BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
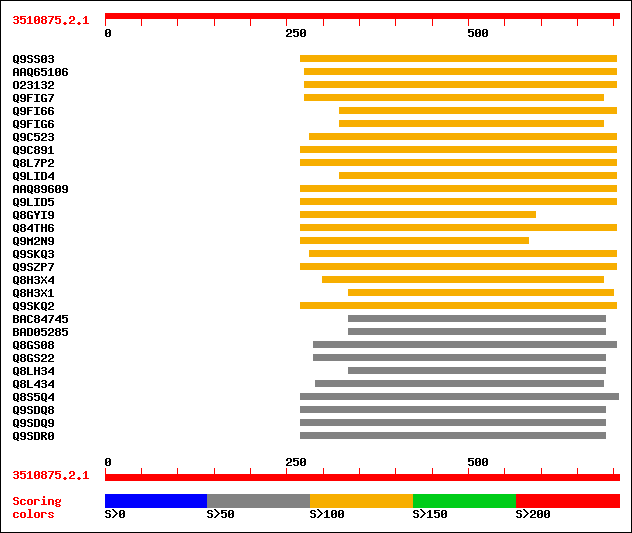
Query= 3510875.2.1
(707 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9SS03|Q9SS03 F12P19.3. 138 6e-32
sptrnew|AAQ65106|AAQ65106 At1g22900. 131 1e-29
sptr|O23132|O23132 F19G10.14 protein. 131 1e-29
sptr|Q9FIG7|Q9FIG7 Disease resistance response protein-like (Hyp... 129 3e-29
sptr|Q9FI66|Q9FI66 Similarity to disease resistance response pro... 124 1e-27
sptr|Q9FIG6|Q9FIG6 Similarity to disease resistance response pro... 123 2e-27
sptr|Q9C523|Q9C523 Dirigent protein, putative (Hypothetical prot... 123 3e-27
sptr|Q9C891|Q9C891 Hypothetical protein. 122 4e-27
sptr|Q8L7P2|Q8L7P2 Hypothetical protein. 122 5e-27
sptr|Q9LID4|Q9LID4 Similarity to disease resistance response pro... 122 5e-27
sptrnew|AAQ89609|AAQ89609 At3g13650. 120 1e-26
sptr|Q9LID5|Q9LID5 Disease resistance response protein-like (Put... 120 1e-26
sptr|Q8GYI9|Q8GYI9 Hypothetical protein. 118 7e-26
sptr|Q84TH6|Q84TH6 Putative disease resistance response protein/... 117 2e-25
sptr|Q9M2N9|Q9M2N9 Hypothetical protein. 117 2e-25
sptr|Q9SKQ3|Q9SKQ3 Putative disease resistance response protein. 114 1e-24
sptr|Q9SZP7|Q9SZP7 Disease resistance response like protein. 111 1e-23
sptr|Q8H3X4|Q8H3X4 P0585H11.3 protein. 105 5e-22
sptr|Q8H3X1|Q8H3X1 P0585H11.8 protein. 105 6e-22
sptr|Q9SKQ2|Q9SKQ2 Putative disease resistance response protein. 104 1e-21
sptrnew|BAC84745|BAC84745 Disease resistance response protein-li... 88 1e-16
sptrnew|BAD05285|BAD05285 Disease resistance response protein-like. 87 2e-16
sptr|Q8GS08|Q8GS08 P0552F09.12 protein (B1056G08.30 protein). 82 9e-15
sptr|Q8GS22|Q8GS22 P0552F09.4 protein (B1056G08.22 protein). 78 1e-13
sptr|Q8LH34|Q8LH34 P0617C02.32 protein (B1317D11.1 protein). 76 5e-13
sptr|Q8L434|Q8L434 Hypothetical protein. 73 3e-12
sptr|Q8S5Q4|Q8S5Q4 Putative disease resistance response protein. 72 1e-11
sptr|Q9SDQ8|Q9SDQ8 Dirigent-like protein. 69 5e-11
sptr|Q9SDQ9|Q9SDQ9 Dirigent protein. 69 8e-11
sptr|Q9SDR0|Q9SDR0 Dirigent-like protein. 67 2e-10
>sptr|Q9SS03|Q9SS03 F12P19.3.
