BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
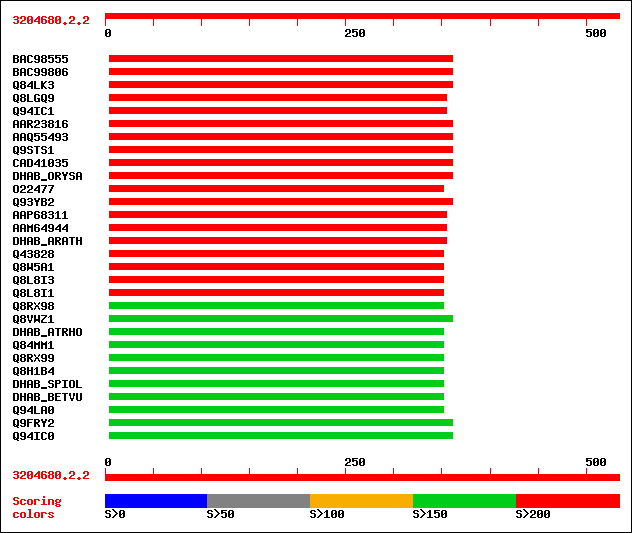
Query= 3204680.2.2
(534 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|BAC98555|BAC98555 Putative betaine-aldehyde dehydrogenase. 246 2e-64
sptrnew|BAC99806|BAC99806 Putative betaine-aldehyde dehydrogenase. 246 2e-64
sptr|Q84LK3|Q84LK3 Betaine aldehyde dehydrogenase. 246 2e-64
sptr|Q8LGQ9|Q8LGQ9 Betaine-aldehyde dehydrogenase. 237 6e-62
sptr|Q94IC1|Q94IC1 Betaine aldehyde dehydrogenase. 236 2e-61
sptrnew|AAR23816|AAR23816 Betaine-aldehyde dehydrogenase. 221 3e-57
sptrnew|AAQ55493|AAQ55493 Betaine aldehyde dehydrogenase. 221 4e-57
sptr|Q9STS1|Q9STS1 Betaine aldehyde dehydrogenase-like protein (... 220 9e-57
sptrnew|CAD41035|CAD41035 OSJNBa0060P14.8 protein. 213 9e-55
sw|O24174|DHAB_ORYSA Betaine-aldehyde dehydrogenase (EC 1.2.1.8)... 213 9e-55
sptr|O22477|O22477 Betaine aldehyde dehydrogenase (EC 1.2.1.8). 212 2e-54
sptr|Q93YB2|Q93YB2 Putative aminoaldehyde dehydrogenase. 206 1e-52
sptrnew|AAP68311|AAP68311 At1g74920. 203 9e-52
sptrnew|AAM64944|AAM64944 Betaine aldehyde dehydrogenase, putative. 203 9e-52
sw|Q9S795|DHAB_ARATH Betaine-aldehyde dehydrogenase, chloroplast... 203 9e-52
sptr|Q43828|Q43828 Betaine aldehyde dehydrogenase (EC 1.2.1.8) (... 201 3e-51
sptr|Q8W5A1|Q8W5A1 Betaine aldehyde dehydrogenase (EC 1.2.1.8). 201 6e-51
sptr|Q8L8I3|Q8L8I3 Betaine aldehyde dehydrogenase. 201 6e-51
sptr|Q8L8I1|Q8L8I1 Betaine aldehyde dehydrogenase. 201 6e-51
sptr|Q8RX98|Q8RX98 Betaine aldehyde dehydrogenase BADH2 (Fragment). 200 8e-51
sptr|Q8VWZ1|Q8VWZ1 Aminoaldehyde dehydrogenase (EC 1.2.1.19). 199 2e-50
sw|P42757|DHAB_ATRHO Betaine-aldehyde dehydrogenase, chloroplast... 199 2e-50
sptr|Q84MM1|Q84MM1 Betaine aldehyde dehydrogenase. 199 2e-50
sptr|Q8RX99|Q8RX99 Betaine aldehyde dehydrogenase BADH1. 199 2e-50
sptr|Q8H1B4|Q8H1B4 Betaine aldehyde dehydrogenase. 198 3e-50
sw|P17202|DHAB_SPIOL Betaine-aldehyde dehydrogenase, chloroplast... 198 3e-50
sw|P28237|DHAB_BETVU Betaine-aldehyde dehydrogenase, chloroplast... 197 8e-50
sptr|Q94LA0|Q94LA0 Betaine aldehyde dehydrogenase. 196 2e-49
sptr|Q9FRY2|Q9FRY2 Betaine aldehyde dehydrogenase. 195 3e-49
sptr|Q94IC0|Q94IC0 Betaine aldehyde dehydrogenase. 194 5e-49
>sptrnew|BAC98555|BAC98555 Putative betaine-aldehyde dehydrogenase.
