BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
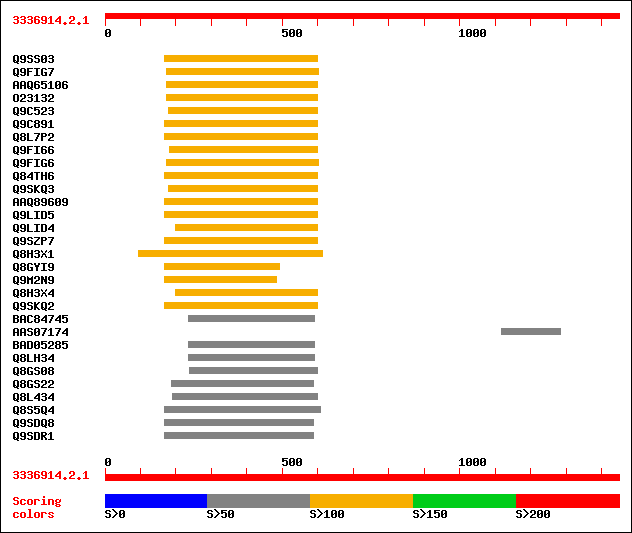
Query= 3336914.2.1
(1448 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9SS03|Q9SS03 F12P19.3. 139 1e-31
sptr|Q9FIG7|Q9FIG7 Disease resistance response protein-like (Hyp... 129 1e-28
sptrnew|AAQ65106|AAQ65106 At1g22900. 128 2e-28
sptr|O23132|O23132 F19G10.14 protein. 128 2e-28
sptr|Q9C523|Q9C523 Dirigent protein, putative (Hypothetical prot... 125 1e-27
sptr|Q9C891|Q9C891 Hypothetical protein. 123 6e-27
sptr|Q8L7P2|Q8L7P2 Hypothetical protein. 122 1e-26
sptr|Q9FI66|Q9FI66 Similarity to disease resistance response pro... 122 1e-26
sptr|Q9FIG6|Q9FIG6 Similarity to disease resistance response pro... 120 4e-26
sptr|Q84TH6|Q84TH6 Putative disease resistance response protein/... 119 9e-26
sptr|Q9SKQ3|Q9SKQ3 Putative disease resistance response protein. 117 6e-25
sptrnew|AAQ89609|AAQ89609 At3g13650. 117 6e-25
sptr|Q9LID5|Q9LID5 Disease resistance response protein-like (Put... 117 6e-25
sptr|Q9LID4|Q9LID4 Similarity to disease resistance response pro... 115 1e-24
sptr|Q9SZP7|Q9SZP7 Disease resistance response like protein. 112 1e-23
sptr|Q8H3X1|Q8H3X1 P0585H11.8 protein. 110 4e-23
sptr|Q8GYI9|Q8GYI9 Hypothetical protein. 109 1e-22
sptr|Q9M2N9|Q9M2N9 Hypothetical protein. 108 2e-22
sptr|Q8H3X4|Q8H3X4 P0585H11.3 protein. 106 8e-22
sptr|Q9SKQ2|Q9SKQ2 Putative disease resistance response protein. 104 3e-21
sptrnew|BAC84745|BAC84745 Disease resistance response protein-li... 90 8e-17
sptrnew|AAS07174|AAS07174 Expressed protein. 88 4e-16
sptrnew|BAD05285|BAD05285 Disease resistance response protein-like. 87 5e-16
sptr|Q8LH34|Q8LH34 P0617C02.32 protein (B1317D11.1 protein). 80 8e-14
sptr|Q8GS08|Q8GS08 P0552F09.12 protein (B1056G08.30 protein). 79 2e-13
sptr|Q8GS22|Q8GS22 P0552F09.4 protein (B1056G08.22 protein). 77 9e-13
sptr|Q8L434|Q8L434 Hypothetical protein. 77 9e-13
sptr|Q8S5Q4|Q8S5Q4 Putative disease resistance response protein. 74 8e-12
sptr|Q9SDQ8|Q9SDQ8 Dirigent-like protein. 73 1e-11
sptr|Q9SDR1|Q9SDR1 Dirigent-like protein. 73 1e-11
>sptr|Q9SS03|Q9SS03 F12P19.3.