Length = 189
Score = 138 bits (348), Expect = 6e-32
Identities = 71/145 (48%), Positives = 87/145 (60%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT YFHD+V+G PT+V++A FG V +DD LT GP EVGRA
Sbjct: 44 LTHLHFYFHDIVSGDKPTSVQVANGPTTNSSATGFGLVAVVDDKLTVGP-EITSEEVGRA 102
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y ADQ GLLM N VFT G+ + ST+++ GRN VLS VREM I+GG+G FR R
Sbjct: 103 QGMYASADQNKLGLLMAFNLVFTKGKFSDSTVAMYGRNPVLSKVREMPIIGGTGAFRFGR 162
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GY A T+ TSG VV+Y V +
Sbjct: 163 GYALAKTLVFNITSGDAVVEYNVYI 187
>sptrnew|AAQ65106|AAQ65106 At1g22900.
Length = 187
Score = 131 bits (329), Expect = 1e-29
Identities = 66/143 (46%), Positives = 86/143 (60%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT YFHD+++G PT +R+A+ FGAV+ +D P+T GP + EVGRA
Sbjct: 44 LTHLHFYFHDIISGDKPTTIRVAEAPGTNSSATVFGAVLIVDAPVTEGP-ELSSKEVGRA 102
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y D +TFG MV NFVFT GE NGST ++ GRN +L + RE+ I+GG+G FR AR
Sbjct: 103 QGLYASTDMKTFGFTMVFNFVFTEGEFNGSTAALYGRNPILLEERELPIIGGTGDFRFAR 162
Query: 343 GYVQAHTIDSGATSGTTVVQYTV 275
GY T + VV+Y V
Sbjct: 163 GYALPKTYK--VVNIDAVVEYNV 183
>sptr|O23132|O23132 F19G10.14 protein.
Length = 187
Score = 131 bits (329), Expect = 1e-29
Identities = 66/143 (46%), Positives = 86/143 (60%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT YFHD+++G PT +R+A+ FGAV+ +D P+T GP + EVGRA
Sbjct: 44 LTHLHFYFHDIISGDKPTTIRVAEAPGTNSSATVFGAVLIVDAPVTEGP-ELSSKEVGRA 102
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y D +TFG MV NFVFT GE NGST ++ GRN +L + RE+ I+GG+G FR AR
Sbjct: 103 QGLYASTDMKTFGFTMVFNFVFTEGEFNGSTAALYGRNPILLEERELPIIGGTGDFRFAR 162
Query: 343 GYVQAHTIDSGATSGTTVVQYTV 275
GY T + VV+Y V
Sbjct: 163 GYALPKTYK--VVNIDAVVEYNV 183
>sptr|Q9FIG7|Q9FIG7 Disease resistance response protein-like
(Hypothetical protein) (At5g42500).
Length = 185
Score = 129 bits (325), Expect = 3e-29
Identities = 67/137 (48%), Positives = 84/137 (61%)
Frame = -2
Query: 685 YFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRAQGTYTF 506
YFHDV++G PTAV++A+ NFG ++ DDPLT GP ++ EVGRAQG Y
Sbjct: 48 YFHDVISGDKPTAVKVAEARRTNSSNVNFGVIMIADDPLTEGPDPSS-KEVGRAQGMYAL 106
Query: 505 ADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARGYVQAH 326
+ MV N FTAGE NGST+++ GRNE+ S VREM I+GG+G FR ARGY QA
Sbjct: 107 TAMKNISFTMVFNLAFTAGEFNGSTVAMYGRNEIFSKVREMPIIGGTGAFRFARGYAQAK 166
Query: 325 TIDSGATSGTTVVQYTV 275
T VV+Y V
Sbjct: 167 TYK--VVGLDAVVEYNV 181
>sptr|Q9FI66|Q9FI66 Similarity to disease resistance response
protein.
Length = 191
Score = 124 bits (311), Expect = 1e-27
Identities = 64/127 (50%), Positives = 81/127 (63%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY+HD+V G NP+++RI FG++ ID+ LT T VG+A
Sbjct: 45 LTHLRVYWHDIVTGRNPSSIRIQGPVAKYSSSSYFGSITMIDNALTLD-VPINSTVVGQA 103
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y A Q+ GLLM MN F G++NGST++ILGRN V+S VREM +VGGSG FR AR
Sbjct: 104 QGMYVGAAQKEIGLLMAMNLAFKTGKYNGSTITILGRNTVMSKVREMPVVGGSGMFRFAR 163
Query: 343 GYVQAHT 323
GYV+A T
Sbjct: 164 GYVEART 170
>sptr|Q9FIG6|Q9FIG6 Similarity to disease resistance response
protein.