Length = 503
Score = 246 bits (627), Expect = 2e-64
Identities = 112/119 (94%), Positives = 117/119 (98%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVKEFSTE+EAIELANDT YGLAGAV+SGDRERCQRL+EEIDAGI
Sbjct: 385 TSMQIWREEVFGPVLCVKEFSTEEEAIELANDTHYGLAGAVLSGDRERCQRLTEEIDAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEY SDEPWGWY+SPSKL
Sbjct: 445 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYASDEPWGWYKSPSKL 503
>sptrnew|BAC99806|BAC99806 Putative betaine-aldehyde dehydrogenase.
Length = 503
Score = 246 bits (627), Expect = 2e-64
Identities = 112/119 (94%), Positives = 117/119 (98%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVKEFSTE+EAIELANDT YGLAGAV+SGDRERCQRL+EEIDAGI
Sbjct: 385 TSMQIWREEVFGPVLCVKEFSTEEEAIELANDTHYGLAGAVLSGDRERCQRLTEEIDAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEY SDEPWGWY+SPSKL
Sbjct: 445 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYASDEPWGWYKSPSKL 503
>sptr|Q84LK3|Q84LK3 Betaine aldehyde dehydrogenase.
Length = 503
Score = 246 bits (627), Expect = 2e-64
Identities = 112/119 (94%), Positives = 117/119 (98%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVKEFSTE+EAIELANDT YGLAGAV+SGDRERCQRL+EEIDAGI
Sbjct: 385 TSMQIWREEVFGPVLCVKEFSTEEEAIELANDTHYGLAGAVLSGDRERCQRLTEEIDAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEY SDEPWGWY+SPSKL
Sbjct: 445 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYASDEPWGWYKSPSKL 503
>sptr|Q8LGQ9|Q8LGQ9 Betaine-aldehyde dehydrogenase.
Length = 503
Score = 237 bits (605), Expect = 6e-62
Identities = 107/117 (91%), Positives = 113/117 (96%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSMEIWREEVFGPVLCVKEFSTE+EAIELANDT YGLAGAVISGDRERCQRL+EEIDAG
Sbjct: 386 TSMEIWREEVFGPVLCVKEFSTEEEAIELANDTHYGLAGAVISGDRERCQRLAEEIDAGC 445
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPS 355
IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLS+KQVTEY SD PWGWY++P+
Sbjct: 446 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSIKQVTEYTSDAPWGWYKAPA 502
>sptr|Q94IC1|Q94IC1 Betaine aldehyde dehydrogenase.
Length = 503
Score = 236 bits (601), Expect = 2e-61
Identities = 106/117 (90%), Positives = 113/117 (96%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSMEIWREEVFGPVLCVKEFSTE+EAIELANDT YGLAGAVISGDRERCQRL+EEI+AG
Sbjct: 386 TSMEIWREEVFGPVLCVKEFSTEEEAIELANDTHYGLAGAVISGDRERCQRLAEEIEAGC 445
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPS 355
IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLS+KQVTEY SD PWGWY++P+
Sbjct: 446 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSIKQVTEYTSDAPWGWYKAPA 502
>sptrnew|AAR23816|AAR23816 Betaine-aldehyde dehydrogenase.
