + S + L P + G A
Sbjct: 22 PRDPRGSLEVFNPTSNTSSPVRSPSNLKNWTETEEPRNEFPDKVTNTSWMAIKEGETGAA 81
Query: 589 TQRAAEWGLVLQTDEHTGRPQGVVARPSG----------SNRTSESGNSIDEXXXXXXXX 738
QRAAEWGLVL TD TG+PQGV R SG S R S +
Sbjct: 82 VQRAAEWGLVLTTDAETGKPQGVAVRHSGGDEPNAVELESKRNSNNTVRTSGESSDGGDP 141
Query: 739 XXLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQ 918
PRVS++L+ ALSAFQQTFVVSDAT+PD+PILYASAGFF MTGY+S EV+GRNCRFLQ
Sbjct: 142 RGFPRVSDDLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGRNCRFLQ 201
Query: 919 GSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQV 1098
G+ TDP ++++IR+AL G ++CGR+LNYKKDGTPFWNLLT++PIKD+DG VLK IGM V
Sbjct: 202 GADTDPNDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLV 261
Query: 1099 EVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLK 1278
EV+K+TEG+K+ LRPNGLPESLI+YDARQK+ A SSVSELL A+K PR++SES +
Sbjct: 262 EVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRAMSESGHRPFI 321
Query: 1279 RKSQESAGSALVPGKRSSETGSRRNSHS----------GMRNSLQKISEVPEGGNKTRKS 1428
RKS G E SRR S S +R+S+++ISE+PE NK + S
Sbjct: 322 RKS---GGGGSSEEDERLENKSRRKSDSVASFRPKPQGKIRHSMERISELPE--NKQKNS 376
Query: 1429 GLRSFMGLIGMGHG---NVEKNILKPREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATT 1599
SFMG + H +++ ++ KE RKG+DLATT
Sbjct: 377 RRGSFMGFMRKSHSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLKEKRKGLDLATT 436
Query: 1600 LERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIR 1779
LERIEKNFVITDPRLPDNPIIFASDSFL LTEY REEILG+NCRFLQG ETD TV+KIR
Sbjct: 437 LERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGQETDPATVRKIR 496
Query: 1780 DAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVR------ 1941
+AIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRD KG+VQYFIGVQLDG++ V
Sbjct: 497 EAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVEPLHNCI 556
Query: 1942 -EAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTASWRAIQ 2118
E +AK+G +LVK+TA+N+ EA KELPDAN +P+DLW NHSK V PKPH KD +WRAIQ
Sbjct: 557 AEESAKEGELLVKETAENVGEAVKELPDANQKPDDLWKNHSKVVRPKPHRKDDDAWRAIQ 616
Query: 2119 KVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRAT 2298
V+ NGE + LKHFRP++PLGSGDTGSVHLVEL GTG YFAMKAMDK VMLNRNKVHRA
Sbjct: 617 NVVGNGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGHYFAMKAMDKGVMLNRNKVHRAC 676
Query: 2299 AERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYA 2478
ER+ILDMLDHPFLP LYASFQTKTH+CLI DY GGELF+LLD+QP KVLKED+VRFYA
Sbjct: 677 TEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDSVRFYA 736
Query: 2479 AEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDIDXXX 2658
AEVV ALEYLHC GIIYRDLKPEN+L+ R+GH+SLTDFDLSCLTSC+PQ+ LP +
Sbjct: 737 AEVVIALEYLHCLGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLILPATEEKKK 796
Query: 2659 XXXXXXXN-------PIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLY 2817
P+F AEPMRASNSFVGTEEYIAPEIITG+GHTSAVDWWALGILLY
Sbjct: 797 RKNKKKKGQPKNQEVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLY 856
Query: 2818 EMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGA 2997
EMLYGYTPFRGKTRQ+TF NILHKD++FP S VS +QLIY LLHRDP NRLGS EGA
Sbjct: 857 EMLYGYTPFRGKTRQKTFGNILHKDLKFPKSKPVSPHGKQLIYWLLHRDPKNRLGSLEGA 916
Query: 2998 MEIKQHPFFRGINWALVRAATPPELEAPL 3084
EIK HPFF+ +NWALVR PPEL+AP+
Sbjct: 917 NEIKNHPFFKNVNWALVRCMKPPELDAPI 945
>sptr|O81204|O81204 NONPHOTOTROPIC HYPOCOTYL 2 (Genomic DNA,
chromosome 5, P1 clone:MCK7).
Length = 915
Score = 1004 bits (2595), Expect = 0.0
Identities = 521/903 (57%), Positives = 625/903 (69%), Gaps = 23/903 (2%)
Frame = +1
Query: 445 RGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQRAAEWGLVLQ 624
R SLE+FNP + +TS PP ++ + T+R AEWGL
Sbjct: 20 RRSLEIFNPSSGKETHGSTSSSSKPPLDGNNKGSSSKWMEFQDSA-KITERTAEWGLSAV 78
Query: 625 TDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRAALSAFQQTFV 804
+ G+ + S S++ + PRVS+EL+ ALS QQTFV
Sbjct: 79 KPD--SGDDGISFKLSSEVERSKNMSRRSSEESTSSESGAFPRVSQELKTALSTLQQTFV 136
Query: 805 VSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALAAGSNY 984
VSDAT+P PI+YAS+GFF MTGYSS E+VGRNCRFLQG TD E++KIR + G +Y
Sbjct: 137 VSDATQPHCPIVYASSGFFTMTGYSSKEIVGRNCRFLQGPDTDKNEVAKIRDCVKNGKSY 196
Query: 985 CGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPES 1164
CGR+LNYKKDGTPFWNLLTV PIKD+ G +KFIGMQVEVSKYTEG D ALRPNGL +S
Sbjct: 197 CGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFIGMQVEVSKYTEGVNDKALRPNGLSKS 256
Query: 1165 LIKYDARQKDHARSSVSELLLALKDPRS-LSESRNNTLK---------------RKSQES 1296
LI+YDARQK+ A S++E++ ++ +S + ES +N R+S E+
Sbjct: 257 LIRYDARQKEKALDSITEVVQTIRHRKSQVQESVSNDTMVKPDSSTTPTPGRQTRQSDEA 316
Query: 1297 AGSALVPGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNV 1476
+ S PG+ S+ TGS+ S + L +
Sbjct: 317 SKSFRTPGRVSTPTGSKLKSSNNRHEDLLR------------------------------ 346
Query: 1477 EKNILKPREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNP 1656
++P E L ++++R+GIDLATTLERIEKNFVI+DPRLPDNP
Sbjct: 347 ----MEPEELMLSTEVIGQRDSWDLSDRERDIRQGIDLATTLERIEKNFVISDPRLPDNP 402
Query: 1657 IIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKS 1836
IIFASDSFL LTEY REEILGRNCRFLQGPETD+ TV+KIRDAI +Q E+TVQLINYTKS
Sbjct: 403 IIFASDSFLELTEYSREEILGRNCRFLQGPETDQATVQKIRDAIRDQREITVQLINYTKS 462
Query: 1837 GKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE-------RVREAAAKDGAILVKKTADNI 1995
GKKFWNLFHLQPMRDQKG++QYFIGVQLDG++ R+ E + LVK TA N+
Sbjct: 463 GKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHVEPLQNRLSERTEMQSSKLVKATATNV 522
Query: 1996 DEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRP 2175
DEA +ELPDAN RPEDLWA HSKPV P PH K++ SW+AI+K+ +GE++ L HF+P++P
Sbjct: 523 DEAVRELPDANTRPEDLWAAHSKPVYPLPHNKESTSWKAIKKIQASGETVGLHHFKPIKP 582
Query: 2176 LGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYA 2355
LGSGDTGSVHLVEL GTGE +AMKAM+K++MLNRNK HRA ER+I+ +LDHPFLPTLYA
Sbjct: 583 LGSGDTGSVHLVELKGTGELYAMKAMEKTMMLNRNKAHRACIEREIISLLDHPFLPTLYA 642
Query: 2356 SFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRD 2535
SFQT TH+CLI D+C GGELF LLDRQPMK+L ED+ RFYAAEVV LEYLHC GI+YRD
Sbjct: 643 SFQTSTHVCLITDFCPGGELFALLDRQPMKILTEDSARFYAAEVVIGLEYLHCLGIVYRD 702
Query: 2536 LKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRAS 2715
LKPENILL +DGHI L DFDLS +T+C PQ+ +P P F AEP S
Sbjct: 703 LKPENILLKKDGHIVLADFDLSFMTTCTPQLIIP-AAPSKRRRSKSQPLPTFVAEPSTQS 761
Query: 2716 NSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDI 2895
NSFVGTEEYIAPEIITGAGHTSA+DWWALGILLYEMLYG TPFRGK RQ+TFANILHKD+
Sbjct: 762 NSFVGTEEYIAPEIITGAGHTSAIDWWALGILLYEMLYGRTPFRGKNRQKTFANILHKDL 821
Query: 2896 RFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELE 3075
FP+SI VSL RQLI LL+RDP++RLGS GA EIKQH FFRGINW L+R +PP L+
Sbjct: 822 TFPSSIPVSLVGRQLINTLLNRDPSSRLGSKGGANEIKQHAFFRGINWPLIRGMSPPPLD 881
Query: 3076 APL 3084
APL
Sbjct: 882 APL 884
>sptr|Q9MB43|Q9MB43 Phototropin.