Length = 182
Score = 123 bits (309), Expect = 2e-27
Identities = 59/121 (48%), Positives = 78/121 (64%)
Frame = -2
Query: 685 YFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRAQGTYTF 506
YFHDV++G PTAV++A+ FG ++ DDPLT GP ++ EVGRAQG Y
Sbjct: 45 YFHDVISGDKPTAVKVAEARPTTTLNVKFGVIMIADDPLTEGPDPSS-KEVGRAQGMYAS 103
Query: 505 ADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARGYVQAH 326
+ MV N+VFTAGE NGST+++ GRN++ S VRE+ I+GG+G FR ARGY
Sbjct: 104 TAMKDIVFTMVFNYVFTAGEFNGSTIAVYGRNDIFSKVRELPIIGGTGAFRFARGYALPK 163
Query: 325 T 323
T
Sbjct: 164 T 164
>sptr|Q9C523|Q9C523 Dirigent protein, putative (Hypothetical
protein) (At1g58170).
Length = 185
Score = 123 bits (308), Expect = 3e-27
Identities = 63/141 (44%), Positives = 85/141 (60%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY+HD+V G + ++V I FG + ID+PLT P + + VGRA
Sbjct: 39 LTHFRVYWHDIVTGQDSSSVSIMNPPKKYTGATGFGLMRMIDNPLTLTP-KLSSKMVGRA 97
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y ++ GLLM MNF G++NGST+++LGRN V VREM ++GGSG FR AR
Sbjct: 98 QGFYAGTSKEEIGLLMAMNFAILDGKYNGSTITVLGRNSVFDKVREMPVIGGSGLFRFAR 157
Query: 343 GYVQAHTIDSGATSGTTVVQY 281
GYVQA T + +G +V+Y
Sbjct: 158 GYVQASTHEFNLKTGNAIVEY 178
>sptr|Q9C891|Q9C891 Hypothetical protein.
Length = 187
Score = 122 bits (307), Expect = 4e-27
Identities = 68/145 (46%), Positives = 89/145 (61%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT +VY+HD+++G NPT++ I FGA+ ID+ LT T +G+A
Sbjct: 44 LTHFKVYWHDILSGPNPTSIMIQPPVTNSSY---FGAISMIDNALTA-KVPMNSTVLGQA 99
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y A Q+ G LM MNF F G++NGST++ILGRN LS+VREM IVGGSG FR AR
Sbjct: 100 QGFYAGAAQKELGFLMAMNFAFKTGKYNGSTITILGRNTALSEVREMPIVGGSGLFRFAR 159
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GYV+A T +G V+Y+ V
Sbjct: 160 GYVEARTKWINLKNGDATVEYSCYV 184
>sptr|Q8L7P2|Q8L7P2 Hypothetical protein.
Length = 187
Score = 122 bits (306), Expect = 5e-27
Identities = 68/145 (46%), Positives = 89/145 (61%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT +VY+HD+++G NPT++ I FGA+ ID+ LT T +G+A
Sbjct: 44 LTHFKVYWHDILSGPNPTSIMIQPP---FTNSSYFGAISMIDNALTA-KVPMNSTVLGQA 99
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y A Q+ G LM MNF F G++NGST++ILGRN LS+VREM IVGGSG FR AR
Sbjct: 100 QGFYAGAAQKELGFLMAMNFAFKTGKYNGSTITILGRNTALSEVREMPIVGGSGLFRFAR 159
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GYV+A T +G V+Y+ V
Sbjct: 160 GYVEARTKWINLKNGDATVEYSCYV 184
>sptr|Q9LID4|Q9LID4 Similarity to disease resistance response
protein.
Length = 186
Score = 122 bits (306), Expect = 5e-27
Identities = 68/127 (53%), Positives = 83/127 (65%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY+H+ V G NP++V I Q G++ +DDPLT R A T VG+A
Sbjct: 42 LTHLRVYWHNSVNGRNPSSVMIQQPVLNSSLS---GSITMMDDPLTFDVPRNA-TVVGQA 97
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y A Q G LMVMNF FT G++NGST++ILGRN V+S VREM +VGGSG FR AR
Sbjct: 98 QGMYVAAAQGEIGFLMVMNFAFTTGKYNGSTITILGRNVVMSKVREMPVVGGSGIFRFAR 157
Query: 343 GYVQAHT 323
GYV+A T
Sbjct: 158 GYVEART 164
>sptrnew|AAQ89609|AAQ89609 At3g13650.