Length = 503
Score = 221 bits (564), Expect = 3e-57
Identities = 98/119 (82%), Positives = 108/119 (90%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK F TE+EA+ELANDT YGL AVIS D ERC R+S+ + AGI
Sbjct: 385 TSMQIWREEVFGPVLCVKTFRTEEEALELANDTHYGLGAAVISNDLERCDRVSKNLQAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+WVNCSQPCFCQAPWGGNKRSGFGRELGE G+DNYLSVKQVT+Y+SDEPWGWYRSPSKL
Sbjct: 445 VWVNCSQPCFCQAPWGGNKRSGFGRELGEWGLDNYLSVKQVTQYVSDEPWGWYRSPSKL 503
>sptrnew|AAQ55493|AAQ55493 Betaine aldehyde dehydrogenase.
Length = 503
Score = 221 bits (563), Expect = 4e-57
Identities = 98/119 (82%), Positives = 111/119 (93%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSMEIWR+EVFGPVLCVK FSTEDEAI+LAND+QYGLAGAV+S D ERC R+S+ +AGI
Sbjct: 385 TSMEIWRDEVFGPVLCVKTFSTEDEAIQLANDSQYGLAGAVLSNDLERCDRVSKAFEAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+WVNCSQPCFCQAPWGG KRSGFGRELGE G++NYLSVKQVT+YIS+EPWGWY+SPSKL
Sbjct: 445 VWVNCSQPCFCQAPWGGTKRSGFGRELGEWGLENYLSVKQVTQYISNEPWGWYKSPSKL 503
>sptr|Q9STS1|Q9STS1 Betaine aldehyde dehydrogenase-like protein
(Putative aldehyde dehydrogenase).
Length = 503
Score = 220 bits (560), Expect = 9e-57
Identities = 98/119 (82%), Positives = 108/119 (90%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSMEIWREEVFGP LCVK FSTEDEAI+LAND+QYGLAGAV+S D ERC R+S+ AGI
Sbjct: 385 TSMEIWREEVFGPALCVKTFSTEDEAIQLANDSQYGLAGAVLSNDLERCDRVSKAFQAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+WVNCSQPCFCQAPWGG KRSGFGRELGE G++NYLSVKQVT+YISDEPWGWY+ PSKL
Sbjct: 445 VWVNCSQPCFCQAPWGGTKRSGFGRELGEWGLENYLSVKQVTQYISDEPWGWYKPPSKL 503
>sptrnew|CAD41035|CAD41035 OSJNBa0060P14.8 protein.
Length = 505
Score = 213 bits (543), Expect = 9e-55
Identities = 95/119 (79%), Positives = 108/119 (90%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPV+CVKEF TE EA+ELANDT YGLAGAVIS D ERC+R+S+ I +GI
Sbjct: 387 TSMQIWREEVFGPVICVKEFRTEREAVELANDTHYGLAGAVISNDLERCERISKAIQSGI 446
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+W+NCSQPCF QAPWGGNKRSGFGRELG+ G+DNYLSVKQVT+Y SDEP+GWYR PSKL
Sbjct: 447 VWINCSQPCFVQAPWGGNKRSGFGRELGQWGLDNYLSVKQVTKYCSDEPYGWYRPPSKL 505
>sw|O24174|DHAB_ORYSA Betaine-aldehyde dehydrogenase (EC 1.2.1.8)
(BADH).
