BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
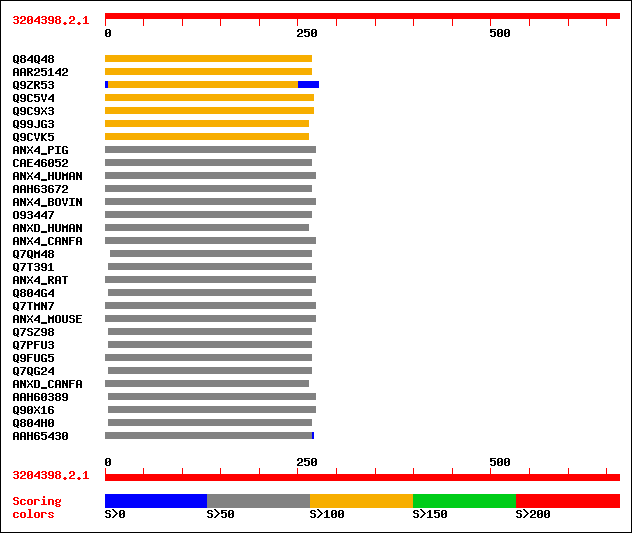
Query= 3204398.2.1
(666 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q84Q48|Q84Q48 P0456B03.13 protein. 145 6e-34
sptrnew|AAR25142|AAR25142 Annexin. 141 9e-33
sptr|Q9ZR53|Q9ZR53 Annexin-like protein. 132 4e-30
sptr|Q9C5V4|Q9C5V4 Calcium-binding protein annexin 5. 127 2e-28
sptr|Q9C9X3|Q9C9X3 Putative annexin. 127 2e-28
sptr|Q99JG3|Q99JG3 Annexin A13 isoform a (Hypothetical protein). 102 3e-21
sptr|Q9CVK5|Q9CVK5 1810034H17Rik protein (Fragment). 102 3e-21
sw|P08132|ANX4_PIG Annexin A4 (Annexin IV) (Lipocortin IV) (Endo... 98 1e-19
sptrnew|CAE46052|CAE46052 Hypothetical protein DKFZp686H02120 (F... 97 2e-19
sw|P09525|ANX4_HUMAN Annexin A4 (Annexin IV) (Lipocortin IV) (En... 97 2e-19
sptrnew|AAH63672|AAH63672 Hypothetical protein. 97 2e-19
sw|P13214|ANX4_BOVIN Annexin A4 (Annexin IV) (Lipocortin IV) (En... 97 2e-19
sptr|O93447|O93447 Annexin max4. 97 2e-19
sw|P27216|ANXD_HUMAN Annexin A13 (Annexin XIII) (Annexin, intest... 96 3e-19
sw|P50994|ANX4_CANFA Annexin A4 (Annexin IV) (Lipocortin IV) (36... 96 4e-19
sptr|Q7QM48|Q7QM48 AgCP15598 (Fragment). 96 6e-19
sptr|Q7T391|Q7T391 Hypothetical protein. 95 7e-19
sw|P55260|ANX4_RAT Annexin A4 (Annexin IV) (Lipocortin IV) (36 k... 95 7e-19
sptr|Q804G4|Q804G4 Annexin 11a. 95 7e-19
sptr|Q7TMN7|Q7TMN7 Annexin A4. 95 9e-19
sw|P97429|ANX4_MOUSE Annexin A4 (Annexin IV). 95 9e-19
sptr|Q7SZ98|Q7SZ98 Hypothetical protein. 94 2e-18
sptr|Q7PFU3|Q7PFU3 ENSANGP00000022424 (Fragment). 92 8e-18
sptr|Q9FUG5|Q9FUG5 Annexin. 92 8e-18
sptr|Q7QG24|Q7QG24 AgCP13874 (Fragment). 91 1e-17
sw|Q29471|ANXD_CANFA Annexin A13 (Annexin XIII) (Annexin, intest... 91 1e-17
sptrnew|AAH60389|AAH60389 MGC68504 protein. 91 1e-17
sptr|Q90X16|Q90X16 Annexin 4. 91 1e-17
sptr|Q804H0|Q804H0 Annexin 1c. 90 2e-17
sptrnew|AAH65430|AAH65430 Hypothetical protein. 90 3e-17
>sptr|Q84Q48|Q84Q48 P0456B03.13 protein.