Length = 186
Score = 120 bits (302), Expect = 1e-26
Identities = 67/145 (46%), Positives = 90/145 (62%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY+HD+++G+NP++V I FG+V ID+ LTT T VG+A
Sbjct: 43 LTHFRVYWHDILSGSNPSSVVINPPISNSSF---FGSVTVIDNRLTT-EVAVNSTLVGQA 98
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y Q+ LMVMNF F G++NGS+++ILGRN VL+ VREM ++GGSG FR AR
Sbjct: 99 QGIYAATGQRDASALMVMNFAFKTGKYNGSSIAILGRNAVLTKVREMPVIGGSGLFRFAR 158
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GYV+A T+ SG V+Y+ V
Sbjct: 159 GYVEARTMWFDQKSGDATVEYSCYV 183
>sptr|Q9LID5|Q9LID5 Disease resistance response protein-like
(Putative dirigent protein).
Length = 186
Score = 120 bits (302), Expect = 1e-26
Identities = 67/145 (46%), Positives = 90/145 (62%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY+HD+++G+NP++V I FG+V ID+ LTT T VG+A
Sbjct: 43 LTHFRVYWHDILSGSNPSSVVINPPISNSSF---FGSVTVIDNRLTT-EVAVNSTLVGQA 98
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y Q+ LMVMNF F G++NGS+++ILGRN VL+ VREM ++GGSG FR AR
Sbjct: 99 QGIYAATGQRDASALMVMNFAFKTGKYNGSSIAILGRNAVLTKVREMPVIGGSGLFRFAR 158
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GYV+A T+ SG V+Y+ V
Sbjct: 159 GYVEARTMWFDQKSGDATVEYSCYV 183
>sptr|Q8GYI9|Q8GYI9 Hypothetical protein.
Length = 110
Score = 118 bits (296), Expect = 7e-26
Identities = 60/108 (55%), Positives = 70/108 (64%)
Frame = -2
Query: 592 VVAIDDPLTTGPTRAAGTEVGRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGR 413
V +DD LT GP EVGRAQG + ADQ FGLLM NFVFT GE +GST+S+ GR
Sbjct: 2 VAVVDDILTVGP-EITSEEVGRAQGIFASADQNNFGLLMAFNFVFTKGEFSGSTVSMYGR 60
Query: 412 NEVLSDVREMSIVGGSGKFRMARGYVQAHTIDSGATSGTTVVQYTVNV 269
N + S VREM I+GG+G FR RGY QA T TSG VV+Y V +
Sbjct: 61 NPIFSKVREMPIIGGTGAFRFGRGYAQAKTFTFNTTSGNAVVKYNVYI 108
>sptr|Q84TH6|Q84TH6 Putative disease resistance response
protein/dirigent protein.
Length = 187
Score = 117 bits (293), Expect = 2e-25
Identities = 61/145 (42%), Positives = 86/145 (59%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
+T + YFHD ++G NPTAV++AQ FGAV +DD LT + VGRA
Sbjct: 42 VTNLQFYFHDTLSGKNPTAVKVAQGTDTEKSPTLFGAVFMVDDALTETADPKSKL-VGRA 100
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y + ++ GL+M M+F F G + ST+S++G+N ++ +REM IVGG+G FRMAR
Sbjct: 101 QGLYGSSCKEEVGLIMAMSFCFEDGPYKDSTISMIGKNSAMNPIREMPIVGGTGMFRMAR 160
Query: 343 GYVQAHTIDSGATSGTTVVQYTVNV 269
GY A T +G +V Y V +
Sbjct: 161 GYAIARTNWFDPKTGDAIVGYNVTI 185
>sptr|Q9M2N9|Q9M2N9 Hypothetical protein.
Length = 249
Score = 117 bits (292), Expect = 2e-25
Identities = 59/105 (56%), Positives = 69/105 (65%)
Frame = -2
Query: 583 IDDPLTTGPTRAAGTEVGRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEV 404
+DD LT GP EVGRAQG + ADQ FGLLM NFVFT GE +GST+S+ GRN +
Sbjct: 144 VDDILTVGP-EITSEEVGRAQGIFASADQNNFGLLMAFNFVFTKGEFSGSTVSMYGRNPI 202
Query: 403 LSDVREMSIVGGSGKFRMARGYVQAHTIDSGATSGTTVVQYTVNV 269
S VREM I+GG+G FR RGY QA T TSG VV+Y V +
Sbjct: 203 FSKVREMPIIGGTGAFRFGRGYAQAKTFTFNTTSGNAVVKYNVYI 247
>sptr|Q9SKQ3|Q9SKQ3 Putative disease resistance response protein.