Length = 189
Score = 139 bits (350), Expect = 1e-31
Identities = 69/144 (47%), Positives = 91/144 (63%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +FHDIV+G PT++++A G +N+S+T FG + +DD LT GP EVGRAQ
Sbjct: 45 THLHFYFHDIVSGDKPTSVQVANGPTTNSSATGFGLVAVVDDKLTVGP-EITSEEVGRAQ 103
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y ADQ GLLM + VFT G+ ST+++ GR VLS VREM I+GG+G FR RG
Sbjct: 104 GMYASADQNKLGLLMAFNLVFTKGKFSDSTVAMYGRNPVLSKVREMPIIGGTGAFRFGRG 163
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
Y A T+ SG+ VV+Y V +
Sbjct: 164 YALAKTLVFNITSGDAVVEYNVYI 187
>sptr|Q9FIG7|Q9FIG7 Disease resistance response protein-like
(Hypothetical protein) (At5g42500).
Length = 185
Score = 129 bits (324), Expect = 1e-28
Identities = 64/143 (44%), Positives = 90/143 (62%)
Frame = -1
Query: 602 FTTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRA 423
FT +FHD+++G PTA+++A+ +N+S+ +FG ++ DDPLT GP ++ EVGRA
Sbjct: 42 FTHLHFYFHDVISGDKPTAVKVAEARRTNSSNVNFGVIMIADDPLTEGPDPSS-KEVGRA 100
Query: 422 QGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMAR 243
QG Y + MV + FTAGE GST+++ GR E+ S VREM I+GG+G FR AR
Sbjct: 101 QGMYALTAMKNISFTMVFNLAFTAGEFNGSTVAMYGRNEIFSKVREMPIIGGTGAFRFAR 160
Query: 242 GYVQAHTIDSGFKSGETVVQYTV 174
GY QA T + VV+Y V
Sbjct: 161 GYAQAKTYK--VVGLDAVVEYNV 181
>sptrnew|AAQ65106|AAQ65106 At1g22900.
Length = 187
Score = 128 bits (322), Expect = 2e-28
Identities = 65/142 (45%), Positives = 90/142 (63%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +FHDI++G PT IR+A+ +N+S+T FGA++ +D P+T GP + EVGRAQ
Sbjct: 45 THLHFYFHDIISGDKPTTIRVAEAPGTNSSATVFGAVLIVDAPVTEGP-ELSSKEVGRAQ 103
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y D +TFG MV +FVFT GE GST ++ GR +L + RE+ I+GG+G FR ARG
Sbjct: 104 GLYASTDMKTFGFTMVFNFVFTEGEFNGSTAALYGRNPILLEERELPIIGGTGDFRFARG 163
Query: 239 YVQAHTIDSGFKSGETVVQYTV 174
Y T + + VV+Y V
Sbjct: 164 YALPKTYK--VVNIDAVVEYNV 183
>sptr|O23132|O23132 F19G10.14 protein.
Length = 187
Score = 128 bits (322), Expect = 2e-28
Identities = 65/142 (45%), Positives = 90/142 (63%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +FHDI++G PT IR+A+ +N+S+T FGA++ +D P+T GP + EVGRAQ
Sbjct: 45 THLHFYFHDIISGDKPTTIRVAEAPGTNSSATVFGAVLIVDAPVTEGP-ELSSKEVGRAQ 103
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y D +TFG MV +FVFT GE GST ++ GR +L + RE+ I+GG+G FR ARG
Sbjct: 104 GLYASTDMKTFGFTMVFNFVFTEGEFNGSTAALYGRNPILLEERELPIIGGTGDFRFARG 163
Query: 239 YVQAHTIDSGFKSGETVVQYTV 174
Y T + + VV+Y V
Sbjct: 164 YALPKTYK--VVNIDAVVEYNV 183
>sptr|Q9C523|Q9C523 Dirigent protein, putative (Hypothetical
protein) (At1g58170).
Length = 185
Score = 125 bits (315), Expect = 1e-27
Identities = 63/140 (45%), Positives = 87/140 (62%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T F+V++HDIV G +++ I T +T FG M ID+PLT P + + VGRAQ
Sbjct: 40 THFRVYWHDIVTGQDSSSVSIMNPPKKYTGATGFGLMRMIDNPLTLTP-KLSSKMVGRAQ 98
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y ++ GLLM M+F G++ GST+++LGR V VREM ++GGSG FR ARG
Sbjct: 99 GFYAGTSKEEIGLLMAMNFAILDGKYNGSTITVLGRNSVFDKVREMPVIGGSGLFRFARG 158
Query: 239 YVQAHTIDSGFKSGETVVQY 180
YVQA T + K+G +V+Y
Sbjct: 159 YVQASTHEFNLKTGNAIVEY 178
>sptr|Q9C891|Q9C891 Hypothetical protein.