Length = 321
Score = 145 bits (365), Expect = 6e-34
Identities = 69/89 (77%), Positives = 79/89 (88%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSGNF F LL ILRCAE+PAKYFAK+L +AMKGLGT+D TLIRVV TR E+DMQYIKAEY
Sbjct: 230 TSGNFGFGLLTILRCAESPAKYFAKVLHEAMKGLGTNDTTLIRVVTTRAEVDMQYIKAEY 289
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ YK+ LA+A+HSETSGNYRTFLLSL+G
Sbjct: 290 HRSYKRSLADAVHSETSGNYRTFLLSLIG 318
Score = 50.8 bits (120), Expect = 2e-05
Identities = 35/97 (36%), Positives = 51/97 (52%), Gaps = 10/97 (10%)
Frame = +1
Query: 4 SGNFEFALLAILRCA--ENP--------AKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEI 153
SG+ + LLA LR E P A+ +L R + LGTD++T IRV R+
Sbjct: 146 SGDHQRLLLAYLRSPRYEGPEVVDMAAAARDARELYRAGERRLGTDERTFIRVFSERSAA 205
Query: 154 DMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
M + A Y Y + L +A+ SETSGN+ LL+++
Sbjct: 206 HMAAVAAAYHHMYDRSLEKAVKSETSGNFGFGLLTIL 242
>sptrnew|AAR25142|AAR25142 Annexin.
Length = 316
Score = 141 bits (355), Expect = 9e-33
Identities = 69/89 (77%), Positives = 76/89 (85%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
T+GNF+F LL ILRCA+ PAKYFAK+L KAMKGLGT + L RV VTRTE+DM+YIKAEY
Sbjct: 225 TTGNFQFGLLTILRCADTPAKYFAKVLHKAMKGLGTSNAALTRVAVTRTEVDMKYIKAEY 284
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
KYK LAEAIHSETSGNYRTFLLSLVG
Sbjct: 285 HNKYKGSLAEAIHSETSGNYRTFLLSLVG 313
Score = 40.4 bits (93), Expect = 0.021
Identities = 20/64 (31%), Positives = 37/64 (57%)
Frame = +1
Query: 73 KLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFL 252
+L + K +GTD++ IR+ R+ M + Y Y + L +A+ SET+GN++ L
Sbjct: 174 ELYQAGEKRVGTDERAFIRIFSERSWAHMVSVANAYQHMYARSLEKAVKSETTGNFQFGL 233
Query: 253 LSLV 264
L+++
Sbjct: 234 LTIL 237
>sptr|Q9ZR53|Q9ZR53 Annexin-like protein.
Length = 333
Score = 132 bits (332), Expect = 4e-30
Identities = 65/82 (79%), Positives = 70/82 (85%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNF ALL I CA NPAKYFAK+L KAMKGLGT+D TLIRV+VTRTEIDMQYIKAEY
Sbjct: 225 SGNFGLALLTITECATNPAKYFAKVLYKAMKGLGTNDSTLIRVIVTRTEIDMQYIKAEYA 284
Query: 184 KKYKKPLAEAIHSETSGNYRTF 249
KKYKK L +A+HSETSGNYR F
Sbjct: 285 KKYKKTLNDAVHSETSGNYRIF 306
Score = 43.1 bits (100), Expect = 0.003
Identities = 30/96 (31%), Positives = 49/96 (51%), Gaps = 9/96 (9%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENP--------AKYFAKLLRKA-MKGLGTDDKTLIRVVVTRTEI 153
TSG+ + LL L + A+ AK+L KA K LGTD+KT +++ R+
Sbjct: 140 TSGDHQKILLRYLTTPRHEGLEVNREIAQKDAKVLYKAGEKKLGTDEKTFVQIFSERSSA 199
Query: 154 DMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSL 261
+ + + Y Y L +A+ +E SGN+ LL++
Sbjct: 200 HLAAVSSYYHDMYGHSLKKAVKNEASGNFGLALLTI 235
Score = 36.6 bits (83), Expect = 0.31
Identities = 25/83 (30%), Positives = 42/83 (50%), Gaps = 1/83 (1%)
Frame = +1
Query: 31 AILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAE 210
A+L +PA A+++RK++ + + V+ +RT +QY+K Y K+ L
Sbjct: 76 AVLLWLPDPAARDAEIIRKSLV-VDRSLEAATEVICSRTPSQLQYLKQLYHSKFGVYLEH 134
Query: 211 AIHSETSGNYRTFLLS-LVGPGH 276
I TSG+++ LL L P H
Sbjct: 135 EIELNTSGDHQKILLRYLTTPRH 157
>sptr|Q9C5V4|Q9C5V4 Calcium-binding protein annexin 5.
Length = 316
Score = 127 bits (318), Expect = 2e-28
Identities = 61/90 (67%), Positives = 72/90 (80%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
T GNFE LL IL+CAEN YFAK LRK+MKGLGTDD LIR+VVTR E+DMQ+I EY
Sbjct: 225 TRGNFEHVLLTILQCAENSCFYFAKALRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEY 284
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVGP 270
K+YKK L A+HS+T+ +YRTFLLSL+GP
Sbjct: 285 RKRYKKTLYNAVHSDTTSHYRTFLLSLLGP 314
Score = 43.1 bits (100), Expect = 0.003
Identities = 29/99 (29%), Positives = 47/99 (47%), Gaps = 12/99 (12%)
Frame = +1
Query: 4 SGNFEFALLAILRCA------------ENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRT 147
SGN + LLA L EN A+ + + K +DD+TLI++ R+
Sbjct: 142 SGNHKRVLLAYLNTTRYEGPEIDNASVENDARTLKSAVARKHK---SDDQTLIQIFTDRS 198
Query: 148 EIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ +++ Y Y K L +AI ET GN+ LL+++
Sbjct: 199 RTHLVAVRSTYRSMYGKELGKAIRDETRGNFEHVLLTIL 237
>sptr|Q9C9X3|Q9C9X3 Putative annexin.