Length = 1092
Score = 995 bits (2573), Expect = 0.0
Identities = 510/873 (58%), Positives = 628/873 (71%), Gaps = 42/873 (4%)
Frame = +1
Query: 592 QRAAEWGLVLQTDEHTGRPQGVVARPSGSNRT----SESGNSIDEXX----------XXX 729
+RAA WGLVL+TD +GR GV R S R SE S++
Sbjct: 198 ERAAGWGLVLKTDGDSGRVDGVRTRTSEEEREFRRLSEERRSLNSSTVRTSDDSGFTSDT 257
Query: 730 XXXXXLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCR 909
+PRVS+++ AL FQQTFV++D T+PD PI+YASAGFF MTGY+S+EV+GRNCR
Sbjct: 258 SNASRIPRVSKDVLQALEGFQQTFVIADGTKPDLPIMYASAGFFKMTGYTSSEVIGRNCR 317
Query: 910 FLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIG 1089
FLQG TDP EI +IR+ ++ GS YCGR+LNYKKDG+ FWNLLT++PIKD DG VLK+IG
Sbjct: 318 FLQGKETDPEEIDRIRECISKGSGYCGRLLNYKKDGSAFWNLLTISPIKDVDGSVLKYIG 377
Query: 1090 MQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNN 1269
MQVEVS++TEG K+ A+RPNGL ESLIKYDARQK+ A V+EL+ A+KDP+ + + +
Sbjct: 378 MQVEVSQFTEGTKENAMRPNGLSESLIKYDARQKERASFQVTELVEAIKDPKQVGDDKKT 437
Query: 1270 TLKR---KSQESAGSALVPGKRSSETGSRRNSHSGMRNSLQ-KISEVPEGGN-------K 1416
++ + +G P R T + S S+Q K S + E G+ +
Sbjct: 438 SISLGVVRPVAPSGPRKGPDLRELLTAQQYTPRSA--TSVQPKGSNLTEPGSHRESISKR 495
Query: 1417 TRKSGLRSFMGL---IGMGHGN-------VEKNILKPREDPLLXXXXXXXXXXXXXXXKK 1566
R SG SF+GL G G GN +E IL +++ K
Sbjct: 496 NRSSGFFSFLGLDKLAGKGPGNQHDAAEFIEPEILMTKDED-----SSEASFELDKARLK 550
Query: 1567 EMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGP 1746
E+R+GIDLATTLERIEKNFVITDPRLPDNPIIFASD+FL LTEY REEILGRNCRFLQGP
Sbjct: 551 EIRRGIDLATTLERIEKNFVITDPRLPDNPIIFASDNFLELTEYSREEILGRNCRFLQGP 610
Query: 1747 ETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDG 1926
+T+R TVK IRDAIDN+ EVTVQL+NYTK+G+ FWNLFHLQPMRD KG++QYF GVQLDG
Sbjct: 611 DTNRETVKLIRDAIDNEKEVTVQLLNYTKTGRTFWNLFHLQPMRDHKGELQYFTGVQLDG 670
Query: 1927 TE-------RVREAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPH 2085
TE R+ + A +GA ++++TA N++EA +ELPDANL+ EDLW HS+ VLPKPH
Sbjct: 671 TEYLEPLTKRLSQQIASEGAKIIRETAANVNEALRELPDANLKVEDLWRIHSRLVLPKPH 730
Query: 2086 MKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSV 2265
+ SW I+K+ +GE + LKHFRP+RPLG GDTGSVHLVEL GTG+ FAMKAM+K+V
Sbjct: 731 KLNHDSWGVIRKIHASGEKVKLKHFRPLRPLGYGDTGSVHLVELRGTGKLFAMKAMEKNV 790
Query: 2266 MLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMK 2445
M+ RNKVHR AER+IL M+DHPFLPTLYASF+T+TH+CLI D+CAGGELF+LL+RQP K
Sbjct: 791 MVKRNKVHRVCAEREILGMMDHPFLPTLYASFETQTHVCLITDFCAGGELFLLLERQPTK 850
Query: 2446 VLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQ 2625
+ +E+ RFY +EVV ALEYLHCQG+IYRDLKPENILL +DGH+ L+DFDLS L+S P+
Sbjct: 851 IFREETARFYTSEVVVALEYLHCQGVIYRDLKPENILLQQDGHVMLSDFDLSYLSSSNPR 910
Query: 2626 VFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALG 2805
+ +P + PIF AEP+ A NSFVGTEEYIAPE+ITG+GH S+VDWWALG
Sbjct: 911 LVVPPRLHKKRSKRKNFPPPIFRAEPIGACNSFVGTEEYIAPEVITGSGHNSSVDWWALG 970
Query: 2806 ILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGS 2985
IL+YEMLYG TPFRGKTRQ+TF NILHKD+ FP I SLAARQLI LL +DP NRLGS
Sbjct: 971 ILMYEMLYGRTPFRGKTRQKTFGNILHKDLVFPRRIPTSLAARQLINGLLQKDPENRLGS 1030
Query: 2986 YEGAMEIKQHPFFRGINWALVRAATPPELEAPL 3084
GA EIK HPFF+G+NW L+R PP +AP+
Sbjct: 1031 QGGANEIKGHPFFQGVNWTLIRCMRPPTFDAPV 1063
>sptr|Q9ST27|Q9ST27 Nonphototrophic hypocotyl 1b.
Length = 907
Score = 975 bits (2520), Expect = 0.0
Identities = 506/834 (60%), Positives = 596/834 (71%), Gaps = 23/834 (2%)
Frame = +1
Query: 694 SGNSIDEXXXXXXXXXXLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTG 873
SG S + LPRVS+EL+ ALS+ QQTFVVSDATRPD PI+YAS GFF MTG
Sbjct: 69 SGKSSVDGGVGRASHDSLPRVSQELKDALSSLQQTFVVSDATRPDCPIIYASEGFFTMTG 128
Query: 874 YSSNEVVGRNCRFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPI 1053
YS EVVGRNCRFLQG TD E++KIR A+ G ++CGR+LNY+KDG PFWNLLTV PI
Sbjct: 129 YSPREVVGRNCRFLQGPDTDAAEVAKIRDAVKHGRSFCGRLLNYRKDGAPFWNLLTVTPI 188
Query: 1054 KDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLAL 1233
+D++G+V+KFIGMQVEVSKYTEG D +RPN LP SLI+YD RQKD A SS++E++ +
Sbjct: 189 RDDNGKVIKFIGMQVEVSKYTEGLSDKRMRPNELPVSLIRYDERQKDKAMSSMTEVVQTV 248
Query: 1234 KDPRSLSESRNNTLKRKSQESAGSALV----------PGKRSSETGSRRNSHSGMRNSLQ 1383
K PR + L + S + P GS ++ ++
Sbjct: 249 KQPRGARAPADAALLTPPKMSDADKMAAMSPVVAPGTPSGGGGGAGSFKSPLWDLKKEES 308
Query: 1384 KISEVPEGGNKTRKSGLRSFMGL-IG----MGHGNVEKNILKPRE-DPLLXXXXXXXXXX 1545
++S + G RKSG S MG IG +G + +P P
Sbjct: 309 RLSRLASG----RKSGRSSLMGFKIGKRSSVGSREAPAVVEEPAPAPPPAPEVVERTDSW 364
Query: 1546 XXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRN 1725
+K++R+GIDLATTLERIEKNFVITDPR+PDNPIIFASDSFL LTEY REEILGRN
Sbjct: 365 ERAEREKDIRQGIDLATTLERIEKNFVITDPRIPDNPIIFASDSFLELTEYTREEILGRN 424
Query: 1726 CRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYF 1905
CRFLQGPETD+GTV KIR+AI Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYF
Sbjct: 425 CRFLQGPETDQGTVDKIREAIREQKEITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYF 484
Query: 1906 IGVQLDGTE-------RVREAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSK 2064
IGVQLDG++ R+ E A LVK TA+N+D+A +ELPDANLRPEDLWA HS
Sbjct: 485 IGVQLDGSDHVEPLRNRLSENTEIQSAKLVKATAENVDDAVRELPDANLRPEDLWAIHSM 544
Query: 2065 PVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAM 2244
V PKPH ++ SW AI+K GE I LKHF+PV+PLG GDTGSVHLVEL G+GE FAM
Sbjct: 545 RVSPKPHKRNNPSWIAIEKATNLGEKIGLKHFKPVKPLGCGDTGSVHLVELQGSGELFAM 604
Query: 2245 KAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFML 2424
KAMDKSVMLNRNKVHRA ER+I +LDHPFLPTLY SFQT TH+CLI D+C GGELF +
Sbjct: 605 KAMDKSVMLNRNKVHRACIEREIYALLDHPFLPTLYTSFQTPTHVCLITDFCPGGELFAV 664
Query: 2425 LDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSC 2604
LDRQPMK+ +E+ RFYAAEVV LEYLHC GIIYRDLKPENILL DGHI LTDFDLS
Sbjct: 665 LDRQPMKIFREECARFYAAEVVIGLEYLHCLGIIYRDLKPENILLQADGHIVLTDFDLSF 724
Query: 2605 LTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSA 2784
LT+ +P V + + P F +EP SNSFVGTEEYIAPE+ITGAGHTSA
Sbjct: 725 LTTSKPHV-IKNSTSLKRRRSQEFLPPTFVSEPSTPSNSFVGTEEYIAPEVITGAGHTSA 783
Query: 2785 VDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRD 2964
+DWWALGILLYEMLYG TPFRGK R++TF NILHKD+ FP+SI VSLAA+QLI+ LL RD
Sbjct: 784 IDWWALGILLYEMLYGRTPFRGKNRKKTFYNILHKDLTFPSSIPVSLAAKQLIHGLLQRD 843
Query: 2965 PANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDTLEATLPPPAAD 3126
P+NR+GS GA +IKQH FF+ INW L+R +PPEL+ PL+ + T P D
Sbjct: 844 PSNRIGSNAGANDIKQHSFFQDINWPLIRCMSPPELDVPLKLIGKETQPKAKPD 897
>sptr|Q7XVR2|Q7XVR2 OSJNBa0079M09.14 protein.