Length = 187
Score = 123 bits (309), Expect = 6e-27
Identities = 69/144 (47%), Positives = 93/144 (64%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T FKV++HDI++G +PT+I I T+S+ FGA+ ID+ LT T +G+AQ
Sbjct: 45 THFKVYWHDILSGPNPTSIMIQPPV---TNSSYFGAISMIDNALTA-KVPMNSTVLGQAQ 100
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y A Q+ G LM M+F F G++ GST++ILGR LS+VREM IVGGSG FR ARG
Sbjct: 101 GFYAGAAQKELGFLMAMNFAFKTGKYNGSTITILGRNTALSEVREMPIVGGSGLFRFARG 160
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
YV+A T K+G+ V+Y+ V
Sbjct: 161 YVEARTKWINLKNGDATVEYSCYV 184
>sptr|Q8L7P2|Q8L7P2 Hypothetical protein.
Length = 187
Score = 122 bits (307), Expect = 1e-26
Identities = 69/144 (47%), Positives = 93/144 (64%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T FKV++HDI++G +PT+I I T+S+ FGA+ ID+ LT T +G+AQ
Sbjct: 45 THFKVYWHDILSGPNPTSIMIQPPF---TNSSYFGAISMIDNALTA-KVPMNSTVLGQAQ 100
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y A Q+ G LM M+F F G++ GST++ILGR LS+VREM IVGGSG FR ARG
Sbjct: 101 GFYAGAAQKELGFLMAMNFAFKTGKYNGSTITILGRNTALSEVREMPIVGGSGLFRFARG 160
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
YV+A T K+G+ V+Y+ V
Sbjct: 161 YVEARTKWINLKNGDATVEYSCYV 184
>sptr|Q9FI66|Q9FI66 Similarity to disease resistance response
protein.
Length = 191
Score = 122 bits (307), Expect = 1e-26
Identities = 66/139 (47%), Positives = 90/139 (64%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +V++HDIV G +P++IRI A +SS+ FG++ ID+ LT T VG+AQ
Sbjct: 46 THLRVYWHDIVTGRNPSSIRIQGPVAKYSSSSYFGSITMIDNALTLD-VPINSTVVGQAQ 104
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y A Q+ GLLM M+ F G++ GST++ILGR V+S VREM +VGGSG FR ARG
Sbjct: 105 GMYVGAAQKEIGLLMAMNLAFKTGKYNGSTITILGRNTVMSKVREMPVVGGSGMFRFARG 164
Query: 239 YVQAHTIDSGFKSGETVVQ 183
YV+A T K+G+ V+
Sbjct: 165 YVEARTKLFDMKTGDATVE 183
>sptr|Q9FIG6|Q9FIG6 Similarity to disease resistance response
protein.
Length = 182
Score = 120 bits (302), Expect = 4e-26
Identities = 60/143 (41%), Positives = 87/143 (60%)
Frame = -1
Query: 602 FTTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRA 423
FT +FHD+++G PTA+++A+ + T + FG ++ DDPLT GP ++ EVGRA
Sbjct: 39 FTHLHFYFHDVISGDKPTAVKVAEARPTTTLNVKFGVIMIADDPLTEGPDPSS-KEVGRA 97
Query: 422 QGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMAR 243
QG Y + MV ++VFTAGE GST+++ GR ++ S VRE+ I+GG+G FR AR
Sbjct: 98 QGMYASTAMKDIVFTMVFNYVFTAGEFNGSTIAVYGRNDIFSKVRELPIIGGTGAFRFAR 157
Query: 242 GYVQAHTIDSGFKSGETVVQYTV 174
GY T + VV+Y V
Sbjct: 158 GYALPKTYK--IVGLDAVVEYNV 178
>sptr|Q84TH6|Q84TH6 Putative disease resistance response
protein/dirigent protein.
