BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
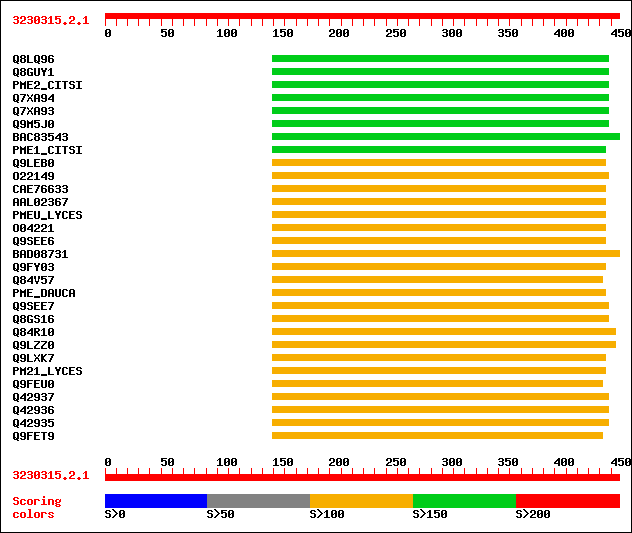
Query= 3230315.2.1
(457 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8LQ96|Q8LQ96 Putative pectinesterase 2.1. 192 1e-48
sptr|Q8GUY1|Q8GUY1 Methyl pectinesterase (Fragment). 186 7e-47
sw|O04887|PME2_CITSI Pectinesterase 2.1 precursor (EC 3.1.1.11) ... 161 2e-39
sptr|Q7XA94|Q7XA94 Pectinesterase (EC 3.1.1.11) (Fragment). 156 1e-37
sptr|Q7XA93|Q7XA93 Pectinesterase (EC 3.1.1.11) (Fragment). 155 1e-37
sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC 3.1.1... 154 4e-37
sptrnew|BAC83543|BAC83543 Putative pectinesterase. 152 1e-36
sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11) ... 151 3e-36
sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11) (Pectines... 150 4e-36
sptr|O22149|O22149 Putative pectinesterase (At2g45220/F4L23.27). 150 4e-36
sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11) (F... 150 4e-36
sptrnew|AAL02367|AAL02367 Pectin methylesterase (EC 3.1.1.11). 150 6e-36
sw|Q43143|PMEU_LYCES Pectinesterase U1 precursor (EC 3.1.1.11) (... 150 6e-36
sptr|O04221|O04221 Pectinesterase (Fragment). 150 8e-36
sptr|Q9SEE6|Q9SEE6 Pectin methyl esterase (EC 3.1.1.11) (Pectine... 149 1e-35
sptrnew|BAD08731|BAD08731 Putative pectinesterase. 149 1e-35
sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC ... 149 1e-35
sptr|Q84V57|Q84V57 Pectin methyl-esterase (EC 3.1.1.11). 149 2e-35
sw|P83218|PME_DAUCA Pectinesterase (EC 3.1.1.11) (Pectin methyle... 148 2e-35
sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11) (Pectine... 148 2e-35
sptr|Q8GS16|Q8GS16 Putative thermostable pectinesterase. 144 3e-34
sptr|Q84R10|Q84R10 Putative pectinesterase. 144 4e-34
sptr|Q9LZZ0|Q9LZZ0 Pectinesterase-like protein. 144 4e-34
sptr|Q9LXK7|Q9LXK7 Putative pectinesterase (EC 3.1.1.11) (Pectin... 143 7e-34
sw|P09607|PM21_LYCES Pectinesterase 2 precursor (EC 3.1.1.11) (P... 142 2e-33
sptr|Q9FEU0|Q9FEU0 Putative pectin methylesterase precursor. 142 2e-33
sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment). 142 2e-33
sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11) (Pectines... 142 2e-33
sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11) (Pectines... 142 2e-33
sptr|Q9FET9|Q9FET9 Putative pectin methylesterase (Fragment). 142 2e-33
>sptr|Q8LQ96|Q8LQ96 Putative pectinesterase 2.1.