Length = 276
Score = 114 bits (286), Expect = 1e-24
Identities = 60/141 (42%), Positives = 84/141 (59%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
+T + YFHD ++G NPTAV++AQ FGAV +DD LT + VGRA
Sbjct: 42 VTNLQFYFHDTLSGKNPTAVKVAQGTDTEKSPTLFGAVFMVDDALTETADPKSKL-VGRA 100
Query: 523 QGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
QG Y + ++ GL+M M+F F G + ST+S++G+N ++ +REM IVGG+G FRMAR
Sbjct: 101 QGLYGSSCKEEVGLIMAMSFCFEDGPYKDSTISMIGKNSAMNPIREMPIVGGTGMFRMAR 160
Query: 343 GYVQAHTIDSGATSGTTVVQY 281
GY A T +G +V Y
Sbjct: 161 GYAIARTNWFDPKTGDAIVGY 181
>sptr|Q9SZP7|Q9SZP7 Disease resistance response like protein.
Length = 176
Score = 111 bits (277), Expect = 1e-23
Identities = 64/151 (42%), Positives = 86/151 (56%), Gaps = 6/151 (3%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXX--NFGAVVAIDDPLTTGPTRAAGTEVG 530
+T Y HD ++G NPTAVRIA FG++ IDDPLT GP + + E+G
Sbjct: 24 VTRVRFYLHDTLSGQNPTAVRIAHANLTGGSASPVGFGSLFVIDDPLTVGPEKHS-KEIG 82
Query: 529 RAQGTYTFA--DQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDV--REMSIVGGSG 362
QG Y D F ++M + FTAG+ NGS++SI RN V +V RE++IVGG G
Sbjct: 83 NGQGMYVSGCKDLSKFTIVMYADLAFTAGKFNGSSISIFSRNPVAEEVGEREIAIVGGRG 142
Query: 361 KFRMARGYVQAHTIDSGATSGTTVVQYTVNV 269
KFRMARG+V+ T +G V++Y V
Sbjct: 143 KFRMARGFVKVKTNKIDMKTGDAVLRYDATV 173
>sptr|Q8H3X4|Q8H3X4 P0585H11.3 protein.
Length = 229
Score = 105 bits (263), Expect = 5e-22
Identities = 55/131 (41%), Positives = 81/131 (61%), Gaps = 2/131 (1%)
Frame = -2
Query: 685 YFHDVVAGTNPTAVRIAQ--XAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRAQGTY 512
+ HDVV+G+ TAV++ + + FG +DD LT + A EVGRAQG Y
Sbjct: 35 FMHDVVSGSGQTAVQVIKGPTSANGGVSTGFGDTTVVDDALTE-TSSATSPEVGRAQGFY 93
Query: 511 TFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARGYVQ 332
+ + L+M +N FTAGE+NGST++++G ++ + VRE+S+VGG+GKFRMA GYV
Sbjct: 94 MMSSLSSPTLMMCVNLYFTAGENNGSTIAVIGHDDTTATVRELSVVGGTGKFRMATGYVV 153
Query: 331 AHTIDSGATSG 299
T A++G
Sbjct: 154 WKTASMSASTG 164
>sptr|Q8H3X1|Q8H3X1 P0585H11.8 protein.
Length = 220
Score = 105 bits (262), Expect = 6e-22
Identities = 57/125 (45%), Positives = 76/125 (60%), Gaps = 3/125 (2%)
Frame = -2
Query: 700 TTSEVYFHDVVAGTNPTAVRIAQX---AXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVG 530
T ++ HDV+ G + TAV + A FG VV +DD LT GP+R++ VG
Sbjct: 50 THLHLFMHDVLTGPDATAVDVVNGTGRAFDVAGGLRFGQVVVMDDVLTEGPSRSS-PRVG 108
Query: 529 RAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRM 350
R QG Y F+D LL MN V TAG + GST++ILGR+ + +RE+S+VGG+G FRM
Sbjct: 109 RTQGFYVFSDMNVPALLFCMNVVLTAGPYAGSTVTILGRDHITQPLRELSVVGGTGAFRM 168
Query: 349 ARGYV 335
A GYV
Sbjct: 169 ATGYV 173
>sptr|Q9SKQ2|Q9SKQ2 Putative disease resistance response protein.