Length = 940
Score = 958 bits (2477), Expect = 0.0
Identities = 506/867 (58%), Positives = 596/867 (68%), Gaps = 56/867 (6%)
Frame = +1
Query: 694 SGNSIDEXXXXXXXXXXLPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTG 873
SG S + LPRVS+EL+ ALS+ QQTFVVSDATRPD PI+YAS GFF MTG
Sbjct: 69 SGKSSVDGGVGRASHDSLPRVSQELKDALSSLQQTFVVSDATRPDCPIIYASEGFFTMTG 128
Query: 874 YSSNEVVGRNCRFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPI 1053
YS EVVGRNCRFLQG TD E++KIR A+ G ++CGR+LNY+KDG PFWNLLTV PI
Sbjct: 129 YSPREVVGRNCRFLQGPDTDAAEVAKIRDAVKHGRSFCGRLLNYRKDGAPFWNLLTVTPI 188
Query: 1054 KDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDA----------------- 1182
+D++G+V+KFIGMQVEVSKYTEG D +RPN LP SLI+YD
Sbjct: 189 RDDNGKVIKFIGMQVEVSKYTEGLSDKRMRPNELPVSLIRYDGEHRRRRRGILCFLDVSL 248
Query: 1183 ----------------RQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALV 1314
RQKD A SS++E++ +K PR + L + S +
Sbjct: 249 ILLSYQCRHWRVWSAERQKDKAMSSMTEVVQTVKQPRGARAPADAALLTPPKMSDADKMA 308
Query: 1315 ----------PGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGL-IG- 1458
P GS ++ ++ ++S + G RKSG S MG IG
Sbjct: 309 AMSPVVAPGTPSGGGGGAGSFKSPLWDLKKEESRLSRLASG----RKSGRSSLMGFKIGK 364
Query: 1459 ---MGHGNVEKNILKPRE-DPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFV 1626
+G + +P P +K++R+GIDLATTLERIEKNFV
Sbjct: 365 RSSVGSREAPAVVEEPAPAPPPAPEVVERTDSWERAEREKDIRQGIDLATTLERIEKNFV 424
Query: 1627 ITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEV 1806
ITDPR+PDNPIIFASDSFL LTEY REEILGRNCRFLQGPETD+GTV KIR+AI Q E+
Sbjct: 425 ITDPRIPDNPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQGTVDKIREAIREQKEI 484
Query: 1807 TVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE-------RVREAAAKDGA 1965
TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++ R+ E A
Sbjct: 485 TVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHVEPLRNRLSENTEIQSA 544
Query: 1966 ILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESI 2145
LVK TA+N+D+A +ELPDANLRPEDLWA HS V PKPH ++ SW AI+K GE I
Sbjct: 545 KLVKATAENVDDAVRELPDANLRPEDLWAIHSMRVSPKPHKRNNPSWIAIEKATNLGEKI 604
Query: 2146 DLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDML 2325
LKHF+PV+PLG GDTGSVHLVEL G+GE FAMKAMDKSVMLNRNKVHRA ER+I +L
Sbjct: 605 GLKHFKPVKPLGCGDTGSVHLVELQGSGELFAMKAMDKSVMLNRNKVHRACIEREIYALL 664
Query: 2326 DHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEY 2505
DHPFLPTLY SFQT TH+CLI D+C GGELF +LDRQPMK+ +E+ RFYAAEVV LEY
Sbjct: 665 DHPFLPTLYTSFQTPTHVCLITDFCPGGELFAVLDRQPMKIFREECARFYAAEVVIGLEY 724
Query: 2506 LHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNP 2685
LHC GIIYRDLKPENILL DGHI LTDFDLS LT+ +P V + + P
Sbjct: 725 LHCLGIIYRDLKPENILLQADGHIVLTDFDLSFLTTSKPHV-IKNSTSLKRRRSQEFLPP 783
Query: 2686 IFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQR 2865
F +EP SNSFVGTEEYIAPE+ITGAGHTSA+DWWALGILLYEMLYG TPFRGK R++
Sbjct: 784 TFVSEPSTPSNSFVGTEEYIAPEVITGAGHTSAIDWWALGILLYEMLYGRTPFRGKNRKK 843
Query: 2866 TFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWAL 3045
TF NILHKD+ FP+SI VSLAA+QLI+ LL RDP+NR+GS GA +IKQH FF+ INW L
Sbjct: 844 TFYNILHKDLTFPSSIPVSLAAKQLIHGLLQRDPSNRIGSNAGANDIKQHSFFQDINWPL 903
Query: 3046 VRAATPPELEAPLQDTLEATLPPPAAD 3126
+R +PPEL+ PL+ + T P D
Sbjct: 904 IRCMSPPELDVPLKLIGKETQPKAKPD 930
>sptr|Q9ZWQ6|Q9ZWQ6 PHY3.
Length = 1465
Score = 842 bits (2174), Expect = 0.0
Identities = 445/782 (56%), Positives = 549/782 (70%), Gaps = 24/782 (3%)
Frame = +1
Query: 778 LSAFQQ-TFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKI 954
+SAFQ +F+V DA +PD PI+YAS GFFN+TGY+S EV+G NCRFLQG T+P +++ I
Sbjct: 669 ISAFQHNSFIVVDALKPDFPIIYASTGFFNLTGYTSREVIGGNCRFLQGPDTNPADVASI 728
Query: 955 RQALAAGSN-YCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKD 1131
R+ALA G+ +CGR+LNY+KDG+ FWNLLT+APIKD+ G ++K IG+Q+EVSKYTEG +
Sbjct: 729 REALAQGTGTFCGRLLNYRKDGSSFWNLLTIAPIKDDLGSIVKLIGVQLEVSKYTEGIRA 788
Query: 1132 TALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSAL 1311
RPNG+P+SLI+YD R +D + +++L+ AL P + R L
Sbjct: 789 NNRRPNGMPQSLIRYDVRHQDKVSAFIAQLVAALTKPDKVETPR---------------L 833
Query: 1312 VPGKRSSETGSRRNSHSGMRNSLQKISEVP-EGGNKTRKSGLRSFMGLIGMGHGNVEKNI 1488
R S TG S L + + +P EGG +TR+ SF+ L+GM EK+I
Sbjct: 834 SSAMRFSLTGQTIES-------LPQPTAIPREGGGRTRRPRSSSFLSLLGM---EKEKDI 883
Query: 1489 LKPREDPL-----LXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDN 1653
P ED L + + R+GIDLATTLERI K+FVITDPRLPDN
Sbjct: 884 --PEEDELQELEVIMLEDASVGRPGSLDDPERTRRGIDLATTLERIGKSFVITDPRLPDN 941
Query: 1654 PIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTK 1833
PIIFASD FL LTEY REE+LG NCRFLQG TDR V+ IRDA+ Q +VTVQ++NYTK
Sbjct: 942 PIIFASDRFLELTEYTREEVLGNNCRFLQGRGTDRKAVQLIRDAVKEQRDVTVQVLNYTK 1001
Query: 1834 SGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGT------------ERVREAAAKDGAILVK 1977
G+ FWNLFHLQ MRD+ GDVQYFIGVQ + + + + ++ A +V+
Sbjct: 1002 GGRAFWNLFHLQVMRDENGDVQYFIGVQQEMVAPRPVHQPPELPDILPDRVEQEKAEVVR 1061
Query: 1978 KTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMK-DTASWRAIQKV---LENGESI 2145
TA +D AA+ELPDANL P+ L+A HSK V P PH K +++SW AI++V L GE +
Sbjct: 1062 ATAQRVDAAARELPDANLVPDHLFAPHSKVVTPLPHSKTNSSSWFAIRRVQRRLRRGERL 1121
Query: 2146 DLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDML 2325
LKHFRP++PLGSGDTGSVHLVEL GTG+ FA+KAMDKS+ML RNKVHRA AER+IL ++
Sbjct: 1122 GLKHFRPIKPLGSGDTGSVHLVELRGTGQVFALKAMDKSMMLQRNKVHRARAEREILAIM 1181
Query: 2326 DHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEY 2505
DHPFLPTLYASFQTKTH+CLI DYC GG+LF+L D+QP + L E FYAAEVV ALEY
Sbjct: 1182 DHPFLPTLYASFQTKTHVCLITDYCPGGDLFLLQDKQPTQTLSERTASFYAAEVVVALEY 1241
Query: 2506 LHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNP 2685
LHC G+IYRDLKPEN+LL ++GHI LTDFDLS LTSCRPQ+ L
Sbjct: 1242 LHCMGVIYRDLKPENVLLQKNGHILLTDFDLSFLTSCRPQLILQGG-KGRSRRSKRRRRV 1300
Query: 2686 IFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQR 2865
F AEP +SNSFVGTEEYIAPEII+G H+SAVDWWALGILLYEMLYG TPF G+ RQ+
Sbjct: 1301 TFCAEPRVSSNSFVGTEEYIAPEIISGEPHSSAVDWWALGILLYEMLYGRTPFVGRNRQK 1360
Query: 2866 TFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWAL 3045
TF N+L+K++ FP SI VSLA RQLI LL RDP RLG+ GA E+K+HPFFR INW L
Sbjct: 1361 TFYNVLNKELIFPTSIPVSLAGRQLIAGLLQRDPTIRLGTLRGASELKKHPFFREINWPL 1420
Query: 3046 VR 3051
+R
Sbjct: 1421 IR 1422
Score = 99.4 bits (246), Expect = 3e-19
Identities = 48/114 (42%), Positives = 70/114 (61%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L ++FV++D PD+PI++AS F +T Y+ EV+G NCRFLQG GTD
Sbjct: 919 DLATTLERIGKSFVITDPRLPDNPIIFASDRFLELTEYTREEVLGNNCRFLQGRGTDRKA 978
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEV 1104
+ IR A+ + ++LNY K G FWNL + ++DE+G V FIG+Q E+
Sbjct: 979 VQLIRDAVKEQRDVTVQVLNYTKGGRAFWNLFHLQVMRDENGDVQYFIGVQQEM 1032
>sptr|Q41384|Q41384 Protein kinase (Fragment).