Length = 295
Score = 192 bits (488), Expect = 1e-48
Identities = 86/100 (86%), Positives = 95/100 (95%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWKQYSRTV++QSELDS+V+PAGWLEW+G+FALDTLYYGEY NTGPGA TS RV
Sbjct: 196 KTYLGRPWKQYSRTVFMQSELDSVVNPAGWLEWSGNFALDTLYYGEYQNTGPGASTSNRV 255
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KWKGYRVITSA+EAS FTVGNFIDGD+WLAGTS+PFT GL
Sbjct: 256 KWKGYRVITSASEASTFTVGNFIDGDVWLAGTSVPFTVGL 295
>sptr|Q8GUY1|Q8GUY1 Methyl pectinesterase (Fragment).
Length = 226
Score = 186 bits (473), Expect = 7e-47
Identities = 84/100 (84%), Positives = 94/100 (94%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
+TYLGRPWKQYSRTVYLQSELDS+VDP GWLEW+G+FALDTLYYGEY NTG GA TS RV
Sbjct: 127 RTYLGRPWKQYSRTVYLQSELDSVVDPKGWLEWDGTFALDTLYYGEYQNTGAGASTSNRV 186
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KWKGYRVI+S++EAS FTVG+FIDGD+WLAGTSIPF+TGL
Sbjct: 187 KWKGYRVISSSSEASTFTVGSFIDGDVWLAGTSIPFSTGL 226
>sw|O04887|PME2_CITSI Pectinesterase 2.1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 510
Score = 161 bits (408), Expect = 2e-39
Identities = 68/100 (68%), Positives = 87/100 (87%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KT+LGRPWKQYSRTVY+++ LDSL++PAGW+EW+G FAL+TLYY EYMNTGPG+ T+ RV
Sbjct: 411 KTFLGRPWKQYSRTVYIKTFLDSLINPAGWMEWSGDFALNTLYYAEYMNTGPGSSTANRV 470
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW+GY V+TS ++ S FTVGNFI G+ WL T++PFT+GL
Sbjct: 471 KWRGYHVLTSPSQVSQFTVGNFIAGNSWLPATNVPFTSGL 510
>sptr|Q7XA94|Q7XA94 Pectinesterase (EC 3.1.1.11) (Fragment).
Length = 193
Score = 156 bits (394), Expect = 1e-37
Identities = 68/100 (68%), Positives = 78/100 (78%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
+TYLGRPW++YSR V ++S LD L+ P GW W+GSF LDTLYYGEYMNTG GA T GRV
Sbjct: 94 RTYLGRPWQKYSRVVIMKSNLDGLIAPQGWFPWSGSFGLDTLYYGEYMNTGAGAATGGRV 153
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW G+RVITSA EA FTVGNF+ GD WL GT +PF GL
Sbjct: 154 KWPGFRVITSATEAGKFTVGNFLAGDAWLPGTGVPFEAGL 193
>sptr|Q7XA93|Q7XA93 Pectinesterase (EC 3.1.1.11) (Fragment).
Length = 209
Score = 155 bits (393), Expect = 1e-37
Identities = 69/100 (69%), Positives = 79/100 (79%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWK+YSRTV +QS + ++DPAGW EW+G+FALDTL+Y EY NTG GA TS RV
Sbjct: 110 KTYLGRPWKEYSRTVIMQSSITDIIDPAGWYEWSGTFALDTLFYAEYANTGAGASTSNRV 169
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKGY+VITSA EA AFT GNFI G WL+ T PFT GL
Sbjct: 170 TWKGYKVITSATEAQAFTPGNFIAGGSWLSATGFPFTLGL 209
>sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC
3.1.1.11) (Fragment).
Length = 277
Score = 154 bits (389), Expect = 4e-37
Identities = 69/100 (69%), Positives = 80/100 (80%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
+ YLGRPWK YSRTV+L S ++ L++PAGWLEWNG+FALDTLYYGEY N GPGA TSGRV
Sbjct: 177 RNYLGRPWKMYSRTVFLNSLMEDLIEPAGWLEWNGTFALDTLYYGEYNNRGPGANTSGRV 236
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
W GYRVIT++ EAS FTV NFI G+ WL IPF +GL
Sbjct: 237 TWPGYRVITNSTEASQFTVQNFIQGNEWLNSYGIPFFSGL 276
>sptrnew|BAC83543|BAC83543 Putative pectinesterase.