Length = 316
Score = 127 bits (318), Expect = 2e-28
Identities = 61/90 (67%), Positives = 72/90 (80%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
T GNFE LL IL+CAEN YFAK LRK+MKGLGTDD LIR+VVTR E+DMQ+I EY
Sbjct: 225 TRGNFEHVLLTILQCAENSCFYFAKALRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEY 284
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVGP 270
K+YKK L A+HS+T+ +YRTFLLSL+GP
Sbjct: 285 RKRYKKTLYNAVHSDTTSHYRTFLLSLLGP 314
Score = 43.1 bits (100), Expect = 0.003
Identities = 29/99 (29%), Positives = 47/99 (47%), Gaps = 12/99 (12%)
Frame = +1
Query: 4 SGNFEFALLAILRCA------------ENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRT 147
SGN + LLA L EN A+ + + K +DD+TLI++ R+
Sbjct: 142 SGNHKRVLLAYLNTTRYEGPEIDNASVENDARTLKSAVARKHK---SDDQTLIQIFTDRS 198
Query: 148 EIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ +++ Y Y K L +AI ET GN+ LL+++
Sbjct: 199 RTHLVAVRSTYRSMYGKELGKAIRDETRGNFEHVLLTIL 237
>sptr|Q99JG3|Q99JG3 Annexin A13 isoform a (Hypothetical protein).
Length = 317
Score = 102 bits (255), Expect = 3e-21
Identities = 47/88 (53%), Positives = 72/88 (81%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+ + A L I+RCA++ YFA LL KAMKG+GTD++TLIR++VTR E+D+Q IKA++
Sbjct: 229 TSGDLKKAYLTIVRCAQDLEGYFADLLYKAMKGMGTDEETLIRIIVTRAEVDLQGIKAKF 288
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+KY+K L++ +HS+TSG++R L++L+
Sbjct: 289 QEKYQKSLSDMVHSDTSGDFRKLLVALL 316
Score = 65.5 bits (158), Expect = 6e-10
Identities = 35/87 (40%), Positives = 55/87 (63%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNF+ LA+L + P +Y A+ L+KAMKG+GTD+ LI ++ TR+ ++ IK Y
Sbjct: 74 SGNFKKTALALL---DRPNEYAARQLQKAMKGVGTDEAMLIEILCTRSNKEIVAIKEAYQ 130
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ + + L + +TSGN R L+SL+
Sbjct: 131 RLFGRSLESDVKEDTSGNLRKILVSLL 157
Score = 60.1 bits (144), Expect = 3e-08
Identities = 30/65 (46%), Positives = 44/65 (67%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
AK L KA KG+GTD+ +I V+ +RT + Q IK +Y +KY K L E ++SE SGN++
Sbjct: 21 AKKLYKACKGMGTDEAAIIEVLSSRTSEERQQIKQKYKEKYGKDLEEVLNSELSGNFKKT 80
Query: 250 LLSLV 264
L+L+
Sbjct: 81 ALALL 85
>sptr|Q9CVK5|Q9CVK5 1810034H17Rik protein (Fragment).
Length = 125
Score = 102 bits (255), Expect = 3e-21
Identities = 47/88 (53%), Positives = 72/88 (81%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+ + A L I+RCA++ YFA LL KAMKG+GTD++TLIR++VTR E+D+Q IKA++
Sbjct: 37 TSGDLKKAYLTIVRCAQDLEGYFADLLYKAMKGMGTDEETLIRIIVTRAEVDLQGIKAKF 96
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+KY+K L++ +HS+TSG++R L++L+
Sbjct: 97 QEKYQKSLSDMVHSDTSGDFRKLLVALL 124
>sw|P08132|ANX4_PIG Annexin A4 (Annexin IV) (Lipocortin IV)
(Endonexin I) (Chromobindin 4) (Protein II) (P32.5)
(Placental anticoagulant protein I) (PAP-II) (PP4-X)
(35-beta calcimedin).