Length = 724
Score = 834 bits (2155), Expect = 0.0
Identities = 427/701 (60%), Positives = 517/701 (73%), Gaps = 17/701 (2%)
Frame = +1
Query: 1036 LTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVS 1215
LTV PI+D+ G V+KFIGMQVEVSK+TEG D ALRPNGLP+SLI+YD RQK+ A S+
Sbjct: 1 LTVTPIRDDKGCVIKFIGMQVEVSKFTEGINDKALRPNGLPKSLIRYDPRQKEAALGSII 60
Query: 1216 ELLLALKDPRSLSESRNNTLKRKSQESAGSA----LVPGKRSSE---TGSRRNSHSGMRN 1374
E++ +K PRSLS+ +N +E AG ++P + T R +S S +
Sbjct: 61 EVVQTVKHPRSLSQPLSNN-DADRREVAGKFNLDYMLPKLAEIDNVGTPGRLSSPSRLST 119
Query: 1375 SLQKISEVPEGGNKTRKSGLRSF-MGLIGMGHGNVEKNILKPREDP--LLXXXXXXXXXX 1545
++ ++ + KS +S + L+G + K+ P +P L+
Sbjct: 120 PGRQTPKIDASSRDSDKSSRKSARISLLGFKGRSSAKHERPPSPEPEFLIPKDIDRDDSW 179
Query: 1546 XXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRN 1725
++++R+GIDLATTLERIEKNFVI+DPRLPDNPIIFASDSFL LTEY REEILGRN
Sbjct: 180 ERAERERDVRQGIDLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYTREEILGRN 239
Query: 1726 CRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYF 1905
CRFLQGPETD+ TV+KIRDAI Q ++TVQLINYTKSG+KFWNLFHLQPMRDQKG++QYF
Sbjct: 240 CRFLQGPETDQTTVQKIRDAIKEQRDITVQLINYTKSGRKFWNLFHLQPMRDQKGELQYF 299
Query: 1906 IGVQLDGTE-------RVREAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSK 2064
IGVQLDG++ R+ E A +VK TA+N+DEA +ELPDAN RPEDLWA HS+
Sbjct: 300 IGVQLDGSDHVEPLRNRLSERTEIQSAKVVKATAENVDEAVRELPDANSRPEDLWAIHSE 359
Query: 2065 PVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAM 2244
PV P+PH + ++SW AIQK+ GE + L+HF P++PLG GDTGSVHLVEL +FAM
Sbjct: 360 PVYPRPHKRGSSSWAAIQKITAAGEKVGLEHFNPIKPLGCGDTGSVHLVELKVPENWFAM 419
Query: 2245 KAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFML 2424
KAMDKSVMLNRNKVHRA ER+I+ LDHPFLPTLYASFQT TH+CLI D+C GGELF L
Sbjct: 420 KAMDKSVMLNRNKVHRACVEREIISTLDHPFLPTLYASFQTSTHVCLITDFCPGGELFAL 479
Query: 2425 LDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSC 2604
LD+QP+K+ KE++ RFYAAEVV LEYLHC GIIYRDLKPENILL +DGH+ LTDFDLS
Sbjct: 480 LDKQPLKIFKEESARFYAAEVVIGLEYLHCLGIIYRDLKPENILLQKDGHLVLTDFDLSF 539
Query: 2605 LTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSA 2784
LTSC P + + P F AEP+ SNSFVGTEEYIAPE+ITGA HTSA
Sbjct: 540 LTSCNPHII--NHPQPKKRRSRSQPPPTFIAEPVTQSNSFVGTEEYIAPEVITGASHTSA 597
Query: 2785 VDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRD 2964
+DWWALG+LLYEMLYG TPFRGK RQ+TFANI+HKD+ FP+SI VSL+ARQLIY LL+RD
Sbjct: 598 IDWWALGVLLYEMLYGRTPFRGKNRQKTFANIMHKDLTFPSSIPVSLSARQLIYALLNRD 657
Query: 2965 PANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQ 3087
PA RLG+ GA EIK+HP+FRGINW L+R PP LEAP +
Sbjct: 658 PATRLGTQGGASEIKEHPYFRGINWPLIRCMDPPTLEAPFK 698
Score = 108 bits (270), Expect = 5e-22
Identities = 58/165 (35%), Positives = 88/165 (53%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L ++ FV+SD PD+PI++AS F +T Y+ E++GRNCRFLQG TD
Sbjct: 193 DLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQTT 252
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEG 1122
+ KIR A+ + +++NY K G FWNL + P++D+ G + FIG+Q++ S + E
Sbjct: 253 VQKIRDAIKEQRDITVQLINYTKSGRKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHVEP 312
Query: 1123 NKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSE 1257
+ N L E A+ +V E + L D S E
Sbjct: 313 LR------NRLSERTEIQSAKVVKATAENVDEAVRELPDANSRPE 351
>sptr|Q40269|Q40269 Protein kinase (Fragment).
Length = 572
Score = 798 bits (2061), Expect = 0.0
Identities = 393/515 (76%), Positives = 436/515 (84%), Gaps = 8/515 (1%)
Frame = +1
Query: 1561 KKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQ 1740
KKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFL LTE+ R EIL RN RFLQ
Sbjct: 33 KKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEFSRAEILARNRRFLQ 92
Query: 1741 GPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQL 1920
GPETD TV KIRDAIDN+T+VTVQLINYTK+GKKFWN+FHLQPMRDQKG+VQYFIGVQL
Sbjct: 93 GPETDPATVAKIRDAIDNETDVTVQLINYTKTGKKFWNVFHLQPMRDQKGEVQYFIGVQL 152
Query: 1921 DGTERVRE-------AAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPK 2079
DG+E V A+ D VK+TA N+D A +ELPDAN +PEDLWANHSK V PK
Sbjct: 153 DGSEHVEPVQNSIPVASVMDSEKQVKETATNVDVAVRELPDANKKPEDLWANHSKVVQPK 212
Query: 2080 PHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDK 2259
PH K+ +SW+AI+K+ E+GE I LKHFRPV+PLG+GDTGSVHLVEL GTGEYFAMKAMDK
Sbjct: 213 PHRKECSSWKAIEKIKESGEQIGLKHFRPVKPLGAGDTGSVHLVELCGTGEYFAMKAMDK 272
Query: 2260 SVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQP 2439
+VMLNRNKVHRA AER+ILDMLDHPFLP LYASFQT THICLI +YC GGELF+LLDRQP
Sbjct: 273 NVMLNRNKVHRACAEREILDMLDHPFLPALYASFQTNTHICLITEYCPGGELFLLLDRQP 332
Query: 2440 MKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCR 2619
KVLKEDAVRFYAAEV+ ALEYLHCQGIIYRDLKPENILL +GH+SLTDFDLSCLTSC+
Sbjct: 333 TKVLKEDAVRFYAAEVIIALEYLHCQGIIYRDLKPENILLQSNGHVSLTDFDLSCLTSCK 392
Query: 2620 PQVFLPHDID-XXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWW 2796
PQ+ +P D PIF AEPMRASNSFVGTEEYIAPEII GAG AVDWW
Sbjct: 393 PQLLIPEIRDKKKQQKAQHQQTPIFMAEPMRASNSFVGTEEYIAPEIIAGAGIQGAVDWW 452
Query: 2797 ALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANR 2976
ALGILLYEMLYG+TPFRGKTRQ+TF+N+L KD++FPA+ QVSL A QLIY+LL +DP +R
Sbjct: 453 ALGILLYEMLYGFTPFRGKTRQKTFSNVLRKDLKFPATKQVSLDASQLIYQLLQKDPKDR 512
Query: 2977 LGSYEGAMEIKQHPFFRGINWALVRAATPPELEAP 3081
LG+ EGA EIK+HPFFRG NWALVR PP L+AP
Sbjct: 513 LGACEGANEIKRHPFFRGANWALVRCMKPPVLDAP 547
Score = 105 bits (263), Expect = 3e-21
Identities = 62/199 (31%), Positives = 110/199 (55%), Gaps = 4/199 (2%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L ++ FV++D PD+PI++AS F +T +S E++ RN RFLQG TDP
Sbjct: 41 DLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEFSRAEILARNRRFLQGPETDPAT 100
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEG 1122
++KIR A+ ++ +++NY K G FWN+ + P++D+ G V FIG+Q++ S++ E
Sbjct: 101 VAKIRDAIDNETDVTVQLINYTKTGKKFWNVFHLQPMRDQKGEVQYFIGVQLDGSEHVEP 160
Query: 1123 NKDTALRPNGLP-ESLIKYDARQKDHARS---SVSELLLALKDPRSLSESRNNTLKRKSQ 1290
+ N +P S++ + + K+ A + +V EL A K P L + + ++ K
Sbjct: 161 VQ------NSIPVASVMDSEKQVKETATNVDVAVRELPDANKKPEDLWANHSKVVQPKPH 214
Query: 1291 ESAGSALVPGKRSSETGSR 1347
S+ ++ E+G +
Sbjct: 215 RKECSSWKAIEKIKESGEQ 233
>sptr|Q07526|Q07526 Protein kinase PSPK-5 (EC 2.7.1.-) (Probable
serine/threonine-protein kinase PK5).
Length = 428
Score = 634 bits (1635), Expect = e-180
Identities = 309/414 (74%), Positives = 344/414 (83%), Gaps = 7/414 (1%)
Frame = +1
Query: 1873 MRDQKGDVQYFIGVQLDGTE-------RVREAAAKDGAILVKKTADNIDEAAKELPDANL 2031
MRDQKG+VQYFIGVQLDG++ R+ E AK+G LVKKTA+N+D+A +ELPDAN+
Sbjct: 1 MRDQKGEVQYFIGVQLDGSQHVEPLHNRIAEDTAKEGENLVKKTAENVDDALRELPDANM 60
Query: 2032 RPEDLWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLV 2211
+PEDLW NHSK V PKPH ++ A+WRAIQK++E+GE I LKHF+P++PLGSGDTGSVHLV
Sbjct: 61 KPEDLWMNHSKMVHPKPHRREDAAWRAIQKIMESGEQIGLKHFKPIKPLGSGDTGSVHLV 120
Query: 2212 ELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIV 2391
EL GT FAMKAMDK V+LNRNK HRA ER+ILDMLDHPFLP LYASFQTKTHICLI
Sbjct: 121 ELCGTDHQFAMKAMDKGVILNRNKEHRACTEREILDMLDHPFLPALYASFQTKTHICLIT 180
Query: 2392 DYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDG 2571
DYC GGELFMLLDRQP KVLKEDAVRFYA EVV ALEYLHCQGIIYRDLKPEN+LL G
Sbjct: 181 DYCPGGELFMLLDRQPAKVLKEDAVRFYATEVVVALEYLHCQGIIYRDLKPENVLLQSTG 240
Query: 2572 HISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAP 2751
H+SLTDFDLSCLTSC+PQ+ +P D PIF AEPMRASNSFVGTEEYIAP
Sbjct: 241 HVSLTDFDLSCLTSCKPQLLVPSTND----KKKGQHGPIFMAEPMRASNSFVGTEEYIAP 296
Query: 2752 EIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAA 2931
EIITG+GHTSAVDWWALGILLYEM YGYTPFRGK RQRTFANILHKD++FP S QVSL A
Sbjct: 297 EIITGSGHTSAVDWWALGILLYEMFYGYTPFRGKNRQRTFANILHKDLKFPKSKQVSLGA 356
Query: 2932 RQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPPELEAPLQDT 3093
+QLIY LL RDP +RLGS GA +IK H FF+GINWALVR PPEL+APL DT
Sbjct: 357 KQLIYYLLQRDPTSRLGSKGGANDIKNHSFFKGINWALVRCTKPPELDAPLFDT 410
>sptr|Q8RWE6|Q8RWE6 Hypothetical protein (At5g58140).