Length = 579
Score = 152 bits (385), Expect = 1e-36
Identities = 68/103 (66%), Positives = 80/103 (77%)
Frame = -1
Query: 457 ARAKTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTS 278
A +YLGRPWK YSRTV+LQS++DSL+ P GWLEWNGSFALDTLYY EYMN G GA TS
Sbjct: 476 ANFSSYLGRPWKTYSRTVFLQSKIDSLIHPRGWLEWNGSFALDTLYYAEYMNRGDGADTS 535
Query: 277 GRVKWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
RV W GY V+T+A +A+ FTV NF+ GDLWL +S P+ GL
Sbjct: 536 ARVSWPGYHVLTNATDAANFTVLNFVQGDLWLNSSSFPYILGL 578
>sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 584
Score = 151 bits (382), Expect = 3e-36
Identities = 67/99 (67%), Positives = 80/99 (80%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + L+ PAGW EW+G+FAL+TL+YGE+ N+G GAGTSGRVK
Sbjct: 486 TYLGRPWKEYSRTVIMQSSITDLIHPAGWHEWDGNFALNTLFYGEHQNSGAGAGTSGRVK 545
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG+RVITSA EA AFT G+FI G WL T PF+ GL
Sbjct: 546 WKGFRVITSATEAQAFTPGSFIAGSSWLGSTGFPFSLGL 584
>sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase).
Length = 579
Score = 150 bits (380), Expect = 4e-36
Identities = 66/99 (66%), Positives = 80/99 (80%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + +++ AGW EWNG+FAL+TL+YGEY NTG GAGTSGRVK
Sbjct: 481 TYLGRPWKEYSRTVIMQSSITDVINSAGWHEWNGNFALNTLFYGEYQNTGAGAGTSGRVK 540
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITSA EA A+T G FI G WL+ T PF+ GL
Sbjct: 541 WKGFKVITSATEAQAYTPGRFIAGGSWLSSTGFPFSLGL 579
>sptr|O22149|O22149 Putative pectinesterase (At2g45220/F4L23.27).
Length = 511
Score = 150 bits (380), Expect = 4e-36
Identities = 66/100 (66%), Positives = 84/100 (84%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPW+QYSRTV++++ LDSL+DP GWLEW+G+FAL TL+Y E+ NTGPGA TSGRV
Sbjct: 413 KTYLGRPWRQYSRTVFMKTSLDSLIDPRGWLEWDGNFALKTLFYAEFQNTGPGASTSGRV 472
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
W G+RV+ SA+EAS FTVG F+ G W+ +S+PFT+GL
Sbjct: 473 TWPGFRVLGSASEASKFTVGTFLAGGSWIP-SSVPFTSGL 511
>sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11)
(Fragment).
Length = 426
Score = 150 bits (380), Expect = 4e-36
Identities = 66/99 (66%), Positives = 77/99 (77%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++DP GW EWNG+FAL+TL Y EY NTGPGAGTS RV
Sbjct: 328 TYLGRPWKEYSRTVIMQSSISDVIDPIGWHEWNGNFALNTLVYREYQNTGPGAGTSKRVN 387
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITSA+EA FT GNFI G WL T PF+ GL
Sbjct: 388 WKGFKVITSASEAQTFTPGNFIGGSTWLGSTGFPFSLGL 426
>sptrnew|AAL02367|AAL02367 Pectin methylesterase (EC 3.1.1.11).
Length = 583
Score = 150 bits (379), Expect = 6e-36
Identities = 66/99 (66%), Positives = 79/99 (79%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++ PAGW EWNG+FALDTL+YGEY NTG GA TSGRVK
Sbjct: 485 TYLGRPWKEYSRTVIMQSSITDVIQPAGWHEWNGNFALDTLFYGEYANTGAGAPTSGRVK 544
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITS+ EA A+T G FI G WL+ T PF+ GL
Sbjct: 545 WKGHKVITSSTEAQAYTPGRFIAGGSWLSSTGFPFSLGL 583
>sw|Q43143|PMEU_LYCES Pectinesterase U1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 583
Score = 150 bits (379), Expect = 6e-36
Identities = 66/99 (66%), Positives = 79/99 (79%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++ PAGW EWNG+FALDTL+YGEY NTG GA TSGRVK
Sbjct: 485 TYLGRPWKEYSRTVIMQSSITDVIQPAGWHEWNGNFALDTLFYGEYANTGAGAPTSGRVK 544
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITS+ EA A+T G FI G WL+ T PF+ GL
Sbjct: 545 WKGHKVITSSTEAQAYTPGRFIAGGSWLSSTGFPFSLGL 583
>sptr|O04221|O04221 Pectinesterase (Fragment).