Length = 505
Score = 213 bits (543), Expect = 9e-55
Identities = 95/119 (79%), Positives = 108/119 (90%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPV+CVKEF TE EA+ELANDT YGLAGAVIS D ERC+R+S+ I +GI
Sbjct: 387 TSMQIWREEVFGPVICVKEFRTEREAVELANDTHYGLAGAVISNDLERCERISKAIQSGI 446
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+W+NCSQPCF QAPWGGNKRSGFGRELG+ G+DNYLSVKQVT+Y SDEP+GWYR PSKL
Sbjct: 447 VWINCSQPCFVQAPWGGNKRSGFGRELGQWGLDNYLSVKQVTKYCSDEPYGWYRPPSKL 505
>sptr|O22477|O22477 Betaine aldehyde dehydrogenase (EC 1.2.1.8).
Length = 500
Score = 212 bits (540), Expect = 2e-54
Identities = 92/116 (79%), Positives = 106/116 (91%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK F +EDEAIELANDTQYGL AV+S D +RC+R+++ + AGI
Sbjct: 385 TSMQIWREEVFGPVLCVKTFGSEDEAIELANDTQYGLGAAVLSKDLDRCERITKALQAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCFCQAPWGG KRSGFGRELGE GI+NYL++KQVTEYISDEPWGWY+SP
Sbjct: 445 VWVNCSQPCFCQAPWGGTKRSGFGRELGEWGIENYLNIKQVTEYISDEPWGWYKSP 500
>sptr|Q93YB2|Q93YB2 Putative aminoaldehyde dehydrogenase.
Length = 503
Score = 206 bits (524), Expect = 1e-52
Identities = 92/119 (77%), Positives = 106/119 (89%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
T+M+IWREEVFGPVLCVK FSTE+EAI+LANDT YGL AVIS D ERC+R+++ AGI
Sbjct: 385 TNMQIWREEVFGPVLCVKTFSTEEEAIDLANDTVYGLGAAVISNDLERCERVTKAFKAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+WVNCSQPCF QAPWGG KRSGFGRELGE G+DNYLSVKQVT+YIS+EPWGWY+ P+KL
Sbjct: 445 VWVNCSQPCFTQAPWGGVKRSGFGRELGEWGLDNYLSVKQVTQYISEEPWGWYQPPAKL 503
>sptrnew|AAP68311|AAP68311 At1g74920.
Length = 501
Score = 203 bits (517), Expect = 9e-52
Identities = 90/117 (76%), Positives = 103/117 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK F++EDEAIELAND+ YGL AVIS D ERC R+SE +AGI
Sbjct: 385 TSMQIWREEVFGPVLCVKTFASEDEAIELANDSHYGLGAAVISNDTERCDRISEAFEAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPS 355
+W+NCSQPCF QAPWGG KRSGFGRELGE G+DNYLSVKQVT Y S++PWGWY+SP+
Sbjct: 445 VWINCSQPCFTQAPWGGVKRSGFGRELGEWGLDNYLSVKQVTLYTSNDPWGWYKSPN 501
>sptrnew|AAM64944|AAM64944 Betaine aldehyde dehydrogenase, putative.
Length = 501
Score = 203 bits (517), Expect = 9e-52
Identities = 90/117 (76%), Positives = 103/117 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK F++EDEAIELAND+ YGL AVIS D ERC R+SE +AGI
Sbjct: 385 TSMQIWREEVFGPVLCVKTFASEDEAIELANDSHYGLGAAVISNDTERCDRISEAFEAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPS 355
+W+NCSQPCF QAPWGG KRSGFGRELGE G+DNYLSVKQVT Y S++PWGWY+SP+
Sbjct: 445 VWINCSQPCFTQAPWGGVKRSGFGRELGEWGLDNYLSVKQVTLYTSNDPWGWYKSPN 501
>sw|Q9S795|DHAB_ARATH Betaine-aldehyde dehydrogenase, chloroplast
precursor (EC 1.2.1.8) (BADH).