Length = 549
Score = 490 bits (1262), Expect = e-137
Identities = 274/551 (49%), Positives = 340/551 (61%), Gaps = 23/551 (4%)
Frame = +1
Query: 445 RGSLEVFNPDAPVSDRATTSPFLLPPAVASHPSLLXXXXXXXXXVGRATQRAAEWGLVLQ 624
R SLE+FNP + +TS PP ++ + T+R AEWGL
Sbjct: 20 RRSLEIFNPSSGKETHGSTSSSSKPPLDGNNKGSSSKWMEFQDSA-KITERTAEWGLSAV 78
Query: 625 TDEHTGRPQGVVARPSGSNRTSESGNSIDEXXXXXXXXXXLPRVSEELRAALSAFQQTFV 804
+ G+ + S S++ + PRVS+EL+ ALS QQTFV
Sbjct: 79 KPD--SGDDGISFKLSSEVERSKNMSRRSSEESTSSESGAFPRVSQELKTALSTLQQTFV 136
Query: 805 VSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQALAAGSNY 984
VSDAT+P PI+YAS+GFF MTGYSS E+VGRNCRFLQG TD E++KIR + G +Y
Sbjct: 137 VSDATQPHCPIVYASSGFFTMTGYSSKEIVGRNCRFLQGPDTDKNEVAKIRDCVKNGKSY 196
Query: 985 CGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTALRPNGLPES 1164
CGR+LNYKKDGTPFWNLLTV PIKD+ G +KFIGMQVEVSKYTEG D ALRPNGL +S
Sbjct: 197 CGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFIGMQVEVSKYTEGVNDKALRPNGLSKS 256
Query: 1165 LIKYDARQKDHARSSVSELLLALKDPRS-LSESRNNTLK---------------RKSQES 1296
LI+YDARQK+ A S++E++ ++ +S + ES +N R+S E+
Sbjct: 257 LIRYDARQKEKALDSITEVVQTIRHRKSQVQESVSNDTMVKPDSSTTPTPGRQTRQSDEA 316
Query: 1297 AGSALVPGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNV 1476
+ S PG+ S+ TGS+ S + L +
Sbjct: 317 SKSFRTPGRVSTPTGSKLKSSNNRHEDLLR------------------------------ 346
Query: 1477 EKNILKPREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNP 1656
++P E L ++++R+GIDLATTLERIEKNFVI+DPRLPDNP
Sbjct: 347 ----MEPEELMLSTEVIGQRDSWDLSDRERDIRQGIDLATTLERIEKNFVISDPRLPDNP 402
Query: 1657 IIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKS 1836
IIFASDSFL LTEY REEILGRNCRFLQGPETD+ TV+KIRDAI +Q E+TVQLINYTKS
Sbjct: 403 IIFASDSFLELTEYSREEILGRNCRFLQGPETDQATVQKIRDAIRDQREITVQLINYTKS 462
Query: 1837 GKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE-------RVREAAAKDGAILVKKTADNI 1995
GKKFWNLFHLQPMRDQKG++QYFIGVQLDG++ R+ E + LVK TA N+
Sbjct: 463 GKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHVEPLQNRLSERTEMQSSKLVKATATNV 522
Query: 1996 DEAAKELPDAN 2028
DEA +ELPDAN
Sbjct: 523 DEAVRELPDAN 533
>sptr|Q8LPD9|Q8LPD9 Putative blue light receptor.
Length = 749
Score = 479 bits (1233), Expect = e-133
Identities = 260/531 (48%), Positives = 334/531 (62%), Gaps = 33/531 (6%)
Frame = +1
Query: 1573 RKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPET 1752
R +DLATT+ERI++NF I+DP LPD PI+FASD+FL LT Y REE+LGRNCRFLQG T
Sbjct: 199 RVALDLATTVERIQQNFCISDPTLPDCPIVFASDAFLELTGYSREEVLGRNCRFLQGAGT 258
Query: 1753 DRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE 1932
DRGTV +IR AI +E+TV+++NYTK+GK FWN+F L PMRDQ G ++F+GVQ+D T
Sbjct: 259 DRGTVDQIRAAIKEGSELTVRILNYTKAGKAFWNMFTLAPMRDQDGHARFFVGVQVDVTA 318
Query: 1933 RVREAAAKDGAILVKKTADN-IDEAAKELPDANLRPEDL-----------WANHSKPVLP 2076
++ + D A + KT + + +A A+L L WA S ++
Sbjct: 319 ---QSTSPDKAPVWNKTPEEEVAKAKMGAEAASLISSALQGMAAPTTANPWAAISGVIMR 375
Query: 2077 -KPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAM 2253
KPH D +++A+ ++ E + L HFR V+ LG+GD G V LV+L G+ FAMK +
Sbjct: 376 RKPHKADDKAYQALLQLQERDGKMKLMHFRRVKQLGAGDVGLVDLVQLQGSELKFAMKTL 435
Query: 2254 DKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDR 2433
DK M RNKV R E IL +DHPFL TLY + QT TH+ +++YC GGEL+ LL+
Sbjct: 436 DKFEMQERNKVARVLTESAILAAVDHPFLATLYCTIQTDTHLHFVMEYCDGGELYGLLNS 495
Query: 2434 QPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTS 2613
QP K LKE+ VRFYA+EV+TAL+YLH G +YRDLKPENILLH GH+ LTDFDLS
Sbjct: 496 QPKKRLKEEHVRFYASEVLTALQYLHLLGYVYRDLKPENILLHHTGHVLLTDFDLSYSKG 555
Query: 2614 C--------------------RPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGT 2733
P+ N + AEP +NSFVGT
Sbjct: 556 STTPRIEKIGGAGAAGGSAPKSPKKSSSKSGGSSSGSALQLENYLLLAEPSARANSFVGT 615
Query: 2734 EEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPASI 2913
EEY+APE+I AGH AVDWW+LGIL++E+LYG TPFRG R TF NI+ ++FP+
Sbjct: 616 EEYLAPEVINAAGHGPAVDWWSLGILIFELLYGTTPFRGARRDETFENIIKSPLKFPSKP 675
Query: 2914 QVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPP 3066
VS R LI +LL +D RLGS GA EIK HP+F+GINWAL+R PP
Sbjct: 676 AVSEECRDLIEKLLVKDVGARLGSRTGANEIKSHPWFKGINWALLRHQQPP 726
Score = 172 bits (435), Expect = 4e-41
Identities = 87/165 (52%), Positives = 115/165 (69%), Gaps = 1/165 (0%)
Frame = +1
Query: 745 LPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGS 924
+P + +L L+ + TFVV+DAT PD P++YAS GF+ MTGY +EV+G NCRFLQG
Sbjct: 4 VPAPASQLTKVLAGLRHTFVVADATLPDCPLVYASEGFYAMTGYGPDEVLGHNCRFLQGE 63
Query: 925 GTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEV 1104
GTDP E+ KIR A+ G R+LNY+KDGTPFWNLLTV PIK DGRV KF+G+QV+V
Sbjct: 64 GTDPKEVQKIRDAIKKGEACSVRLLNYRKDGTPFWNLLTVTPIKTPDGRVSKFVGVQVDV 123
Query: 1105 SKYTEGNKDTALRPNGLPESLIKYDARQKDH-ARSSVSELLLALK 1236
+ TEG AL N L+KYD R +D+ AR+ V ++ +A++
Sbjct: 124 TSKTEGK---ALADNSGVPLLVKYDHRLRDNVARTIVDDVTIAVE 165
Score = 133 bits (335), Expect = 2e-29
Identities = 72/165 (43%), Positives = 99/165 (60%)
Frame = +1
Query: 748 PRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSG 927
PRV+ +L + QQ F +SD T PD PI++AS F +TGYS EV+GRNCRFLQG+G
Sbjct: 198 PRVALDLATTVERIQQNFCISDPTLPDCPIVFASDAFLELTGYSREEVLGRNCRFLQGAG 257
Query: 928 TDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVS 1107
TD + +IR A+ GS RILNY K G FWN+ T+AP++D+DG F+G+QV+V+
Sbjct: 258 TDRGTVDQIRAAIKEGSELTVRILNYTKAGKAFWNMFTLAPMRDQDGHARFFVGVQVDVT 317
Query: 1108 KYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDP 1242
+ + D A N PE + A+ A S +S L + P
Sbjct: 318 AQST-SPDKAPVWNKTPEEEVA-KAKMGAEAASLISSALQGMAAP 360
Score = 112 bits (280), Expect = 4e-23
Identities = 60/148 (40%), Positives = 83/148 (56%), Gaps = 3/148 (2%)
Frame = +1
Query: 1588 LATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTV 1767
L L + FV+ D LPD P+++AS+ F +T Y +E+LG NCRFLQG TD V
Sbjct: 11 LTKVLAGLRHTFVVADATLPDCPLVYASEGFYAMTGYGPDEVLGHNCRFLQGEGTDPKEV 70
Query: 1768 KKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVREA 1947
+KIRDAI +V+L+NY K G FWNL + P++ G V F+GVQ+D T +
Sbjct: 71 QKIRDAIKKGEACSVRLLNYRKDGTPFWNLLTVTPIKTPDGRVSKFVGVQVDVTSKTEGK 130
Query: 1948 AAKDGA---ILVKKTADNIDEAAKELPD 2022
A D + +LVK D A+ + D
Sbjct: 131 ALADNSGVPLLVKYDHRLRDNVARTIVD 158
>sptr|Q8LPE0|Q8LPE0 Putative blue light receptor.