Length = 318
Score = 97.8 bits (242), Expect = 1e-19
Identities = 51/89 (57%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 227 TSGSFEDALLAIVKCMRNKSAYFAERLYKSMKGLGTDDNTLIRVMVSRAEIDMMDIRANF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRLYGKSLYSFIKGDTSGDYRKVLLILCG 315
Score = 53.5 bits (127), Expect = 2e-06
Identities = 32/90 (35%), Positives = 51/90 (56%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE +L ++ Y + LR+AMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 72 SGNFEQVILGMMTPT---VLYDVQELRRAMKGAGTDEGCLIEILASRTPEEIRRINQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
+Y + L + I S+TS ++ L+SL G
Sbjct: 129 LQYGRSLEDDIRSDTSFMFQRVLVSLSAGG 158
Score = 52.0 bits (123), Expect = 7e-06
Identities = 26/67 (38%), Positives = 39/67 (58%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I V+ R+ Q I+ Y + L + + SE SGN+
Sbjct: 19 AQTLRKAMKGLGTDEDAIISVLAYRSTAQRQEIRTAYKSTIGRDLLDDLKSELSGNFEQV 78
Query: 250 LLSLVGP 270
+L ++ P
Sbjct: 79 ILGMMTP 85
Score = 36.2 bits (82), Expect = 0.40
Identities = 18/57 (31%), Positives = 34/57 (59%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + V+ +R + ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 183 KKWGTDEVKFLTVLCSRNRNHLLHVFDEYKRISQKDIEQSIKSETSGSFEDALLAIV 239
>sptrnew|CAE46052|CAE46052 Hypothetical protein DKFZp686H02120
(Fragment).
Length = 112
Score = 97.1 bits (240), Expect = 2e-19
Identities = 51/89 (57%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 21 TSGSFEDALLAIVKCMRNKSAYFAEKLYKSMKGLGTDDNTLIRVMVSRAEIDMLDIRAHF 80
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 81 KRLYGKSLYSFIKGDTSGDYRKVLLVLCG 109
>sw|P09525|ANX4_HUMAN Annexin A4 (Annexin IV) (Lipocortin IV)
(Endonexin I) (Chromobindin 4) (Protein II) (P32.5)
(Placental anticoagulant protein II) (PAP-II) (PP4-X)
(35-beta calcimedin) (Carbohydrate-binding protein
P33/P41) (P33/41).
Length = 318
Score = 97.1 bits (240), Expect = 2e-19
Identities = 51/89 (57%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 227 TSGSFEDALLAIVKCMRNKSAYFAEKLYKSMKGLGTDDNTLIRVMVSRAEIDMLDIRAHF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRLYGKSLYSFIKGDTSGDYRKVLLVLCG 315
Score = 53.5 bits (127), Expect = 2e-06
Identities = 31/90 (34%), Positives = 52/90 (57%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE ++ ++ Y + LR+AMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 72 SGNFEQVIVGMMTPT---VLYDVQELRRAMKGAGTDEGCLIEILASRTPEEIRRISQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
++Y + L + I S+TS ++ L+SL G
Sbjct: 129 QQYGRSLEDDIRSDTSFMFQRVLVSLSAGG 158
Score = 50.4 bits (119), Expect = 2e-05
Identities = 25/67 (37%), Positives = 38/67 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I V+ R Q I+ Y + L + + SE SGN+
Sbjct: 19 AQTLRKAMKGLGTDEDAIISVLAYRNTAQRQEIRTAYKSTIGRDLIDDLKSELSGNFEQV 78
Query: 250 LLSLVGP 270
++ ++ P
Sbjct: 79 IVGMMTP 85
Score = 36.2 bits (82), Expect = 0.40
Identities = 18/57 (31%), Positives = 34/57 (59%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + V+ +R + ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 183 KKWGTDEVKFLTVLCSRNRNHLLHVFDEYKRISQKDIEQSIKSETSGSFEDALLAIV 239
>sptrnew|AAH63672|AAH63672 Hypothetical protein.
Length = 299
Score = 97.1 bits (240), Expect = 2e-19
Identities = 51/89 (57%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 208 TSGSFEDALLAIVKCMRNKSAYFAEKLYKSMKGLGTDDNTLIRVMVSRAEIDMLDIRAHF 267
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 268 KRLYGKSLYSFIKGDTSGDYRKVLLVLCG 296
Score = 47.0 bits (110), Expect = 2e-04
Identities = 24/57 (42%), Positives = 33/57 (57%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNY 240
A+ LRKAMKGLGTD+ +I V+ R Q I+ Y + L + + SE SGN+
Sbjct: 22 AQTLRKAMKGLGTDEDAIISVLAYRNTAQRQEIRTAYKSTIGRDLIDDLKSELSGNF 78
Score = 36.2 bits (82), Expect = 0.40
Identities = 18/57 (31%), Positives = 34/57 (59%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + V+ +R + ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 164 KKWGTDEVKFLTVLCSRNRNHLLHVFDEYKRISQKDIEQSIKSETSGSFEDALLAIV 220
>sw|P13214|ANX4_BOVIN Annexin A4 (Annexin IV) (Lipocortin IV)
(Endonexin I) (Chromobindin 4) (Protein II) (P32.5)
(Placental anticoagulant protein II) (PAP-II) (PP4-X)
(35-beta calcimedin) (Carbohydrate-binding protein
P33/P41) (P33/41).