Length = 186
Score = 104 bits (260), Expect = 1e-21
Identities = 53/149 (35%), Positives = 87/149 (58%), Gaps = 4/149 (2%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXN----FGAVVAIDDPLTTGPTRAAGTE 536
+T +FHD + NP+A+ IA+ + FG++ A+DDPLT GP + +
Sbjct: 36 VTNLHFFFHDTLTAPNPSAILIAKPTHTRGDNDSSPSPFGSLFALDDPLTVGPDPKS-EK 94
Query: 535 VGRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKF 356
+G A+G Y + + L M ++F FT+G+ NGS++++ RN + RE+++VGG G+F
Sbjct: 95 IGNARGMYVSSGKHVPTLTMYVDFGFTSGKFNGSSIAVFSRNTITEKEREVAVVGGRGRF 154
Query: 355 RMARGYVQAHTIDSGATSGTTVVQYTVNV 269
RMARG Q +T T+G +V+Y V +
Sbjct: 155 RMARGVAQLNTYYVNLTNGDAIVEYNVTL 183
>sptrnew|BAC84745|BAC84745 Disease resistance response protein-like
protein.
Length = 209
Score = 88.2 bits (217), Expect = 1e-16
Identities = 50/122 (40%), Positives = 70/122 (57%), Gaps = 4/122 (3%)
Frame = -2
Query: 688 VYFHDVVAGTNPTAVRIAQXAXXXXXXXN----FGAVVAIDDPLTTGPTRAAGTEVGRAQ 521
VY HD+V G T+V + + + FG V IDD +T GP+ A+ EVGRAQ
Sbjct: 42 VYMHDIVGGAGQTSVVVVKGPGPANPSMSPGNNFGDTVIIDDVVTEGPSLAS-REVGRAQ 100
Query: 520 GTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARG 341
GTY A ++ + V T G +NGST+ I GR++ +VRE+++VGGSG R A G
Sbjct: 101 GTYMLASMARPVFIVDITVVLTDGPYNGSTILIAGRDDTSEEVRELAVVGGSGMLRRASG 160
Query: 340 YV 335
+V
Sbjct: 161 HV 162
>sptrnew|BAD05285|BAD05285 Disease resistance response protein-like.
Length = 207
Score = 87.0 bits (214), Expect = 2e-16
Identities = 50/123 (40%), Positives = 71/123 (57%), Gaps = 5/123 (4%)
Frame = -2
Query: 688 VYFHDVVAGTNPTAVRIAQXAX---XXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRAQG 518
+Y HDV+ G TA+R+ A FG VA+DD +T GP+ A+ VGRAQG
Sbjct: 51 LYMHDVIDGPGQTAIRLIWGAGPPHASMPGRAFGDTVAVDDLVTEGPSIASAA-VGRAQG 109
Query: 517 TYTFADQQTFGLLMVMNFVFTA--GEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMAR 344
TY + Q+ L++ + T+ G +NGS L I GR+ V + RE+++VGG+G R A
Sbjct: 110 TYMLSSQREAVLVVAITVALTSAGGPYNGSNLVIAGRDRVRDETRELAVVGGTGALRGAA 169
Query: 343 GYV 335
GYV
Sbjct: 170 GYV 172
>sptr|Q8GS08|Q8GS08 P0552F09.12 protein (B1056G08.30 protein).
Length = 189
Score = 81.6 bits (200), Expect = 9e-15
Identities = 54/143 (37%), Positives = 71/143 (49%), Gaps = 4/143 (2%)
Frame = -2
Query: 703 LTTSEVYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRA 524
LT VY H+ AG N TA+ + FG V +DD L GP RA+ +GR
Sbjct: 38 LTRIRVYMHERFAGANATALAVVPSPLGANEA--FGRVAVLDDELRDGPDRASSALIGRF 95
Query: 523 QG----TYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKF 356
QG T ++ VFTAGEH GSTLS++G + E +VGG+G F
Sbjct: 96 QGVVAGTSLPGTAPPASFQSAISLVFTAGEHAGSTLSMVGPVLGFAGAIERPLVGGTGAF 155
Query: 355 RMARGYVQAHTIDSGATSGTTVV 287
RMARGY T + A++ +VV
Sbjct: 156 RMARGYC-VMTAAAAASTAVSVV 177
>sptr|Q8GS22|Q8GS22 P0552F09.4 protein (B1056G08.22 protein).