Length = 750
Score = 476 bits (1225), Expect = e-132
Identities = 260/532 (48%), Positives = 335/532 (62%), Gaps = 34/532 (6%)
Frame = +1
Query: 1573 RKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPET 1752
R +DLATT+ERI++NF I+DP LPD PI+FASD+FL LT Y REE+LGRNCRFLQG T
Sbjct: 199 RVALDLATTVERIQQNFCISDPTLPDCPIVFASDAFLELTGYSREEVLGRNCRFLQGAGT 258
Query: 1753 DRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE 1932
DRGTV +IR AI +E+TV+++NYTK+GK FWN+F L PMRDQ G ++F+GVQ+D T
Sbjct: 259 DRGTVDQIRAAIKEGSELTVRILNYTKAGKAFWNMFTLAPMRDQDGHARFFVGVQVDVTA 318
Query: 1933 RVREAAAKDGAILVKKTADN-IDEAAKELPDANLRPEDL-----------WANHSKPVLP 2076
++ + D A + KT + + +A A+L L WA S ++
Sbjct: 319 ---QSTSPDKAPVWNKTPEEEVAKAKMGAEAASLISSALQGMAAPTTANPWAAISGVIMR 375
Query: 2077 -KPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAM 2253
KPH D +++A+ ++ E + L HFR V+ LG+GD G V LV+L G+ FAMK +
Sbjct: 376 RKPHKADDKAYQALLQLQERDGKMKLMHFRRVKQLGAGDVGLVDLVQLQGSELKFAMKTL 435
Query: 2254 DKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDR 2433
DK M RNKV R E IL +DHPFL TLY + QT TH+ +++YC GGEL+ LL+
Sbjct: 436 DKFEMQERNKVARVLTESAILAAVDHPFLATLYCTIQTDTHLHFVMEYCEGGELYGLLNS 495
Query: 2434 QPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTS 2613
QP K LKE+ VRFYA+EV+TAL+YLH G +YRDLKPENILLH GH+ LTDFDLS
Sbjct: 496 QPKKRLKEEHVRFYASEVLTALQYLHLLGYVYRDLKPENILLHHTGHVLLTDFDLSYSKG 555
Query: 2614 C--------------------RPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGT 2733
P+ N + AEP +NSFVGT
Sbjct: 556 STTPRIEKIGGAGAAGGSAPKSPKKSSSKSGGSSSGSALQLENYLLLAEPSARANSFVGT 615
Query: 2734 EEYIAPEIITGAGH-TSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPAS 2910
EEY+APE+I AGH +AVDWW+LGIL++E+LYG TPFRG R TF NI+ ++FP+
Sbjct: 616 EEYLAPEVINAAGHGPAAVDWWSLGILIFELLYGTTPFRGARRDETFENIIKSPLKFPSK 675
Query: 2911 IQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPP 3066
VS R LI +LL +D RLGS GA EIK HP+F+GINWAL+R PP
Sbjct: 676 PAVSEECRDLIEKLLVKDVGARLGSRTGANEIKSHPWFKGINWALLRHQQPP 727
Score = 172 bits (435), Expect = 4e-41
Identities = 87/165 (52%), Positives = 115/165 (69%), Gaps = 1/165 (0%)
Frame = +1
Query: 745 LPRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGS 924
+P + +L L+ + TFVV+DAT PD P++YAS GF+ MTGY +EV+G NCRFLQG
Sbjct: 4 VPAPASQLTKVLAGLRHTFVVADATLPDCPLVYASEGFYAMTGYGPDEVLGHNCRFLQGE 63
Query: 925 GTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEV 1104
GTDP E+ KIR A+ G R+LNY+KDGTPFWNLLTV PIK DGRV KF+G+QV+V
Sbjct: 64 GTDPKEVQKIRDAIKKGEACSVRLLNYRKDGTPFWNLLTVTPIKTPDGRVSKFVGVQVDV 123
Query: 1105 SKYTEGNKDTALRPNGLPESLIKYDARQKDH-ARSSVSELLLALK 1236
+ TEG AL N L+KYD R +D+ AR+ V ++ +A++
Sbjct: 124 TSKTEGK---ALADNSGVPLLVKYDHRLRDNVARTIVDDVTIAVE 165
Score = 133 bits (335), Expect = 2e-29
Identities = 72/165 (43%), Positives = 99/165 (60%)
Frame = +1
Query: 748 PRVSEELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSG 927
PRV+ +L + QQ F +SD T PD PI++AS F +TGYS EV+GRNCRFLQG+G
Sbjct: 198 PRVALDLATTVERIQQNFCISDPTLPDCPIVFASDAFLELTGYSREEVLGRNCRFLQGAG 257
Query: 928 TDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVS 1107
TD + +IR A+ GS RILNY K G FWN+ T+AP++D+DG F+G+QV+V+
Sbjct: 258 TDRGTVDQIRAAIKEGSELTVRILNYTKAGKAFWNMFTLAPMRDQDGHARFFVGVQVDVT 317
Query: 1108 KYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDP 1242
+ + D A N PE + A+ A S +S L + P
Sbjct: 318 AQST-SPDKAPVWNKTPEEEVA-KAKMGAEAASLISSALQGMAAP 360
Score = 112 bits (280), Expect = 4e-23
Identities = 60/148 (40%), Positives = 83/148 (56%), Gaps = 3/148 (2%)
Frame = +1
Query: 1588 LATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRGTV 1767
L L + FV+ D LPD P+++AS+ F +T Y +E+LG NCRFLQG TD V
Sbjct: 11 LTKVLAGLRHTFVVADATLPDCPLVYASEGFYAMTGYGPDEVLGHNCRFLQGEGTDPKEV 70
Query: 1768 KKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTERVREA 1947
+KIRDAI +V+L+NY K G FWNL + P++ G V F+GVQ+D T +
Sbjct: 71 QKIRDAIKKGEACSVRLLNYRKDGTPFWNLLTVTPIKTPDGRVSKFVGVQVDVTSKTEGK 130
Query: 1948 AAKDGA---ILVKKTADNIDEAAKELPD 2022
A D + +LVK D A+ + D
Sbjct: 131 ALADNSGVPLLVKYDHRLRDNVARTIVD 158
>sptr|P93025|P93025 Putative serine/threonine protein kinase
(Fragment).
Length = 356
Score = 473 bits (1217), Expect = e-132
Identities = 228/326 (69%), Positives = 264/326 (80%)
Frame = +1
Query: 2107 RAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKV 2286
+AI+K+ +GE++ L HF+P++PLGSGDTGSVHLVEL GTGE +AMKAM+K++MLNRNK
Sbjct: 1 KAIKKIQASGETVGLHHFKPIKPLGSGDTGSVHLVELKGTGELYAMKAMEKTMMLNRNKA 60
Query: 2287 HRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAV 2466
HRA ER+I+ +LDHPFLPTLYASFQT TH+CLI D+C GGELF LLDRQPMK+L ED+
Sbjct: 61 HRACIEREIISLLDHPFLPTLYASFQTSTHVCLITDFCPGGELFALLDRQPMKILTEDSA 120
Query: 2467 RFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPHDI 2646
RFYAAEVV LEYLHC GI+YRDLKPENILL +DGHI L DF LS +T+C PQ+ +P
Sbjct: 121 RFYAAEVVIGLEYLHCLGIVYRDLKPENILLKKDGHIVLADFYLSFMTTCTPQLIIP-AA 179
Query: 2647 DXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEML 2826
P F AEP SNSFVGTEEYIAPEIITGAGHTSA+DWWALGILLYEML
Sbjct: 180 PSKRRRSKSQPLPTFVAEPSTQSNSFVGTEEYIAPEIITGAGHTSAIDWWALGILLYEML 239
Query: 2827 YGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEI 3006
YG TPFRGK RQ+TFANILHKD+ FP+SI VSL RQLI LL+RDP++RLGS GA EI
Sbjct: 240 YGRTPFRGKNRQKTFANILHKDLTFPSSIPVSLVGRQLINTLLNRDPSSRLGSKGGANEI 299
Query: 3007 KQHPFFRGINWALVRAATPPELEAPL 3084
KQH FFRGINW L+R +PP L+APL
Sbjct: 300 KQHAFFRGINWPLIRGMSPPPLDAPL 325
>sptr|Q84KS5|Q84KS5 Phytochrome 3 (Fragment).
Length = 686
Score = 444 bits (1141), Expect = e-123
Identities = 236/476 (49%), Positives = 314/476 (65%), Gaps = 9/476 (1%)
Frame = +1
Query: 775 ALSAFQQT-FVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISK 951
A+S FQQT FVV DA +PD PI++AS GFFN+TGY+S EV+G NCRFLQG T+P +++
Sbjct: 223 AISVFQQTSFVVVDALKPDLPIIFASTGFFNLTGYTSREVIGGNCRFLQGPDTNPEDVAS 282
Query: 952 IRQALA--AGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGN 1125
IR A+ +CGR+LNY+KDG+ FWNLLT+APIKD+ G ++K +G+Q+EVSKYTEG+
Sbjct: 283 IRDAVVPRGTGTFCGRLLNYRKDGSNFWNLLTIAPIKDDTGTIVKLVGVQLEVSKYTEGS 342
Query: 1126 KDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGS 1305
+ LRPNGLP+SLIKYD R +D + V++++ AL P + R + R S G
Sbjct: 343 RANRLRPNGLPQSLIKYDVRHQDKVSALVAQIVAALTKPYKVEPPRPSYAMRASL--TGQ 400
Query: 1306 ALVPGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKN 1485
+ P +R S S + + + +P G R+ +F+ L+GM + E++
Sbjct: 401 TIQPLSPGRAAAARPYSAS----DVPQTAAIPREGGGRRRHRSSTFLSLLGMEEKDSEED 456
Query: 1486 ILKPREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIF 1665
E L+ ++ R+GIDLATTLERI +FVITDPRLPDNPIIF
Sbjct: 457 QFP--EPELIMMDNASVGRPGSLDDRERTRRGIDLATTLERIGHSFVITDPRLPDNPIIF 514
Query: 1666 ASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKK 1845
ASD FL LTEY REE+LG NCRFLQG +TD V+ IRDA+ ++TVQL+NYT+ G+
Sbjct: 515 ASDQFLELTEYSREEVLGENCRFLQGRDTDLKAVQLIRDAVKEGRDITVQLLNYTRGGRP 574
Query: 1846 FWNLFHLQPMRDQKGDVQYFIGVQ--LDGTERVREAAAKDGAILVKKTADNIDEAAKELP 2019
FWNLFHLQ MRD+KG++QYFIGVQ D +RV + A+ +V+ TA N+D AA+ELP
Sbjct: 575 FWNLFHLQAMRDKKGNLQYFIGVQQETDTLDRVEQEKAE----VVRATAQNVDVAARELP 630
Query: 2020 DANLRPEDLWANHSKPVLPKPHMK-DTASWRAIQKV---LENGESIDLKHFRPVRP 2175
DANL P+ LW HSK V P PH K ++ W AI+KV L GE + LKHFRP++P
Sbjct: 631 DANLTPDHLWERHSKVVTPLPHSKINSPCWYAIRKVQRRLRRGERLGLKHFRPIKP 686
Score = 100 bits (250), Expect = 1e-19
Identities = 57/160 (35%), Positives = 85/160 (53%), Gaps = 13/160 (8%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L +FV++D PD+PI++AS F +T YS EV+G NCRFLQG TD
Sbjct: 488 DLATTLERIGHSFVITDPRLPDNPIIFASDQFLELTEYSREEVLGENCRFLQGRDTDLKA 547
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQ--------- 1095
+ IR A+ G + ++LNY + G PFWNL + ++D+ G + FIG+Q
Sbjct: 548 VQLIRDAVKEGRDITVQLLNYTRGGRPFWNLFHLQAMRDKKGNLQYFIGVQQETDTLDRV 607
Query: 1096 ----VEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHAR 1203
EV + T N D A R LP++ + D + H++
Sbjct: 608 EQEKAEVVRATAQNVDVAARE--LPDANLTPDHLWERHSK 645
>sptr|Q84KS3|Q84KS3 Phytochrome 3 (Fragment).