Length = 187
Score = 119 bits (299), Expect = 9e-26
Identities = 61/144 (42%), Positives = 90/144 (62%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T + +FHD ++G +PTA+++AQG + S T FGA+ +DD LT + VGRAQ
Sbjct: 43 TNLQFYFHDTLSGKNPTAVKVAQGTDTEKSPTLFGAVFMVDDALTETADPKSKL-VGRAQ 101
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y + ++ GL+M M F F G +K ST+S++G+ ++ +REM IVGG+G FRMARG
Sbjct: 102 GLYGSSCKEEVGLIMAMSFCFEDGPYKDSTISMIGKNSAMNPIREMPIVGGTGMFRMARG 161
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
Y A T K+G+ +V Y V +
Sbjct: 162 YAIARTNWFDPKTGDAIVGYNVTI 185
>sptr|Q9SKQ3|Q9SKQ3 Putative disease resistance response protein.
Length = 276
Score = 117 bits (292), Expect = 6e-25
Identities = 60/140 (42%), Positives = 88/140 (62%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T + +FHD ++G +PTA+++AQG + S T FGA+ +DD LT + VGRAQ
Sbjct: 43 TNLQFYFHDTLSGKNPTAVKVAQGTDTEKSPTLFGAVFMVDDALTETADPKSKL-VGRAQ 101
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y + ++ GL+M M F F G +K ST+S++G+ ++ +REM IVGG+G FRMARG
Sbjct: 102 GLYGSSCKEEVGLIMAMSFCFEDGPYKDSTISMIGKNSAMNPIREMPIVGGTGMFRMARG 161
Query: 239 YVQAHTIDSGFKSGETVVQY 180
Y A T K+G+ +V Y
Sbjct: 162 YAIARTNWFDPKTGDAIVGY 181
>sptrnew|AAQ89609|AAQ89609 At3g13650.
Length = 186
Score = 117 bits (292), Expect = 6e-25
Identities = 63/144 (43%), Positives = 95/144 (65%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T F+V++HDI++G++P+++ I ++ S+ FG++ ID+ LTT T VG+AQ
Sbjct: 44 THFRVYWHDILSGSNPSSVVINPPISN---SSFFGSVTVIDNRLTT-EVAVNSTLVGQAQ 99
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y Q+ LMVM+F F G++ GS+++ILGR VL+ VREM ++GGSG FR ARG
Sbjct: 100 GIYAATGQRDASALMVMNFAFKTGKYNGSSIAILGRNAVLTKVREMPVIGGSGLFRFARG 159
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
YV+A T+ KSG+ V+Y+ V
Sbjct: 160 YVEARTMWFDQKSGDATVEYSCYV 183
>sptr|Q9LID5|Q9LID5 Disease resistance response protein-like
(Putative dirigent protein).
Length = 186
Score = 117 bits (292), Expect = 6e-25
Identities = 63/144 (43%), Positives = 95/144 (65%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T F+V++HDI++G++P+++ I ++ S+ FG++ ID+ LTT T VG+AQ
Sbjct: 44 THFRVYWHDILSGSNPSSVVINPPISN---SSFFGSVTVIDNRLTT-EVAVNSTLVGQAQ 99
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y Q+ LMVM+F F G++ GS+++ILGR VL+ VREM ++GGSG FR ARG
Sbjct: 100 GIYAATGQRDASALMVMNFAFKTGKYNGSSIAILGRNAVLTKVREMPVIGGSGLFRFARG 159
Query: 239 YVQAHTIDSGFKSGETVVQYTVNV 168
YV+A T+ KSG+ V+Y+ V
Sbjct: 160 YVEARTMWFDQKSGDATVEYSCYV 183
>sptr|Q9LID4|Q9LID4 Similarity to disease resistance response
protein.
Length = 186
Score = 115 bits (289), Expect = 1e-24
Identities = 64/134 (47%), Positives = 88/134 (65%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +V++H+ V G +P+++ I Q +++ S G++ +DDPLT R A T VG+AQ
Sbjct: 43 THLRVYWHNSVNGRNPSSVMIQQPVLNSSLS---GSITMMDDPLTFDVPRNA-TVVGQAQ 98
Query: 419 GTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARG 240
G Y A Q G LMVM+F FT G++ GST++ILGR V+S VREM +VGGSG FR ARG
Sbjct: 99 GMYVAAAQGEIGFLMVMNFAFTTGKYNGSTITILGRNVVMSKVREMPVVGGSGIFRFARG 158
Query: 239 YVQAHTIDSGFKSG 198
YV+A T K+G
Sbjct: 159 YVEARTKSFDLKAG 172
>sptr|Q9SZP7|Q9SZP7 Disease resistance response like protein.