Length = 175
Score = 78.2 bits (191), Expect = 1e-13
Identities = 53/134 (39%), Positives = 64/134 (47%)
Frame = -2
Query: 688 VYFHDVVAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVGRAQGTYT 509
+Y H+ +G N T + FG V D+ L TG RAA VGR QG
Sbjct: 34 LYMHETFSGPNATEGGVVASPFNTT----FGQVAVFDNELRTGEDRAASPLVGRYQGFIV 89
Query: 508 FADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARGYVQA 329
+ + G L FTAGE NGSTLS+ G + E SIVGG+GK R+ARGY
Sbjct: 90 GTGRSSPGYLTSATVAFTAGELNGSTLSLEGPFFGFAGTAERSIVGGTGKLRLARGYYLL 149
Query: 328 HTIDSGATSGTTVV 287
I G TS T V
Sbjct: 150 KLI--GKTSPETAV 161
>sptr|Q8LH34|Q8LH34 P0617C02.32 protein (B1317D11.1 protein).
Length = 204
Score = 75.9 bits (185), Expect = 5e-13
Identities = 43/122 (35%), Positives = 66/122 (54%), Gaps = 4/122 (3%)
Frame = -2
Query: 688 VYFHDVVAGTNPTAVRIAQXAXXXXXX----XNFGAVVAIDDPLTTGPTRAAGTEVGRAQ 521
+Y HD+ G TAV++ +FG + +DD LT VGRAQ
Sbjct: 32 LYMHDITGGPGQTAVQVVNGTGPLHPAMPPGSHFGDTMVVDDLLTDVDD---SKPVGRAQ 88
Query: 520 GTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFRMARG 341
G+YT A + ++ + V T G + GST+ I GR+++ +VRE+++VGG+GK R A G
Sbjct: 89 GSYTLACLRAPVFVVSITLVLTDGPYKGSTILIAGRDDISEEVRELAVVGGTGKLRRATG 148
Query: 340 YV 335
+V
Sbjct: 149 HV 150
>sptr|Q8L434|Q8L434 Hypothetical protein.
Length = 192
Score = 73.2 bits (178), Expect = 3e-12
Identities = 52/146 (35%), Positives = 69/146 (47%), Gaps = 14/146 (9%)
Frame = -2
Query: 685 YFHDVVAGTNPTAVRIAQX-------------AXXXXXXXNFGAVVAIDDPLTTGPTRAA 545
Y HD G NPTA I Q + FG + +++ LT GP R +
Sbjct: 42 YMHDQYTGENPTAALIMQGRPPLSNSNATTTGSSDDDDPRRFGDIAVMNNALTEGPERGS 101
Query: 544 GTEVGRAQG-TYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGG 368
VG AQG T A++ + L M+ V AGEH G +L + GR + VRE IVGG
Sbjct: 102 A-RVGTAQGFTVRVAERGSVNALS-MHLVMEAGEHAGISLVVSGRVDTDLAVRESVIVGG 159
Query: 367 SGKFRMARGYVQAHTIDSGATSGTTV 290
+G+FR ARGY + + D G V
Sbjct: 160 TGRFRAARGYALSRSYDYDLAKGGVV 185
>sptr|Q8S5Q4|Q8S5Q4 Putative disease resistance response protein.
Length = 188
Score = 71.6 bits (174), Expect = 1e-11
Identities = 54/152 (35%), Positives = 74/152 (48%), Gaps = 6/152 (3%)
Frame = -2
Query: 706 GLTTSEVYFHDV-VAGTNPTAVRIAQXAXXXXXXXNFGAVVAIDDPLTTGPTRAAGTEVG 530
G+T +FH+V AG N T +A A G V +DD L G A+ VG
Sbjct: 35 GMTHLHFFFHEVFTAGPNGTTATVAPPARSGDGSS-LGFVGVVDDMLREGADPASRL-VG 92
Query: 529 RAQG----TYTFADQQTFGLLMVMNFVFTA-GEHNGSTLSILGRNEVLSDVREMSIVGGS 365
RAQG T A + +++ FT G + GSTL + GR VL V E +VGG+
Sbjct: 93 RAQGVTAGTSLAAADGAGAITTMLSLAFTEEGPYAGSTLQVFGR-AVLGTVMERPVVGGT 151
Query: 364 GKFRMARGYVQAHTIDSGATSGTTVVQYTVNV 269
GKFRMARG+ + ++S V++Y V
Sbjct: 152 GKFRMARGHTLSRRVNSSDPDNLLVIEYDAYV 183
>sptr|Q9SDQ8|Q9SDQ8 Dirigent-like protein.