Length = 692
Score = 441 bits (1135), Expect = e-122
Identities = 237/476 (49%), Positives = 310/476 (65%), Gaps = 9/476 (1%)
Frame = +1
Query: 775 ALSAFQQT-FVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISK 951
A+S FQQT FVV DA + D PI+YAS GFFN+TGY+S EV+G NCRFLQG T+P I
Sbjct: 228 AISVFQQTSFVVVDALKLDLPIIYASTGFFNLTGYTSREVIGGNCRFLQGPETNPAVIDS 287
Query: 952 IRQALA--AGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGN 1125
IR+AL +CGR+LNY+KDG+ FWNLLT+APIKD+ G ++ IG+Q+EVSKYTEG+
Sbjct: 288 IREALVPQGTGTFCGRLLNYRKDGSSFWNLLTIAPIKDDSGTIVNLIGVQLEVSKYTEGS 347
Query: 1126 KDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGS 1305
++ LRPNGLP+SLIKYD R +D + +++L+ AL P + R + R S G
Sbjct: 348 RENRLRPNGLPQSLIKYDVRHQDKVSALIAQLVAALTKPHKVEPPRPSYTMRFSL--TGQ 405
Query: 1306 ALVPGKRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGNVEKN 1485
+ P + R + + + + + +P G R+ +F+ L+GM + E++
Sbjct: 406 TIEPLSPGQAAAAARPCST---SDVPQTTSIPREGRGRRRHRSSTFLSLLGMEEKDSEED 462
Query: 1486 ILKPREDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIF 1665
E L+ + R+GIDLATTLERI +FVITDPRLP NPIIF
Sbjct: 463 QFP--EPELIMVADASVGRLRSSDDPERTRRGIDLATTLERIGHSFVITDPRLPGNPIIF 520
Query: 1666 ASDSFLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKK 1845
ASD FL LTEY REE+LG NCRFLQG +TDR V+ IRDA++ +VTVQL+NYTK G+
Sbjct: 521 ASDQFLELTEYSREEVLGENCRFLQGRDTDRKAVQLIRDAVEEGRDVTVQLLNYTKGGRP 580
Query: 1846 FWNLFHLQPMRDQKGDVQYFIGVQ--LDGTERVREAAAKDGAILVKKTADNIDEAAKELP 2019
FWNLFHLQ MRD+KG++QYFIGVQ D +RV AK +V+ TA N+D AA+ELP
Sbjct: 581 FWNLFHLQAMRDKKGNLQYFIGVQQETDTPDRVEHEKAK----VVRATAQNVDVAARELP 636
Query: 2020 DANLRPEDLWANHSKPVLPKPHMK-DTASWRAIQKV---LENGESIDLKHFRPVRP 2175
DANL + LW HSK V P PH K ++ W AI++V L GE + LKHFRP++P
Sbjct: 637 DANLTLDHLWERHSKEVTPLPHSKINSPCWYAIRRVQRRLRRGERLGLKHFRPIKP 692
Score = 99.8 bits (247), Expect = 2e-19
Identities = 57/160 (35%), Positives = 84/160 (52%), Gaps = 13/160 (8%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L +FV++D P +PI++AS F +T YS EV+G NCRFLQG TD
Sbjct: 494 DLATTLERIGHSFVITDPRLPGNPIIFASDQFLELTEYSREEVLGENCRFLQGRDTDRKA 553
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVE------- 1101
+ IR A+ G + ++LNY K G PFWNL + ++D+ G + FIG+Q E
Sbjct: 554 VQLIRDAVEEGRDVTVQLLNYTKGGRPFWNLFHLQAMRDKKGNLQYFIGVQQETDTPDRV 613
Query: 1102 ------VSKYTEGNKDTALRPNGLPESLIKYDARQKDHAR 1203
V + T N D A R LP++ + D + H++
Sbjct: 614 EHEKAKVVRATAQNVDVAARE--LPDANLTLDHLWERHSK 651
>sptr|Q84KS4|Q84KS4 Phytochrome 3 (Fragment).
Length = 657
Score = 426 bits (1094), Expect = e-117
Identities = 228/467 (48%), Positives = 303/467 (64%), Gaps = 4/467 (0%)
Frame = +1
Query: 787 FQQT-FVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVEISKIRQA 963
FQQT FVV DA +PD PI++AS GFFN+TGY+S EV+G NCR LQG T+P +++ IR+A
Sbjct: 217 FQQTSFVVVDALKPDLPIIFASTGFFNLTGYTSTEVIGANCRLLQGPDTNPEDVASIREA 276
Query: 964 LAAGSN-YCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEGNKDTAL 1140
LA + +C ++LNY+KDG+ FWNLLT+APIKD+ G ++K IG+Q+EVSKYTEG++ L
Sbjct: 277 LAQDTGTFCRKLLNYRKDGSSFWNLLTIAPIKDDRGSIVKLIGVQLEVSKYTEGSRANRL 336
Query: 1141 RPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRNNTLKRKSQESAGSALVPG 1320
RPNGLP+SLIKYD R +D V+EL+ AL P + ++ ++ + S G + P
Sbjct: 337 RPNGLPQSLIKYDVRHRDKVSVFVAELVAALTKPDKVELTKPSSTMQFSL--TGQTIKP- 393
Query: 1321 KRSSETGSRRNSHSGMRNSLQKISEVPEGGNKTRKSGLRSFMGLIGMGHGN-VEKNILKP 1497
L K + +P G R+ G +F+ L+G+ + VE +P
Sbjct: 394 -------------------LSKTAAMPREGGGRRRHGSNTFLSLLGVEKKDPVEDQFPEP 434
Query: 1498 REDPLLXXXXXXXXXXXXXXXKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDS 1677
R L+ + R+GIDLATTLERI ++FVITDPRLP+NPIIFASD
Sbjct: 435 R---LIMVDDNSVGRPESLDDPERTRRGIDLATTLERIGQSFVITDPRLPNNPIIFASDQ 491
Query: 1678 FLRLTEYCREEILGRNCRFLQGPETDRGTVKKIRDAIDNQTEVTVQLINYTKSGKKFWNL 1857
FL LTEY REE+LG NC FLQG +TD TV+ IRDA+ Q +VTVQL+NYT+ G+ FWNL
Sbjct: 492 FLELTEYSREEVLGNNCSFLQGRDTDANTVQLIRDAVAEQRDVTVQLLNYTRGGRPFWNL 551
Query: 1858 FHLQPMRDQKGDVQYFIGVQLDGTERVREAAAKDGAILVKKTADNIDEAAKELPDANLRP 2037
FHL MRD+KG++QYFIGVQ + D V TA N+D AA+ELPDANL P
Sbjct: 552 FHLHAMRDEKGELQYFIGVQQETVAPRVPEDMPDRVKQVHTTAQNVDVAARELPDANLSP 611
Query: 2038 EDLWANHSKPVLPKPHMK-DTASWRAIQKVLENGESIDLKHFRPVRP 2175
+ LW HSK + P PH K + +SW AI++V + + LKHFRP++P
Sbjct: 612 DHLWVRHSKVITPLPHSKMNNSSWYAIRRV-QRRVRLGLKHFRPIKP 657
Score = 97.1 bits (240), Expect = 2e-18
Identities = 47/113 (41%), Positives = 68/113 (60%)
Frame = +1
Query: 763 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 942
+L L Q+FV++D P++PI++AS F +T YS EV+G NC FLQG TD
Sbjct: 461 DLATTLERIGQSFVITDPRLPNNPIIFASDQFLELTEYSREEVLGNNCSFLQGRDTDANT 520
Query: 943 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVE 1101
+ IR A+A + ++LNY + G PFWNL + ++DE G + FIG+Q E
Sbjct: 521 VQLIRDAVAEQRDVTVQLLNYTRGGRPFWNLFHLHAMRDEKGELQYFIGVQQE 573
>sptr|Q9SA43|Q9SA43 F3O9.24 protein.
Length = 497
Score = 330 bits (846), Expect = 9e-89
Identities = 177/396 (44%), Positives = 229/396 (57%), Gaps = 39/396 (9%)
Frame = +1
Query: 2077 KPHMKDTASWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGE--YFAMKA 2250
KPH W A+ + G + + FR ++ LG GD GSV+LVEL G YFAMK
Sbjct: 84 KPHTGGDIRWDAVNSLKSRGIKLGISDFRVLKRLGYGDIGSVYLVELKGANPTTYFAMKV 143
Query: 2251 MDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLD 2430
MDK+ +++RNK+ RA ER+IL LDHPFLPTLY+ F+T CL++++C+GG L+ L
Sbjct: 144 MDKASLVSRNKLLRAQTEREILSQLDHPFLPTLYSHFETDKFYCLVMEFCSGGNLYSLRQ 203
Query: 2431 RQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLT 2610
+QP K EDA RF+A+EV+ ALEYLH GI+YRDLKPEN+L+ DGHI L+DFDLS
Sbjct: 204 KQPNKCFTEDAARFFASEVLLALEYLHMLGIVYRDLKPENVLVRDDGHIMLSDFDLSLRC 263
Query: 2611 SC---------------------------RPQVFLPHDIDXXXXXXXXXXN-----PIFF 2694
S +P F P + + P
Sbjct: 264 SVNPTLVKSFNGGGTTGIIDDNAAVQGCYQPSAFFPRMLQSSKKNRKSKSDFDGSLPELM 323
Query: 2695 AEPMRA-SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTF 2871
AEP S SFVGT EY+APEII GH SAVDWW GI +YE+L+G TPF+G+ + T
Sbjct: 324 AEPTNVKSMSFVGTHEYLAPEIIKNEGHGSAVDWWTFGIFIYELLHGATPFKGQGNKATL 383
Query: 2872 ANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVR 3051
N++ + +RFP QVS A+ LI LL ++P NR+ GA EIKQHPFF G+NWAL+R
Sbjct: 384 YNVIGQPLRFPEYSQVSSTAKDLIKGLLVKEPQNRIAYKRGATEIKQHPFFEGVNWALIR 443
Query: 3052 AATPPELEAPLQDTL----EATLPPPAADAAHTDMF 3147
TPP L P+ + E PPAA + MF
Sbjct: 444 GETPPHLPEPVDFSCYVKKEKESLPPAATEKKSKMF 479
>sptr|Q9USX7|Q9USX7 Putative ser/thr protein kinase.