Length = 290
Score = 150 bits (378), Expect = 8e-36
Identities = 66/99 (66%), Positives = 79/99 (79%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++ PAGW EW+G+FAL+TL+YGE+ N G GAGTSGRVK
Sbjct: 192 TYLGRPWKEYSRTVIMQSSITDVIHPAGWHEWDGNFALNTLFYGEHQNAGAGAGTSGRVK 251
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG+RVITSA EA AFT G+FI G WL T PF+ GL
Sbjct: 252 WKGFRVITSATEAQAFTPGSFIAGSSWLGSTGFPFSLGL 290
>sptr|Q9SEE6|Q9SEE6 Pectin methyl esterase (EC 3.1.1.11)
(Pectinesterase) (Pectin methylesterase).
Length = 576
Score = 149 bits (377), Expect = 1e-35
Identities = 65/99 (65%), Positives = 80/99 (80%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++ PAGW EWNG+FAL+TL+YGEY NTG GA TSGRVK
Sbjct: 478 TYLGRPWKEYSRTVIMQSSITDVIQPAGWHEWNGNFALNTLFYGEYANTGAGAATSGRVK 537
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITS+ EA A+T G+FI G WL+ T PF+ GL
Sbjct: 538 WKGHKVITSSTEAQAYTPGSFIAGGSWLSSTGFPFSLGL 576
>sptrnew|BAD08731|BAD08731 Putative pectinesterase.
Length = 557
Score = 149 bits (377), Expect = 1e-35
Identities = 65/103 (63%), Positives = 80/103 (77%)
Frame = -1
Query: 457 ARAKTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTS 278
A +TYLGRPWKQYSR V++QS + ++V P GWL W+G FALDTLYYGEYMNTGPGAG
Sbjct: 453 AVTQTYLGRPWKQYSRVVFMQSYIGAVVRPEGWLAWDGQFALDTLYYGEYMNTGPGAGVG 512
Query: 277 GRVKWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
GRVKW G+ V+TS A+A FTV FI+G++WL T + +T GL
Sbjct: 513 GRVKWPGFHVMTSPAQAGNFTVAQFIEGNMWLPPTGVKYTAGL 555
>sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC
3.1.1.11).
Length = 579
Score = 149 bits (376), Expect = 1e-35
Identities = 66/99 (66%), Positives = 77/99 (77%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++ PAGW EW+GSFAL TL+Y EY N+G GAGTS RVK
Sbjct: 481 TYLGRPWKEYSRTVIMQSSITDVIQPAGWHEWSGSFALSTLFYAEYQNSGAGAGTSSRVK 540
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
W+GY+VITSA EA AF GNFI G WL TS PF+ GL
Sbjct: 541 WEGYKVITSATEAQAFAPGNFIAGSSWLGSTSFPFSLGL 579
>sptr|Q84V57|Q84V57 Pectin methyl-esterase (EC 3.1.1.11).
Length = 579
Score = 149 bits (375), Expect = 2e-35
Identities = 65/98 (66%), Positives = 79/98 (80%)
Frame = -1
Query: 442 YLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVKW 263
YLGRPWK+YSRTV +QS + +++ AGW EWNG+FAL+TL+YGEY NTG GAGTSGRVKW
Sbjct: 482 YLGRPWKEYSRTVIMQSSITDVINSAGWHEWNGNFALNTLFYGEYQNTGAGAGTSGRVKW 541
Query: 262 KGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KG++VITSA EA A+T G FI G WL+ T PF+ GL
Sbjct: 542 KGFKVITSATEAQAYTPGRFIAGGSWLSSTGFPFSLGL 579
>sw|P83218|PME_DAUCA Pectinesterase (EC 3.1.1.11) (Pectin
methylesterase) (PE).