Length = 176
Score = 112 bits (281), Expect = 1e-23
Identities = 62/150 (41%), Positives = 88/150 (58%), Gaps = 6/150 (4%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQG--AASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGR 426
T + + HD ++G +PTA+RIA + S FG++ IDDPLT GP + + E+G
Sbjct: 25 TRVRFYLHDTLSGQNPTAVRIAHANLTGGSASPVGFGSLFVIDDPLTVGPEKHS-KEIGN 83
Query: 425 AQGTYTFA--DQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDV--REMSIVGGSGK 258
QG Y D F ++M D FTAG+ GS++SI R V +V RE++IVGG GK
Sbjct: 84 GQGMYVSGCKDLSKFTIVMYADLAFTAGKFNGSSISIFSRNPVAEEVGEREIAIVGGRGK 143
Query: 257 FRMARGYVQAHTIDSGFKSGETVVQYTVNV 168
FRMARG+V+ T K+G+ V++Y V
Sbjct: 144 FRMARGFVKVKTNKIDMKTGDAVLRYDATV 173
>sptr|Q8H3X1|Q8H3X1 P0585H11.8 protein.
Length = 220
Score = 110 bits (276), Expect = 4e-23
Identities = 69/178 (38%), Positives = 96/178 (53%), Gaps = 4/178 (2%)
Frame = -1
Query: 614 DDSGFTT-FKVFFHDIVAGTSPTAIRIAQG---AASNTSSTSFGAMVAIDDPLTTGPTRA 447
+D G T +F HD++ G TA+ + G A FG +V +DD LT GP+R+
Sbjct: 44 EDRGVATHLHLFMHDVLTGPDATAVDVVNGTGRAFDVAGGLRFGQVVVMDDVLTEGPSRS 103
Query: 446 AGTEVGRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGG 267
+ VGR QG Y F+D LL M+ V TAG + GST++ILGR + +RE+S+VGG
Sbjct: 104 S-PRVGRTQGFYVFSDMNVPALLFCMNVVLTAGPYAGSTVTILGRDHITQPLRELSVVGG 162
Query: 266 SGKFRMARGYVQAHTIDSGFKSGETVVQYTVNVRA*LPARSTLLAPPAHGRLPARLLV 93
+G FRMA GYV T F++ + V++ V V P PP H R P +V
Sbjct: 163 TGAFRMATGYVLWRTASWQFRA-DAVLELDVFVHT-RPEYLQSPPPPPHHRPPPPTVV 218
>sptr|Q8GYI9|Q8GYI9 Hypothetical protein.
Length = 110
Score = 109 bits (272), Expect = 1e-22
Identities = 56/108 (51%), Positives = 67/108 (62%)
Frame = -1
Query: 491 MVAIDDPLTTGPTRAAGTEVGRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGR 312
+ +DD LT GP EVGRAQG + ADQ FGLLM +FVFT GE GST+S+ GR
Sbjct: 2 VAVVDDILTVGP-EITSEEVGRAQGIFASADQNNFGLLMAFNFVFTKGEFSGSTVSMYGR 60
Query: 311 XEVLSDVREMSIVGGSGKFRMARGYVQAHTIDSGFKSGETVVQYTVNV 168
+ S VREM I+GG+G FR RGY QA T SG VV+Y V +
Sbjct: 61 NPIFSKVREMPIIGGTGAFRFGRGYAQAKTFTFNTTSGNAVVKYNVYI 108
>sptr|Q9M2N9|Q9M2N9 Hypothetical protein.
Length = 249
Score = 108 bits (271), Expect = 2e-22
Identities = 56/105 (53%), Positives = 66/105 (62%)
Frame = -1
Query: 482 IDDPLTTGPTRAAGTEVGRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEV 303
+DD LT GP EVGRAQG + ADQ FGLLM +FVFT GE GST+S+ GR +
Sbjct: 144 VDDILTVGP-EITSEEVGRAQGIFASADQNNFGLLMAFNFVFTKGEFSGSTVSMYGRNPI 202
Query: 302 LSDVREMSIVGGSGKFRMARGYVQAHTIDSGFKSGETVVQYTVNV 168
S VREM I+GG+G FR RGY QA T SG VV+Y V +
Sbjct: 203 FSKVREMPIIGGTGAFRFGRGYAQAKTFTFNTTSGNAVVKYNVYI 247
>sptr|Q8H3X4|Q8H3X4 P0585H11.3 protein.