Length = 526
Score = 303 bits (775), Expect = 1e-80
Identities = 163/323 (50%), Positives = 201/323 (62%), Gaps = 5/323 (1%)
Frame = +1
Query: 2158 FRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPF 2337
F +R LG GD G V+LV FAMK ++K M+ R+KV+R AE++IL HPF
Sbjct: 155 FEKIRLLGQGDVGKVYLVRQKSNHRLFAMKILNKREMIKRHKVNRVLAEQEILTKSKHPF 214
Query: 2338 LPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQ 2517
+ TLY SFQ++ ++ L ++YCAGGE F L P +L E FYAAEV ALEYLH
Sbjct: 215 IVTLYHSFQSRDYLYLCMEYCAGGEFFRALHSLPKHILPEKDACFYAAEVTAALEYLHLM 274
Query: 2518 GIIYRDLKPENILLHRDGHISLTDFDLSCLTS--CRPQVFLPHDIDXXXXXXXXXXNPIF 2691
G IYRDLKPENILLH+ GHI L+DFDLS S P V LP N F
Sbjct: 275 GFIYRDLKPENILLHQSGHIMLSDFDLSKPISIVTHPTVVLPKHSTFSQEKPALDTNSYF 334
Query: 2692 FAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTF 2871
+NSFVGTEEYIAPE+I GHT AVDWW LGI +YE+LYG TPF+GK R TF
Sbjct: 335 ---SNFRTNSFVGTEEYIAPEVIRSCGHTVAVDWWTLGIFIYEILYGTTPFKGKNRHATF 391
Query: 2872 ANILHKDIRFP---ASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWA 3042
+NIL+ D+ FP + VS + LI RLL +D + R GS GA +IKQHPFFR I WA
Sbjct: 392 SNILYSDVSFPEYHGAPNVSSTCKSLIRRLLVKDESKRCGSVAGASDIKQHPFFRHIQWA 451
Query: 3043 LVRAATPPELEAPLQDTLEATLP 3111
L+R+ PP + ++D +EA P
Sbjct: 452 LLRSMKPP-IIPKIEDGMEAVEP 473
>sptrnew|EAA27239|EAA27239 Hypothetical protein.
Length = 623
Score = 301 bits (771), Expect = 4e-80
Identities = 158/311 (50%), Positives = 203/311 (65%), Gaps = 8/311 (2%)
Frame = +1
Query: 2158 FRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPF 2337
F ++ +G GD G V+LV+ +G +AMK + K M+ RNK+ RA AE++IL +HPF
Sbjct: 242 FDKIKLIGKGDVGKVYLVKEKKSGRLYAMKVLSKKEMIKRNKIKRALAEQEILATSNHPF 301
Query: 2338 LPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQ 2517
+ TLY SFQ++ ++ L ++YC+GGE F L +P K + ED RFYAAEV ALEYLH
Sbjct: 302 IVTLYHSFQSEDYLYLCMEYCSGGEFFRALQTRPGKCIPEDDARFYAAEVTAALEYLHLM 361
Query: 2518 GIIYRDLKPENILLHRDGHISLTDFDLSCLT--SCRPQVFLPHDIDXXXXXXXXXXNPIF 2691
G IYRDLKPENILLH+ GHI L+DFDLS + +P + + + P
Sbjct: 362 GFIYRDLKPENILLHQSGHIMLSDFDLSKQSDPGGKPTMIIGKN------GTSTSSLPTI 415
Query: 2692 FAEPMRA---SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQ 2862
+ A +NSFVGTEEYIAPE+I G+GHTSAVDWW LGIL+YEMLYG TPF+GK R
Sbjct: 416 DTKSCIANFRTNSFVGTEEYIAPEVIKGSGHTSAVDWWTLGILIYEMLYGTTPFKGKNRN 475
Query: 2863 RTFANILHKDIRFP---ASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGI 3033
TFANIL +DI FP + Q+S + LI +LL +D RLG+ GA +IK HPFFR
Sbjct: 476 ATFANILREDIPFPDHAGAPQISNLCKSLIRKLLIKDENRRLGARAGASDIKTHPFFRTT 535
Query: 3034 NWALVRAATPP 3066
WAL+R PP
Sbjct: 536 QWALIRHMKPP 546
>sw|O42626|NRC2_NEUCR Serine/threonine protein kinase nrc-2 (EC
2.7.1.37) (Nonrepressible conidiation protein 2).
Length = 623
Score = 301 bits (771), Expect = 4e-80
Identities = 158/311 (50%), Positives = 203/311 (65%), Gaps = 8/311 (2%)
Frame = +1
Query: 2158 FRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPF 2337
F ++ +G GD G V+LV+ +G +AMK + K M+ RNK+ RA AE++IL +HPF
Sbjct: 242 FDKIKLIGKGDVGKVYLVKEKKSGRLYAMKVLSKKEMIKRNKIKRALAEQEILATSNHPF 301
Query: 2338 LPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQ 2517
+ TLY SFQ++ ++ L ++YC+GGE F L +P K + ED RFYAAEV ALEYLH
Sbjct: 302 IVTLYHSFQSEDYLYLCMEYCSGGEFFRALQTRPGKCIPEDDARFYAAEVTAALEYLHLM 361
Query: 2518 GIIYRDLKPENILLHRDGHISLTDFDLSCLT--SCRPQVFLPHDIDXXXXXXXXXXNPIF 2691
G IYRDLKPENILLH+ GHI L+DFDLS + +P + + + P
Sbjct: 362 GFIYRDLKPENILLHQSGHIMLSDFDLSKQSDPGGKPTMIIGKN------GTSTSSLPTI 415
Query: 2692 FAEPMRA---SNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQ 2862
+ A +NSFVGTEEYIAPE+I G+GHTSAVDWW LGIL+YEMLYG TPF+GK R
Sbjct: 416 DTKSCIANFRTNSFVGTEEYIAPEVIKGSGHTSAVDWWTLGILIYEMLYGTTPFKGKNRN 475
Query: 2863 RTFANILHKDIRFP---ASIQVSLAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGI 3033
TFANIL +DI FP + Q+S + LI +LL +D RLG+ GA +IK HPFFR
Sbjct: 476 ATFANILREDIPFPDHAGAPQISNLCKSLIRKLLIKDENRRLGARAGASDIKTHPFFRTT 535
Query: 3034 NWALVRAATPP 3066
WAL+R PP
Sbjct: 536 QWALIRHMKPP 546
>sw|Q09831|KAD5_SCHPO Probable serine/threonine-protein kinase C4G8.05
(EC 2.7.1.-).
Length = 566
Score = 298 bits (764), Expect = 3e-79
Identities = 173/408 (42%), Positives = 234/408 (57%), Gaps = 24/408 (5%)
Frame = +1
Query: 1915 QLDGTERVREA---AAKDGAILVKKTADNIDEAAKELP-----------DANLRPE---D 2043
++DGT E + D ++KK+ K++P D +++ E D
Sbjct: 94 KIDGTGSSAEGGDGSGTDSISVIKKSFFKSGRKKKDVPKSRNVSRSNGADTSVQREKLKD 153
Query: 2044 LWANHSKPVLPKPHMKDTASWRAIQKVLENGESIDLK----HFRPVRPLGSGDTGSVHLV 2211
+++ H K H+K T + RA + + D++ F V LG GD G V+LV
Sbjct: 154 IFSPHGKEK-ELAHIKKTVATRARTYSSNSIKICDVEVGPSSFEKVFLLGKGDVGRVYLV 212
Query: 2212 ELLGTGEYFAMKAMDKSVMLNRNKVHRATAERQILDMLDHPFLPTLYASFQTKTHICLIV 2391
+G+++AMK + K M+ RNK RA AE+ IL +HPF+ TLY SFQ+ ++ L +
Sbjct: 213 REKKSGKFYAMKVLSKQEMIKRNKSKRAFAEQHILATSNHPFIVTLYHSFQSDEYLYLCM 272
Query: 2392 DYCAGGELFMLLDRQPMKVLKEDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDG 2571
+YC GGE F L R+P + L E+ +FY AEV ALEYLH G IYRDLKPENILLH G
Sbjct: 273 EYCMGGEFFRALQRRPGRCLSENEAKFYIAEVTAALEYLHLMGFIYRDLKPENILLHESG 332
Query: 2572 HISLTDFDLSCLTSCRPQVFLPHDIDXXXXXXXXXXNPIFFAEPMRASNSFVGTEEYIAP 2751
HI L+DFDLS ++ + + + R +NSFVGTEEYIAP
Sbjct: 333 HIMLSDFDLSKQSNSAGAPTVIQARNAPSAQNAYALDTKSCIADFR-TNSFVGTEEYIAP 391
Query: 2752 EIITGAGHTSAVDWWALGILLYEMLYGYTPFRGKTRQRTFANILHKDIRFPA---SIQVS 2922
E+I G GHTSAVDWW LGIL YEMLY TPF+GK R TF+NILHKD+ FP + +S
Sbjct: 392 EVIKGCGHTSAVDWWTLGILFYEMLYATTPFKGKNRNMTFSNILHKDVIFPEYADAPSIS 451
Query: 2923 LAARQLIYRLLHRDPANRLGSYEGAMEIKQHPFFRGINWALVRAATPP 3066
+ LI +LL +D +RLGS GA ++K HPFF+ + WAL+R PP
Sbjct: 452 SLCKNLIRKLLVKDENDRLGSQAGAADVKLHPFFKNVQWALLRHTEPP 499
Database: /db/trembl-ebi/tmp/swall
Posted date: Nov 7, 2003 8:22 PM
Number of letters in database: 415,658,970
Number of sequences in database: 1,302,759
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,577,286,855
Number of Sequences: 1302759
Number of extensions: 58774421
Number of successful extensions: 183095
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 147979
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 176001
length of database: 415,658,970
effective HSP length: 134
effective length of database: 241,089,264
effective search space used: 235062032400
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