Length = 501
Score = 203 bits (517), Expect = 9e-52
Identities = 90/117 (76%), Positives = 103/117 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK F++EDEAIELAND+ YGL AVIS D ERC R+SE +AGI
Sbjct: 385 TSMQIWREEVFGPVLCVKTFASEDEAIELANDSHYGLGAAVISNDTERCDRISEAFEAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPS 355
+W+NCSQPCF QAPWGG KRSGFGRELGE G+DNYLSVKQVT Y S++PWGWY+SP+
Sbjct: 445 VWINCSQPCFTQAPWGGVKRSGFGRELGEWGLDNYLSVKQVTLYTSNDPWGWYKSPN 501
>sptr|Q43828|Q43828 Betaine aldehyde dehydrogenase (EC 1.2.1.8)
(Fragment).
Length = 424
Score = 201 bits (512), Expect = 3e-51
Identities = 100/116 (86%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSMEI V LCVK STEDEAIELAN TQYGLAGAV+SGDRERCQRLSEEIDAG
Sbjct: 292 TSMEILEGGVLVS-LCVK-ISTEDEAIELANHTQYGLAGAVLSGDRERCQRLSEEIDAGC 349
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
IWVNCSQPCFCQAPWGGNKRS FGRELGEGGIDNYLSVKQVTEYISDE WGWY+SP
Sbjct: 350 IWVNCSQPCFCQAPWGGNKRSDFGRELGEGGIDNYLSVKQVTEYISDEKWGWYQSP 405
>sptr|Q8W5A1|Q8W5A1 Betaine aldehyde dehydrogenase (EC 1.2.1.8).
Length = 501
Score = 201 bits (510), Expect = 6e-51
Identities = 87/116 (75%), Positives = 103/116 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPVLCVK FSTE+EA+ELANDT+YGLA AV S D ERC+R+++ ++ G
Sbjct: 386 TSMQIWKEEVFGPVLCVKTFSTEEEALELANDTEYGLAAAVFSKDLERCERVTKALEVGA 445
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCFC APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 446 VWVNCSQPCFCHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 501
>sptr|Q8L8I3|Q8L8I3 Betaine aldehyde dehydrogenase.
Length = 500
Score = 201 bits (510), Expect = 6e-51
Identities = 88/116 (75%), Positives = 103/116 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPV+CVK FSTEDEAIELANDT+YGLAGAV S D ERC+R+++ ++ G
Sbjct: 385 TSMQIWKEEVFGPVICVKTFSTEDEAIELANDTEYGLAGAVFSKDLERCERVTKALEVGA 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 445 VWVNCSQPCFVHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 500
>sptr|Q8L8I1|Q8L8I1 Betaine aldehyde dehydrogenase.
Length = 500
Score = 201 bits (510), Expect = 6e-51
Identities = 88/116 (75%), Positives = 103/116 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPV+CVK FSTEDEAIELANDT+YGLAGAV S D ERC+R+++ ++ G
Sbjct: 385 TSMQIWKEEVFGPVICVKTFSTEDEAIELANDTEYGLAGAVFSKDLERCERVTKALEVGA 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 445 VWVNCSQPCFVHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 500
>sptr|Q8RX98|Q8RX98 Betaine aldehyde dehydrogenase BADH2 (Fragment).
Length = 424
Score = 200 bits (509), Expect = 8e-51
Identities = 86/116 (74%), Positives = 105/116 (90%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPVLCVK FS+++EAIELANDTQYGL AV+S + ERC+++++ ++ GI
Sbjct: 309 TSMQIWKEEVFGPVLCVKTFSSDEEAIELANDTQYGLGAAVLSKNLERCEKVTKALEVGI 368
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCFCQAPWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 369 VWVNCSQPCFCQAPWGGAKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 424
>sptr|Q8VWZ1|Q8VWZ1 Aminoaldehyde dehydrogenase (EC 1.2.1.19).