Length = 192
Score = 69.3 bits (168), Expect = 5e-11
Identities = 48/148 (32%), Positives = 71/148 (47%), Gaps = 8/148 (5%)
Frame = -2
Query: 688 VYFHDVV------AGTNPTAVRIAQXAXXXXXXXN--FGAVVAIDDPLTTGPTRAAGTEV 533
+YFHD++ T V Q A N FG + DDP+T V
Sbjct: 44 LYFHDIIYNGKNAGNATSTLVAAPQGANLTIMTGNYHFGDLAVFDDPITVD-NNLHSPPV 102
Query: 532 GRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFR 353
GRAQG Y + + TF + FV + ++ G T++ G + +L+ R++S+VGG+G F
Sbjct: 103 GRAQGFYFYDMKNTFSAWLGFTFVLNSTDYKG-TITFGGADPILAKYRDISVVGGTGDFL 161
Query: 352 MARGYVQAHTIDSGATSGTTVVQYTVNV 269
MARG TID+ A G + VN+
Sbjct: 162 MARGIA---TIDTDAYEGDVYFRLRVNI 186
>sptr|Q9SDQ9|Q9SDQ9 Dirigent protein.
Length = 192
Score = 68.6 bits (166), Expect = 8e-11
Identities = 48/148 (32%), Positives = 71/148 (47%), Gaps = 8/148 (5%)
Frame = -2
Query: 688 VYFHDVV------AGTNPTAVRIAQXAXXXXXXXN--FGAVVAIDDPLTTGPTRAAGTEV 533
+YFHD++ T V Q A N FG + DDP+T V
Sbjct: 44 LYFHDIIYNGKNAGNATSTLVAAPQGANLTIMTGNYHFGDLSVFDDPITVD-NNLHSPPV 102
Query: 532 GRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFR 353
GRAQG Y + + TF + FV + ++ G T++ G + +L+ R++S+VGG+G F
Sbjct: 103 GRAQGFYFYDMKNTFSAWLGFTFVLNSTDYKG-TITFGGADPILAKYRDISVVGGTGDFL 161
Query: 352 MARGYVQAHTIDSGATSGTTVVQYTVNV 269
MARG TID+ A G + VN+
Sbjct: 162 MARGIA---TIDTDAYEGDVYFRLRVNI 186
>sptr|Q9SDR0|Q9SDR0 Dirigent-like protein.
Length = 190
Score = 67.4 bits (163), Expect = 2e-10
Identities = 45/148 (30%), Positives = 70/148 (47%), Gaps = 8/148 (5%)
Frame = -2
Query: 688 VYFHDVVAG----TNPTAVRIAQXAXXXXXXXN----FGAVVAIDDPLTTGPTRAAGTEV 533
+YFHD++ N T+ +A FG + DDP+T V
Sbjct: 42 LYFHDIIYNGKNAENATSALVAAPEGANLTIMTGNNHFGNLAVFDDPITLD-NNLHSPPV 100
Query: 532 GRAQGTYTFADQQTFGLLMVMNFVFTAGEHNGSTLSILGRNEVLSDVREMSIVGGSGKFR 353
GRAQG Y + + TF + FV + +H G T++ G + +L+ R++S+VGG+G F
Sbjct: 101 GRAQGFYFYDMKNTFSAWLGFTFVLNSTDHKG-TITFNGADPILTKYRDISVVGGTGDFL 159
Query: 352 MARGYVQAHTIDSGATSGTTVVQYTVNV 269
MARG TI + + G + VN+
Sbjct: 160 MARGIA---TISTDSYEGDVYFRLRVNI 184
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 411,422,825
Number of Sequences: 1395590
Number of extensions: 7496316
Number of successful extensions: 21789
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 20941
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 21700
length of database: 442,889,342
effective HSP length: 119
effective length of database: 276,814,132
effective search space used: 32110439312
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