Length = 318
Score = 96.7 bits (239), Expect = 2e-19
Identities = 51/89 (57%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 227 TSGSFEDALLAIVKCMRNKSAYFAERLYKSMKGLGTDDDTLIRVMVSRAEIDMLDIRANF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRLYGKSLYSFIKGDTSGDYRKVLLILCG 315
Score = 54.7 bits (130), Expect = 1e-06
Identities = 33/90 (36%), Positives = 51/90 (56%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE +L ++ Y + LRKAMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 72 SGNFEQVILGMMTPT---VLYDVQELRKAMKGAGTDEGCLIEILASRTPEEIRRINQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
+Y + L + I S+TS ++ L+SL G
Sbjct: 129 LQYGRSLEDDIRSDTSFMFQRVLVSLSAGG 158
Score = 52.0 bits (123), Expect = 7e-06
Identities = 26/67 (38%), Positives = 39/67 (58%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I V+ R+ Q I+ Y + L + + SE SGN+
Sbjct: 19 AQTLRKAMKGLGTDEDAIINVLAYRSTAQRQEIRTAYKTTIGRDLMDDLKSELSGNFEQV 78
Query: 250 LLSLVGP 270
+L ++ P
Sbjct: 79 ILGMMTP 85
Score = 36.2 bits (82), Expect = 0.40
Identities = 18/57 (31%), Positives = 34/57 (59%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + V+ +R + ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 183 KKWGTDEVKFLTVLCSRNRNHLLHVFDEYKRIAQKDIEQSIKSETSGSFEDALLAIV 239
>sptr|O93447|O93447 Annexin max4.
Length = 508
Score = 96.7 bits (239), Expect = 2e-19
Identities = 47/89 (52%), Positives = 63/89 (70%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSGN E ++A+++C +N YFA+ LRKAMKG GT D+TLIRV+V+R+E+DM I+ EY
Sbjct: 417 TSGNLEDGMVAVVKCIKNTPAYFAERLRKAMKGAGTKDRTLIRVMVSRSEVDMLDIRQEY 476
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K Y K L I +TSG+Y+ LL L G
Sbjct: 477 LKAYGKSLYTDISGDTSGDYKNLLLKLCG 505
Score = 62.0 bits (149), Expect = 7e-09
Identities = 33/86 (38%), Positives = 56/86 (65%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
+GNFE ++A+L+ P ++ A LR+A+KG GTD+ LI ++ +R+ ++ I Y
Sbjct: 262 TGNFEDLVVAMLK---TPTQFDASELREAIKGAGTDEACLIEILSSRSNAEIIEINKVYK 318
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSL 261
+Y K L ++I S+TSG++R L+SL
Sbjct: 319 AEYGKTLEDSISSDTSGHFRRLLVSL 344
Score = 47.0 bits (110), Expect = 2e-04
Identities = 23/64 (35%), Positives = 38/64 (59%)
Frame = +1
Query: 73 KLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFL 252
++LRKAMKG GTD+K +I ++ RT + A Y Y K L + SE +GN+ +
Sbjct: 210 EVLRKAMKGFGTDEKAIIELLGNRTNKQRVPLVAAYKTTYGKDLFRDLKSELTGNFEDLV 269
Query: 253 LSLV 264
++++
Sbjct: 270 VAML 273
>sw|P27216|ANXD_HUMAN Annexin A13 (Annexin XIII) (Annexin,
intestine-specific) (ISA).
Length = 315
Score = 96.3 bits (238), Expect = 3e-19
Identities = 44/88 (50%), Positives = 70/88 (79%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+ + A L ++RCA++ YFA+ L K+MKG GTD++TLIR+VVTR E+D+Q IKA++
Sbjct: 227 TSGDLQKAYLTLVRCAQDCEDYFAERLYKSMKGAGTDEETLIRIVVTRAEVDLQGIKAKF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+KY+K L++ + S+TSG++R L++L+
Sbjct: 287 QEKYQKSLSDMVRSDTSGDFRKLLVALL 314
Score = 68.9 bits (167), Expect = 6e-11
Identities = 37/87 (42%), Positives = 55/87 (63%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE LA+L + P++Y A+ L+KAMKGLGTD+ LI + TRT ++ IK Y
Sbjct: 72 SGNFEKTALALL---DRPSEYAARQLQKAMKGLGTDESVLIEFLCTRTNKEIIAIKEAYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ + + L + +TSGN + L+SL+
Sbjct: 129 RLFDRSLESDVKGDTSGNLKKILVSLL 155
Score = 52.8 bits (125), Expect = 4e-06
Identities = 27/65 (41%), Positives = 39/65 (60%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
AK L KA KG+GT++ +I ++ RT + Q IK +Y Y K L E + SE SGN+
Sbjct: 19 AKKLNKACKGMGTNEAAIIEILSGRTSDERQQIKQKYKATYGKELEEVLKSELSGNFEKT 78
Query: 250 LLSLV 264
L+L+
Sbjct: 79 ALALL 83
Score = 32.0 bits (71), Expect = 7.5
Identities = 30/97 (30%), Positives = 45/97 (46%), Gaps = 9/97 (9%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENP--------AKYFAKLLRKAMKGL-GTDDKTLIRVVVTRTEI 153
TSGN + L+++L+ N A AK L A +G GTD+ V+ R+
Sbjct: 143 TSGNLKKILVSLLQANRNEGDDVDKDLAGQDAKDLYDAGEGRWGTDELAFNEVLAKRSYK 202
Query: 154 DMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
++ Y K + EAI ETSG+ + L+LV
Sbjct: 203 QLRATFQAYQILIGKDIEEAIEEETSGDLQKAYLTLV 239
>sw|P50994|ANX4_CANFA Annexin A4 (Annexin IV) (Lipocortin IV) (36
kDa zymogen granule membrane associated protein)
(ZAP36).