Length = 503
Score = 199 bits (506), Expect = 2e-50
Identities = 90/119 (75%), Positives = 102/119 (85%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVL VK FSTE+EAI LANDT YGL AV+S D ERC+RLS+ + AGI
Sbjct: 385 TSMQIWREEVFGPVLAVKTFSTEEEAINLANDTHYGLGSAVMSNDLERCERLSKALQAGI 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+W+NC+QP F QAPWGG KRSGFGRELGE G++NYLSVKQVT Y SDEPWGWY+ PSKL
Sbjct: 445 VWINCAQPSFIQAPWGGIKRSGFGRELGEWGLENYLSVKQVTRYTSDEPWGWYQPPSKL 503
>sw|P42757|DHAB_ATRHO Betaine-aldehyde dehydrogenase, chloroplast
precursor (EC 1.2.1.8) (BADH).
Length = 502
Score = 199 bits (506), Expect = 2e-50
Identities = 87/116 (75%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPV+CVK F TEDEAIELANDT+YGLAGAV S D ERC+R+++ ++ G
Sbjct: 387 TSMQIWKEEVFGPVICVKTFKTEDEAIELANDTEYGLAGAVFSKDLERCERVTKALEVGA 446
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 447 VWVNCSQPCFVHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 502
>sptr|Q84MM1|Q84MM1 Betaine aldehyde dehydrogenase.
Length = 500
Score = 199 bits (506), Expect = 2e-50
Identities = 87/116 (75%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPV+CVK F TEDEAIELANDT+YGLAGAV S D ERC+R+++ ++ G
Sbjct: 385 TSMQIWKEEVFGPVICVKTFKTEDEAIELANDTEYGLAGAVFSKDLERCERVTKALEVGA 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 445 VWVNCSQPCFVHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 500
>sptr|Q8RX99|Q8RX99 Betaine aldehyde dehydrogenase BADH1.
Length = 500
Score = 199 bits (506), Expect = 2e-50
Identities = 87/116 (75%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPV+CVK F TEDEAIELANDT+YGLAGAV S D ERC+R+++ ++ G
Sbjct: 385 TSMQIWKEEVFGPVICVKTFKTEDEAIELANDTEYGLAGAVFSKDLERCERVTKALEVGA 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT ISDEPWGWY+SP
Sbjct: 445 VWVNCSQPCFVHAPWGGVKRSGFGRELGEWGIENYLNIKQVTSDISDEPWGWYKSP 500
>sptr|Q8H1B4|Q8H1B4 Betaine aldehyde dehydrogenase.
Length = 497
Score = 198 bits (504), Expect = 3e-50
Identities = 87/116 (75%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPVLCVK FS+EDEAI LANDT+YGLA AV S D ERC+R+++ ++ G
Sbjct: 382 TSMQIWKEEVFGPVLCVKTFSSEDEAIALANDTEYGLAAAVFSNDLERCERITKALEVGA 441
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF QAPWGG KRSGFGRELGE GI NYL++KQVT+ ISDEPWGWY+SP
Sbjct: 442 VWVNCSQPCFVQAPWGGIKRSGFGRELGEWGIQNYLNIKQVTQDISDEPWGWYKSP 497
>sw|P17202|DHAB_SPIOL Betaine-aldehyde dehydrogenase, chloroplast
precursor (EC 1.2.1.8) (BADH).
Length = 497
Score = 198 bits (504), Expect = 3e-50
Identities = 87/116 (75%), Positives = 102/116 (87%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW+EEVFGPVLCVK FS+EDEAI LANDT+YGLA AV S D ERC+R+++ ++ G
Sbjct: 382 TSMQIWKEEVFGPVLCVKTFSSEDEAIALANDTEYGLAAAVFSNDLERCERITKALEVGA 441
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF QAPWGG KRSGFGRELGE GI NYL++KQVT+ ISDEPWGWY+SP
Sbjct: 442 VWVNCSQPCFVQAPWGGIKRSGFGRELGEWGIQNYLNIKQVTQDISDEPWGWYKSP 497
>sw|P28237|DHAB_BETVU Betaine-aldehyde dehydrogenase, chloroplast
precursor (EC 1.2.1.8) (BADH).