Length = 319
Score = 148 bits (374), Expect = 2e-35
Identities = 64/99 (64%), Positives = 79/99 (79%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV +QS + ++++PAGW W+G+FALDTLYYGEY NTG GA TSGRV
Sbjct: 221 TYLGRPWKEYSRTVVMQSSITNVINPAGWFPWDGNFALDTLYYGEYQNTGAGAATSGRVT 280
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
WKG++VITS+ EA FT G+FI G WL T+ PF+ GL
Sbjct: 281 WKGFKVITSSTEAQGFTPGSFIAGGSWLKATTFPFSLGL 319
>sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11)
(Pectinesterase) (Pectin methylesterase) (Fragment).
Length = 530
Score = 148 bits (374), Expect = 2e-35
Identities = 67/100 (67%), Positives = 74/100 (74%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWK+YSRTV +QS L L+DPAGW EW+G FAL TLYYGEYMN GPGAGTS RV
Sbjct: 429 KTYLGRPWKEYSRTVVMQSYLGGLIDPAGWAEWSGEFALKTLYYGEYMNNGPGAGTSKRV 488
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW GY VIT AEA FTV I G WL+ T + + GL
Sbjct: 489 KWPGYHVITDPAEAMPFTVAELIQGGSWLSSTGVAYVDGL 528
>sptr|Q8GS16|Q8GS16 Putative thermostable pectinesterase.
Length = 631
Score = 144 bits (364), Expect = 3e-34
Identities = 62/100 (62%), Positives = 77/100 (77%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
+TYLGRPWK+YSRTV +QS++ +++PAGW EW+G+FALDTL+Y EY NTG GA TS RV
Sbjct: 532 ETYLGRPWKRYSRTVVMQSDISDVINPAGWYEWSGNFALDTLFYAEYQNTGAGADTSNRV 591
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW ++VITSAAEA +T NFI G WL T PF+ GL
Sbjct: 592 KWSTFKVITSAAEAQTYTAANFIAGSTWLGSTGFPFSLGL 631
>sptr|Q84R10|Q84R10 Putative pectinesterase.
Length = 519
Score = 144 bits (363), Expect = 4e-34
Identities = 61/102 (59%), Positives = 78/102 (76%)
Frame = -1
Query: 454 RAKTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSG 275
R K+YLGRPWK+YSRTV+L++++D L+DP GW EW+GS+AL TLYYGE+MNTG GAGT
Sbjct: 418 RFKSYLGRPWKKYSRTVFLKTDIDELIDPRGWREWSGSYALSTLYYGEFMNTGAGAGTGR 477
Query: 274 RVKWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
RV W G+ V+ EAS FTV FI GD W+ T +PF+ G+
Sbjct: 478 RVNWPGFHVLRGEEEASPFTVSRFIQGDSWIPITGVPFSAGV 519
>sptr|Q9LZZ0|Q9LZZ0 Pectinesterase-like protein.
Length = 496
Score = 144 bits (363), Expect = 4e-34
Identities = 61/102 (59%), Positives = 78/102 (76%)
Frame = -1
Query: 454 RAKTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSG 275
R K+YLGRPWK+YSRTV+L++++D L+DP GW EW+GS+AL TLYYGE+MNTG GAGT
Sbjct: 395 RFKSYLGRPWKKYSRTVFLKTDIDELIDPRGWREWSGSYALSTLYYGEFMNTGAGAGTGR 454
Query: 274 RVKWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
RV W G+ V+ EAS FTV FI GD W+ T +PF+ G+
Sbjct: 455 RVNWPGFHVLRGEEEASPFTVSRFIQGDSWIPITGVPFSAGV 496
>sptr|Q9LXK7|Q9LXK7 Putative pectinesterase (EC 3.1.1.11) (Pectin
methylesterase) (PE).
Length = 527
Score = 143 bits (361), Expect = 7e-34
Identities = 62/99 (62%), Positives = 75/99 (75%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK YSRTV++Q+ + ++P GWLEWNG+FALDTLYYGEYMN+GPGA RVK
Sbjct: 427 TYLGRPWKLYSRTVFMQNYMSDAINPVGWLEWNGNFALDTLYYGEYMNSGPGASLDRRVK 486
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
W GY V+ ++AEA+ FTV I G+LWL T I F GL
Sbjct: 487 WPGYHVLNTSAEANNFTVSQLIQGNLWLPSTGITFIAGL 525
>sw|P09607|PM21_LYCES Pectinesterase 2 precursor (EC 3.1.1.11)
(Pectin methylesterase 2) (PE 2).