Length = 318
Score = 95.9 bits (237), Expect = 4e-19
Identities = 50/89 (56%), Positives = 63/89 (70%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+ +
Sbjct: 227 TSGSFEDALLAIVKCMRNKSAYFAERLYKSMKGLGTDDNTLIRVMVSRAEIDMMDIRESF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRLYGKSLYSFIKGDTSGDYRKVLLILCG 315
Score = 53.1 bits (126), Expect = 3e-06
Identities = 31/90 (34%), Positives = 51/90 (56%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE ++ ++ Y + LR+AMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 72 SGNFERVIVGMITPT---VLYDVQELRRAMKGSGTDEGCLIEILASRTPEELRCINQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
+Y + L + I S+TS ++ L+SL G
Sbjct: 129 LQYGRSLEDVIRSDTSFMFQRVLVSLSAGG 158
Score = 51.2 bits (121), Expect = 1e-05
Identities = 25/67 (37%), Positives = 38/67 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I V+ R Q I+ Y + L + + SE SGN+
Sbjct: 19 AQTLRKAMKGLGTDEDAIISVLAPRNTSQRQEIRTAYKSTIGRDLMDDLKSELSGNFERV 78
Query: 250 LLSLVGP 270
++ ++ P
Sbjct: 79 IVGMITP 85
Score = 35.8 bits (81), Expect = 0.52
Identities = 18/57 (31%), Positives = 33/57 (57%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + V+ +R + ++ EY + +K + + I SETSG++ LL++V
Sbjct: 183 KKWGTDEVKFLTVLCSRNRNHLLHVFDEYKRISQKDIEQGIKSETSGSFEDALLAIV 239
>sptr|Q7QM48|Q7QM48 AgCP15598 (Fragment).
Length = 89
Score = 95.5 bits (236), Expect = 6e-19
Identities = 48/87 (55%), Positives = 62/87 (71%)
Frame = +1
Query: 7 GNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFK 186
G+F AL AI+ C + +FAK L KAM GLGTDDKTLIR++VTR EID+Q IK E+ +
Sbjct: 1 GDFYDALSAIVECVQMAPHFFAKKLFKAMDGLGTDDKTLIRIIVTRAEIDLQNIKDEFEQ 60
Query: 187 KYKKPLAEAIHSETSGNYRTFLLSLVG 267
Y K L A+ SETSG+Y+ L +L+G
Sbjct: 61 MYNKTLLSAVKSETSGDYKRVLCALIG 87
>sptr|Q7T391|Q7T391 Hypothetical protein.
Length = 483
Score = 95.1 bits (235), Expect = 7e-19
Identities = 46/88 (52%), Positives = 62/88 (70%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG+ E +LA+++C +N YFA+ L KAMKG GT D+TLIR++VTR+E+DM I+ EY
Sbjct: 393 SGDLESGMLAVVKCIKNTPAYFAERLHKAMKGAGTKDRTLIRIMVTRSEVDMLDIRQEYA 452
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K Y K L AI +TSG+Y+ LL L G
Sbjct: 453 KNYGKSLYTAISGDTSGDYKKLLLKLCG 480
Score = 63.9 bits (154), Expect = 2e-09
Identities = 34/86 (39%), Positives = 57/86 (66%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE +LA+L+ P++Y A L++A+KG GTD+ LI ++ +R+ +++ I +
Sbjct: 237 SGNFEKLVLAMLK---TPSQYDAYELKEAIKGAGTDEACLIEILASRSNAEIREINQVFK 293
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSL 261
+ KK L +AI +TSG++R L+SL
Sbjct: 294 AENKKSLEDAISGDTSGHFRRLLVSL 319
Score = 47.8 bits (112), Expect = 1e-04
Identities = 23/65 (35%), Positives = 40/65 (61%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A++LRKAMKG GTD++ +I ++ +R+ + Y Y K L + + SE SGN+
Sbjct: 184 AEVLRKAMKGFGTDEQAIINLLGSRSNKQRVPLLVSYKTAYGKDLIKDLKSELSGNFEKL 243
Query: 250 LLSLV 264
+L+++
Sbjct: 244 VLAML 248
>sw|P55260|ANX4_RAT Annexin A4 (Annexin IV) (Lipocortin IV) (36 kDa
zymogen granule membrane associated protein) (ZAP36).