Length = 500
Score = 197 bits (500), Expect = 8e-50
Identities = 87/116 (75%), Positives = 103/116 (88%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPVLCVK FS+EDEA+ELANDT+YGLA AV S D ERC+R+S+ +++G
Sbjct: 385 TSMQIWREEVFGPVLCVKTFSSEDEALELANDTEYGLASAVFSKDLERCERVSKLLESGA 444
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF APWGG KRSGFGRELGE GI+NYL++KQVT IS+EPWGWY+SP
Sbjct: 445 VWVNCSQPCFVHAPWGGIKRSGFGRELGEWGIENYLNIKQVTSDISNEPWGWYKSP 500
>sptr|Q94LA0|Q94LA0 Betaine aldehyde dehydrogenase.
Length = 496
Score = 196 bits (497), Expect = 2e-49
Identities = 86/116 (74%), Positives = 99/116 (85%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW EEVFGPVLCVK F+TEDEA+ELANDT YGLA AVIS D +RC+R+++ G
Sbjct: 381 TSMQIWIEEVFGPVLCVKTFATEDEAVELANDTHYGLAAAVISKDLDRCERMAKAFQVGA 440
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSP 352
+WVNCSQPCF QAPWGG KRSGFGRELGE G+D YL+VKQVT Y+S EPWGWY+SP
Sbjct: 441 VWVNCSQPCFYQAPWGGKKRSGFGRELGERGLDIYLNVKQVTRYVSSEPWGWYKSP 496
>sptr|Q9FRY2|Q9FRY2 Betaine aldehyde dehydrogenase.
Length = 502
Score = 195 bits (495), Expect = 3e-49
Identities = 87/119 (73%), Positives = 99/119 (83%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IW EEVFGPVL VK FS EDE ELAND YG A AV+S D ERC+R+++ AGI
Sbjct: 384 TSMQIWIEEVFGPVLGVKTFSKEDEPFELANDPHYGWAAAVLSQDLERCERMTKAFQAGI 443
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+WVNCSQPCFCQAPWGG KRSGFGR+LGE G+DNYL+VKQVT Y+S EPW WY+SPSKL
Sbjct: 444 VWVNCSQPCFCQAPWGGKKRSGFGRDLGEWGLDNYLNVKQVTRYVSSEPWDWYKSPSKL 502
>sptr|Q94IC0|Q94IC0 Betaine aldehyde dehydrogenase.
Length = 506
Score = 194 bits (493), Expect = 5e-49
Identities = 87/119 (73%), Positives = 101/119 (84%)
Frame = +2
Query: 5 TSMEIWREEVFGPVLCVKEFSTEDEAIELANDTQYGLAGAVISGDRERCQRLSEEIDAGI 184
TSM+IWREEVFGPV+CVK F TE EA+ELANDT YGLAG VIS D ERC+R+++ I +GI
Sbjct: 388 TSMQIWREEVFGPVICVKVFKTESEAVELANDTHYGLAGGVISDDLERCERIAKVIHSGI 447
Query: 185 IWVNCSQPCFCQAPWGGNKRSGFGRELGEGGIDNYLSVKQVTEYISDEPWGWYRSPSKL 361
+W+NCSQP QAPWGGNKRSGFGRELGE G++NYLSVKQVT Y DE +GWY+ PSKL
Sbjct: 448 VWINCSQPTLVQAPWGGNKRSGFGRELGEWGLENYLSVKQVTRYCKDELYGWYQRPSKL 506
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 440,748,422
Number of Sequences: 1395590
Number of extensions: 8850122
Number of successful extensions: 22335
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 21502
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 21965
length of database: 442,889,342
effective HSP length: 115
effective length of database: 282,396,492
effective search space used: 17508582504
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