Length = 229
Score = 106 bits (265), Expect = 8e-22
Identities = 56/136 (41%), Positives = 83/136 (61%), Gaps = 2/136 (1%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQG--AASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGR 426
T F HD+V+G+ TA+++ +G +A+ ST FG +DD LT + A EVGR
Sbjct: 30 THLHFFMHDVVSGSGQTAVQVIKGPTSANGGVSTGFGDTTVVDDALTE-TSSATSPEVGR 88
Query: 425 AQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMA 246
AQG Y + + L+M ++ FTAGE+ GST++++G + + VRE+S+VGG+GKFRMA
Sbjct: 89 AQGFYMMSSLSSPTLMMCVNLYFTAGENNGSTIAVIGHDDTTATVRELSVVGGTGKFRMA 148
Query: 245 RGYVQAHTIDSGFKSG 198
GYV T +G
Sbjct: 149 TGYVVWKTASMSASTG 164
>sptr|Q9SKQ2|Q9SKQ2 Putative disease resistance response protein.
Length = 186
Score = 104 bits (260), Expect = 3e-21
Identities = 56/148 (37%), Positives = 87/148 (58%), Gaps = 4/148 (2%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGA---ASNTSSTS-FGAMVAIDDPLTTGPTRAAGTEV 432
T FFHD + +P+AI IA+ N SS S FG++ A+DDPLT GP + ++
Sbjct: 37 TNLHFFFHDTLTAPNPSAILIAKPTHTRGDNDSSPSPFGSLFALDDPLTVGPDPKS-EKI 95
Query: 431 GRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFR 252
G A+G Y + + L M +DF FT+G+ GS++++ R + RE+++VGG G+FR
Sbjct: 96 GNARGMYVSSGKHVPTLTMYVDFGFTSGKFNGSSIAVFSRNTITEKEREVAVVGGRGRFR 155
Query: 251 MARGYVQAHTIDSGFKSGETVVQYTVNV 168
MARG Q +T +G+ +V+Y V +
Sbjct: 156 MARGVAQLNTYYVNLTNGDAIVEYNVTL 183
>sptrnew|BAC84745|BAC84745 Disease resistance response protein-like
protein.
Length = 209
Score = 90.1 bits (222), Expect = 8e-17
Identities = 51/123 (41%), Positives = 71/123 (57%), Gaps = 4/123 (3%)
Frame = -1
Query: 590 KVFFHDIVAGTSPTAIRIAQGAASNTSSTS----FGAMVAIDDPLTTGPTRAAGTEVGRA 423
+V+ HDIV G T++ + +G S S FG V IDD +T GP+ A+ EVGRA
Sbjct: 41 RVYMHDIVGGAGQTSVVVVKGPGPANPSMSPGNNFGDTVIIDDVVTEGPSLAS-REVGRA 99
Query: 422 QGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMAR 243
QGTY A ++ + V T G + GST+ I GR + +VRE+++VGGSG R A
Sbjct: 100 QGTYMLASMARPVFIVDITVVLTDGPYNGSTILIAGRDDTSEEVRELAVVGGSGMLRRAS 159
Query: 242 GYV 234
G+V
Sbjct: 160 GHV 162
>sptrnew|AAS07174|AAS07174 Expressed protein.
Length = 92
Score = 87.8 bits (216), Expect = 4e-16
Identities = 45/56 (80%), Positives = 49/56 (87%)
Frame = -3
Query: 1284 KGDREAAIDAXXVRVNDLPAHSSYAIXRMKVLNKLRHLLSIKRTTSQDEELELLFA 1117
+GDRE A+ A RVN LPA+SSYAI R+KVLNKLRHLLSIKRTTSQDEELELLFA
Sbjct: 33 QGDRENAVSAEIERVNKLPANSSYAIHRLKVLNKLRHLLSIKRTTSQDEELELLFA 88
>sptrnew|BAD05285|BAD05285 Disease resistance response protein-like.