Length = 318
Score = 95.1 bits (235), Expect = 7e-19
Identities = 51/89 (57%), Positives = 62/89 (69%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C N YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I A +
Sbjct: 227 TSGSFEDALLAIVKCMRNKPAYFAERLYKSMKGLGTDDSTLIRVMVSRAEIDMLDIPANF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRVYGKSLYSFIKGDTSGDYRKVLLILCG 315
Score = 51.2 bits (121), Expect = 1e-05
Identities = 31/90 (34%), Positives = 50/90 (55%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
S NFE +L ++ Y + LR+AMKG GTD+ LI ++ +R +++ I Y
Sbjct: 72 SSNFEQVILGMMTPT---VLYDVQELRRAMKGAGTDEGCLIEILASRNPEEIRRINQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
++Y + L E I S+TS ++ L+SL G
Sbjct: 129 QQYGRSLEEDICSDTSFMFQRVLVSLTAGG 158
Score = 51.2 bits (121), Expect = 1e-05
Identities = 26/67 (38%), Positives = 38/67 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A++LRKAMKGLGTD+ +I V+ R Q I+ Y + L E + SE S N+
Sbjct: 19 AQVLRKAMKGLGTDEDAIIGVLACRNTAQRQEIRTAYKSTIGRDLLEDLKSELSSNFEQV 78
Query: 250 LLSLVGP 270
+L ++ P
Sbjct: 79 ILGMMTP 85
Score = 35.8 bits (81), Expect = 0.52
Identities = 17/57 (29%), Positives = 34/57 (59%)
Frame = +1
Query: 94 KGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
K GTD+ + ++ +R + ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 183 KRWGTDEVKFLSILCSRNRNHLLHVFDEYKRISQKDIEQSIKSETSGSFEDALLAIV 239
>sptr|Q804G4|Q804G4 Annexin 11a.
Length = 526
Score = 95.1 bits (235), Expect = 7e-19
Identities = 46/88 (52%), Positives = 62/88 (70%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG+ E +LA+++C +N YFA+ L KAMKG GT D+TLIR++VTR+E+DM I+ EY
Sbjct: 436 SGDLESGMLAVVKCIKNTPAYFAERLHKAMKGAGTKDRTLIRIMVTRSEVDMLDIRQEYA 495
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K Y K L AI +TSG+Y+ LL L G
Sbjct: 496 KNYGKSLYTAISGDTSGDYKKLLLKLCG 523
Score = 63.9 bits (154), Expect = 2e-09
Identities = 34/86 (39%), Positives = 57/86 (66%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE +LA+L+ P++Y A L++A+KG GTD+ LI ++ +R+ +++ I +
Sbjct: 280 SGNFEKLVLAMLK---TPSQYDAYELKEAIKGAGTDEACLIEILASRSNAEIREINQVFK 336
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSL 261
+ KK L +AI +TSG++R L+SL
Sbjct: 337 AENKKSLEDAISGDTSGHFRRLLVSL 362
Score = 47.8 bits (112), Expect = 1e-04
Identities = 23/65 (35%), Positives = 40/65 (61%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A++LRKAMKG GTD++ +I ++ +R+ + Y Y K L + + SE SGN+
Sbjct: 227 AEVLRKAMKGFGTDEQAIINLLGSRSNKQRVPLLVSYKTAYGKDLIKDLKSELSGNFEKL 286
Query: 250 LLSLV 264
+L+++
Sbjct: 287 VLAML 291
>sptr|Q7TMN7|Q7TMN7 Annexin A4.
Length = 319
Score = 94.7 bits (234), Expect = 9e-19
Identities = 50/89 (56%), Positives = 63/89 (70%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 228 TSGSFEDALLAIVKCMRSKPSYFAERLYKSMKGLGTDDNTLIRVMVSRAEIDMLDIRASF 287
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 288 KRLYGKSLYSFIKGDTSGDYRKVLLILCG 316
Score = 53.1 bits (126), Expect = 3e-06
Identities = 32/90 (35%), Positives = 51/90 (56%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
S NFE +L ++ Y + LR+AMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 73 SSNFEQVILGLMTPT---VLYDVQELRRAMKGAGTDEGCLIEILASRTPEEIRRINQTYQ 129
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
++Y + L E I S+TS ++ L+SL G
Sbjct: 130 QQYGRSLEEDICSDTSFMFQRVLVSLSAAG 159
Score = 50.8 bits (120), Expect = 2e-05
Identities = 26/67 (38%), Positives = 38/67 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I ++ R Q I++ Y + L E + SE S N+
Sbjct: 20 AQTLRKAMKGLGTDEDAIIGILAYRNTAQRQEIRSAYKSTIGRDLIEDLKSELSSNFEQV 79
Query: 250 LLSLVGP 270
+L L+ P
Sbjct: 80 ILGLMTP 86
Score = 37.0 bits (84), Expect = 0.23
Identities = 24/97 (24%), Positives = 46/97 (47%), Gaps = 9/97 (9%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRK---------AMKGLGTDDKTLIRVVVTRTEI 153
TS F+ L+++ + Y L K K GTD+ + ++ +R
Sbjct: 144 TSFMFQRVLVSLSAAGRDEGNYLDDALMKQDAQELYEAGEKRWGTDEVKFLSILCSRNRN 203
Query: 154 DMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ ++ EY + +K + ++I SETSG++ LL++V
Sbjct: 204 HLLHVFDEYKRISQKDIEQSIKSETSGSFEDALLAIV 240
>sw|P97429|ANX4_MOUSE Annexin A4 (Annexin IV).