Length = 550
Score = 142 bits (358), Expect = 2e-33
Identities = 63/99 (63%), Positives = 73/99 (73%)
Frame = -1
Query: 445 TYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVK 266
TYLGRPWK+YSRTV ++S L L+DP+GW EW+G FAL TLYYGE+MN GPGAGTS RVK
Sbjct: 450 TYLGRPWKKYSRTVVMESYLGGLIDPSGWAEWHGDFALKTLYYGEFMNNGPGAGTSKRVK 509
Query: 265 WKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
W GY VIT AEA +FTV I G WL T + + GL
Sbjct: 510 WPGYHVITDPAEAMSFTVAKLIQGGSWLRSTDVAYVDGL 548
>sptr|Q9FEU0|Q9FEU0 Putative pectin methylesterase precursor.
Length = 574
Score = 142 bits (358), Expect = 2e-33
Identities = 62/98 (63%), Positives = 74/98 (75%)
Frame = -1
Query: 442 YLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVKW 263
+LGRPWK+YSRTV +QS + ++ PAGW EW G FALDTL++ EY N+G GAGTSGRV W
Sbjct: 477 FLGRPWKEYSRTVIMQSSISDVISPAGWREWKGRFALDTLHFAEYENSGAGAGTSGRVPW 536
Query: 262 KGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KGY+VIT A EA AFT NFI G WL T+ PF+ GL
Sbjct: 537 KGYKVITDATEAQAFTARNFITGSSWLKSTTFPFSLGL 574
>sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment).
Length = 274
Score = 142 bits (357), Expect = 2e-33
Identities = 64/100 (64%), Positives = 75/100 (75%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWK YSRTV++QS + +DP GW W+G FAL TLYYGEYMN GPGAGTS RV
Sbjct: 176 KTYLGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRV 235
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW GY ++ SAAEA+ FTVG I G +WL T + +T GL
Sbjct: 236 KWPGYHIL-SAAEATKFTVGQLIQGGVWLKSTGVAYTEGL 274
>sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 142 bits (357), Expect = 2e-33
Identities = 64/100 (64%), Positives = 75/100 (75%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWK YSRTV++QS + +DP GW W+G FAL TLYYGEYMN GPGAGTS RV
Sbjct: 217 KTYLGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRV 276
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW GY ++ SAAEA+ FTVG I G +WL T + +T GL
Sbjct: 277 KWPGYHIL-SAAEATKFTVGQLIQGGVWLKSTGVAYTEGL 315
>sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 142 bits (357), Expect = 2e-33
Identities = 64/100 (64%), Positives = 75/100 (75%)
Frame = -1
Query: 448 KTYLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRV 269
KTYLGRPWK YSRTV++QS + +DP GW W+G FAL TLYYGEYMN GPGAGTS RV
Sbjct: 217 KTYLGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRV 276
Query: 268 KWKGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KW GY ++ SAAEA+ FTVG I G +WL T + +T GL
Sbjct: 277 KWPGYHIL-SAAEATKFTVGQLIQGGVWLKSTGVAYTEGL 315
>sptr|Q9FET9|Q9FET9 Putative pectin methylesterase (Fragment).
Length = 536
Score = 142 bits (357), Expect = 2e-33
Identities = 62/98 (63%), Positives = 74/98 (75%)
Frame = -1
Query: 442 YLGRPWKQYSRTVYLQSELDSLVDPAGWLEWNGSFALDTLYYGEYMNTGPGAGTSGRVKW 263
YLGRPWK+YSRTV +QS + ++ PAGW EW G FAL+TL++ EY N+G GAGTSGRV W
Sbjct: 439 YLGRPWKEYSRTVIMQSSISDVISPAGWREWKGRFALNTLHFAEYENSGAGAGTSGRVPW 498
Query: 262 KGYRVITSAAEASAFTVGNFIDGDLWLAGTSIPFTTGL 149
KGY+VIT A EA AFT NFI G WL T+ PF+ GL
Sbjct: 499 KGYKVITDATEAQAFTARNFITGSSWLKSTTFPFSLGL 536
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 311,909,627
Number of Sequences: 1395590
Number of extensions: 6102286
Number of successful extensions: 21175
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 20139
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 21044
length of database: 442,889,342
effective HSP length: 112
effective length of database: 286,583,262
effective search space used: 11176747218
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