Length = 207
Score = 87.4 bits (215), Expect = 5e-16
Identities = 50/124 (40%), Positives = 74/124 (59%), Gaps = 5/124 (4%)
Frame = -1
Query: 590 KVFFHDIVAGTSPTAIRIAQGAASNTSST---SFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
+++ HD++ G TAIR+ GA +S +FG VA+DD +T GP+ A+ VGRAQ
Sbjct: 50 RLYMHDVIDGPGQTAIRLIWGAGPPHASMPGRAFGDTVAVDDLVTEGPSIASAA-VGRAQ 108
Query: 419 GTYTFADQQTFGLLMVMDFVFTA--GEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMA 246
GTY + Q+ L++ + T+ G + GS L I GR V + RE+++VGG+G R A
Sbjct: 109 GTYMLSSQREAVLVVAITVALTSAGGPYNGSNLVIAGRDRVRDETRELAVVGGTGALRGA 168
Query: 245 RGYV 234
GYV
Sbjct: 169 AGYV 172
>sptr|Q8LH34|Q8LH34 P0617C02.32 protein (B1317D11.1 protein).
Length = 204
Score = 80.1 bits (196), Expect = 8e-14
Identities = 44/123 (35%), Positives = 69/123 (56%), Gaps = 4/123 (3%)
Frame = -1
Query: 590 KVFFHDIVAGTSPTAIRIAQGAA----SNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRA 423
+++ HDI G TA+++ G + + FG + +DD LT VGRA
Sbjct: 31 RLYMHDITGGPGQTAVQVVNGTGPLHPAMPPGSHFGDTMVVDDLLTDVDD---SKPVGRA 87
Query: 422 QGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMAR 243
QG+YT A + ++ + V T G +KGST+ I GR ++ +VRE+++VGG+GK R A
Sbjct: 88 QGSYTLACLRAPVFVVSITLVLTDGPYKGSTILIAGRDDISEEVRELAVVGGTGKLRRAT 147
Query: 242 GYV 234
G+V
Sbjct: 148 GHV 150
>sptr|Q8GS08|Q8GS08 P0552F09.12 protein (B1056G08.30 protein).
Length = 189
Score = 79.0 bits (193), Expect = 2e-13
Identities = 46/125 (36%), Positives = 65/125 (52%), Gaps = 4/125 (3%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQ 420
T +V+ H+ AG + TA+ + ++ +FG + +DD L GP RA+ +GR Q
Sbjct: 39 TRIRVYMHERFAGANATALAVVPSPLG--ANEAFGRVAVLDDELRDGPDRASSALIGRFQ 96
Query: 419 G----TYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFR 252
G T + VFTAGEH GSTLS++G + E +VGG+G FR
Sbjct: 97 GVVAGTSLPGTAPPASFQSAISLVFTAGEHAGSTLSMVGPVLGFAGAIERPLVGGTGAFR 156
Query: 251 MARGY 237
MARGY
Sbjct: 157 MARGY 161
>sptr|Q8GS22|Q8GS22 P0552F09.4 protein (B1056G08.22 protein).
Length = 175
Score = 76.6 bits (187), Expect = 9e-13
Identities = 51/134 (38%), Positives = 68/134 (50%)
Frame = -1
Query: 587 VFFHDIVAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEVGRAQGTYT 408
++ H+ +G + T G ++ +T+FG + D+ L TG RAA VGR QG
Sbjct: 34 LYMHETFSGPNATE----GGVVASPFNTTFGQVAVFDNELRTGEDRAASPLVGRYQGFIV 89
Query: 407 FADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFRMARGYVQA 228
+ + G L FTAGE GSTLS+ G + E SIVGG+GK R+ARGY
Sbjct: 90 GTGRSSPGYLTSATVAFTAGELNGSTLSLEGPFFGFAGTAERSIVGGTGKLRLARGYYLL 149
Query: 227 HTIDSGFKSGETVV 186
I G S ET V
Sbjct: 150 KLI--GKTSPETAV 161
>sptr|Q8L434|Q8L434 Hypothetical protein.
Length = 192
Score = 76.6 bits (187), Expect = 9e-13
Identities = 55/151 (36%), Positives = 76/151 (50%), Gaps = 14/151 (9%)
Frame = -1
Query: 599 TTFKVFFHDIVAGTSPTAIRIAQGAA--SNTSSTS-----------FGAMVAIDDPLTTG 459
T + HD G +PTA I QG SN+++T+ FG + +++ LT G
Sbjct: 37 THLHFYMHDQYTGENPTAALIMQGRPPLSNSNATTTGSSDDDDPRRFGDIAVMNNALTEG 96
Query: 458 PTRAAGTEVGRAQG-TYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREM 282
P R + VG AQG T A++ + L M V AGEH G +L + GR + VRE
Sbjct: 97 PERGSA-RVGTAQGFTVRVAERGSVNALS-MHLVMEAGEHAGISLVVSGRVDTDLAVRES 154
Query: 281 SIVGGSGKFRMARGYVQAHTIDSGFKSGETV 189
IVGG+G+FR ARGY + + D G V
Sbjct: 155 VIVGGTGRFRAARGYALSRSYDYDLAKGGVV 185
>sptr|Q8S5Q4|Q8S5Q4 Putative disease resistance response protein.
Length = 188
Score = 73.6 bits (179), Expect = 8e-12
Identities = 55/153 (35%), Positives = 76/153 (49%), Gaps = 6/153 (3%)
Frame = -1
Query: 608 SGFTTFKVFFHDI-VAGTSPTAIRIAQGAASNTSSTSFGAMVAIDDPLTTGPTRAAGTEV 432
+G T FFH++ AG + T +A A S S S G + +DD L G A+ V
Sbjct: 34 AGMTHLHFFFHEVFTAGPNGTTATVAPPARSGDGS-SLGFVGVVDDMLREGADPASRL-V 91
Query: 431 GRAQG----TYTFADQQTFGLLMVMDFVFTA-GEHKGSTLSILGRXEVLSDVREMSIVGG 267
GRAQG T A + ++ FT G + GSTL + GR VL V E +VGG
Sbjct: 92 GRAQGVTAGTSLAAADGAGAITTMLSLAFTEEGPYAGSTLQVFGRA-VLGTVMERPVVGG 150
Query: 266 SGKFRMARGYVQAHTIDSGFKSGETVVQYTVNV 168
+GKFRMARG+ + ++S V++Y V
Sbjct: 151 TGKFRMARGHTLSRRVNSSDPDNLLVIEYDAYV 183
>sptr|Q9SDQ8|Q9SDQ8 Dirigent-like protein.
Length = 192
Score = 73.2 bits (178), Expect = 1e-11
Identities = 49/148 (33%), Positives = 76/148 (51%), Gaps = 8/148 (5%)
Frame = -1
Query: 587 VFFHDIV-----AGTSPTAIRIAQGAASNTSSTS---FGAMVAIDDPLTTGPTRAAGTEV 432
++FHDI+ AG + + + A A+ T T FG + DDP+T V
Sbjct: 44 LYFHDIIYNGKNAGNATSTLVAAPQGANLTIMTGNYHFGDLAVFDDPITVD-NNLHSPPV 102
Query: 431 GRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFR 252
GRAQG Y + + TF + FV + ++KG T++ G +L+ R++S+VGG+G F
Sbjct: 103 GRAQGFYFYDMKNTFSAWLGFTFVLNSTDYKG-TITFGGADPILAKYRDISVVGGTGDFL 161
Query: 251 MARGYVQAHTIDSGFKSGETVVQYTVNV 168
MARG TID+ G+ + VN+
Sbjct: 162 MARGIA---TIDTDAYEGDVYFRLRVNI 186
>sptr|Q9SDR1|Q9SDR1 Dirigent-like protein.
Length = 190
Score = 72.8 bits (177), Expect = 1e-11
Identities = 48/148 (32%), Positives = 74/148 (50%), Gaps = 8/148 (5%)
Frame = -1
Query: 587 VFFHDIVAG----TSPTAIRIAQGAASN----TSSTSFGAMVAIDDPLTTGPTRAAGTEV 432
++FHDI+ + T+ +A +N T + FG + DDP+T V
Sbjct: 42 LYFHDIIYNGKNAENATSALVAAPEGANLTIMTGNNHFGNLAVFDDPITLD-NNLHSPPV 100
Query: 431 GRAQGTYTFADQQTFGLLMVMDFVFTAGEHKGSTLSILGRXEVLSDVREMSIVGGSGKFR 252
GRAQG Y + + TF + FV + +HKG T++ G +L+ R++S+VGG+G F
Sbjct: 101 GRAQGFYFYDMKNTFSAWLGFTFVLNSTDHKG-TITFNGADPILTKYRDISVVGGTGDFL 159
Query: 251 MARGYVQAHTIDSGFKSGETVVQYTVNV 168
MARG TI + GE + VN+
Sbjct: 160 MARGIA---TISTDSYEGEVYFRLRVNI 184
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 849,589,839
Number of Sequences: 1395590
Number of extensions: 15204109
Number of successful extensions: 42843
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 40649
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 42734
length of database: 442,889,342
effective HSP length: 127
effective length of database: 265,649,412
effective search space used: 94305541260
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