Length = 318
Score = 94.7 bits (234), Expect = 9e-19
Identities = 50/89 (56%), Positives = 63/89 (70%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+FE ALLAI++C + YFA+ L K+MKGLGTDD TLIRV+V+R EIDM I+A +
Sbjct: 227 TSGSFEDALLAIVKCMRSKPSYFAERLYKSMKGLGTDDNTLIRVMVSRAEIDMLDIRASF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L I +TSG+YR LL L G
Sbjct: 287 KRLYGKSLYSFIKGDTSGDYRKVLLILCG 315
Score = 50.8 bits (120), Expect = 2e-05
Identities = 31/90 (34%), Positives = 50/90 (55%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
S NFE +L ++ Y + LR+AMKG GTD+ LI ++ +RT +++ I Y
Sbjct: 72 SSNFEQVILGLMTPT---VLYDVQELRRAMKGAGTDEGCLIEILASRTPEEIRRINQTYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
++Y + L E I S+TS ++ L+ L G
Sbjct: 129 QQYGRSLEEDICSDTSFMFQRVLVFLSAAG 158
Score = 50.8 bits (120), Expect = 2e-05
Identities = 26/67 (38%), Positives = 38/67 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A+ LRKAMKGLGTD+ +I ++ R Q I++ Y + L E + SE S N+
Sbjct: 19 AQTLRKAMKGLGTDEDAIIGILAYRNTAQRQEIRSAYKSTIGRDLIEDLKSELSSNFEQV 78
Query: 250 LLSLVGP 270
+L L+ P
Sbjct: 79 ILGLMTP 85
Score = 36.2 bits (82), Expect = 0.40
Identities = 18/64 (28%), Positives = 36/64 (56%)
Frame = +1
Query: 73 KLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFL 252
+L K GTD+ + ++ +R + ++ EY + +K + ++I SETSG++ L
Sbjct: 176 ELYEAGEKRWGTDEVKFLSILCSRNRNHLLHVFDEYKRISQKDIEQSIKSETSGSFEDAL 235
Query: 253 LSLV 264
L++V
Sbjct: 236 LAIV 239
>sptr|Q7SZ98|Q7SZ98 Hypothetical protein.
Length = 338
Score = 93.6 bits (231), Expect = 2e-18
Identities = 49/87 (56%), Positives = 62/87 (71%)
Frame = +1
Query: 7 GNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFK 186
G+ E L AI++CA N A +FA+ L KAMKG GT DK LIRV+V+R+EIDM IKA+Y K
Sbjct: 250 GDIENCLTAIVKCASNRAAFFAEKLYKAMKGSGTRDKDLIRVMVSRSEIDMNEIKAQYQK 309
Query: 187 KYKKPLAEAIHSETSGNYRTFLLSLVG 267
Y K L +AI ET G+Y T L++L G
Sbjct: 310 LYGKSLHQAILDETKGDYETILIALCG 336
Score = 62.4 bits (150), Expect = 5e-09
Identities = 34/86 (39%), Positives = 55/86 (63%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG FE +LA+L+ PA++ A L+ A KGLGTD+ TLI ++ +R D++ I Y
Sbjct: 93 SGEFEEVVLALLK---TPAEFDAYELKHATKGLGTDEDTLIEILASRNNKDIREINRVYK 149
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSL 261
+ YK L + + S+TSG+++ L++L
Sbjct: 150 EVYKSELTKDLTSDTSGDFQKALVAL 175
Score = 45.1 bits (105), Expect = 9e-04
Identities = 23/65 (35%), Positives = 34/65 (52%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A +L KA+K G D+ T+I ++ R Q IK Y K KPL E + SG +
Sbjct: 40 AAVLDKAIKAKGVDEATIIDILTKRNNAQRQEIKVAYQKSVGKPLEECLKKALSGEFEEV 99
Query: 250 LLSLV 264
+L+L+
Sbjct: 100 VLALL 104
>sptr|Q7PFU3|Q7PFU3 ENSANGP00000022424 (Fragment).
Length = 308
Score = 91.7 bits (226), Expect = 8e-18
Identities = 45/88 (51%), Positives = 61/88 (69%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG AL AI+ C + +FAK L KAM G+GTDD TLIR++V+R+EID+Q IK E+
Sbjct: 219 SGELYDALSAIVECVQMAPHFFAKRLHKAMDGVGTDDATLIRIIVSRSEIDLQNIKDEFE 278
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L A+ SETSG+Y+ L +L+G
Sbjct: 279 QMYNKTLVSAVRSETSGDYKRVLCALIG 306
Score = 63.5 bits (153), Expect = 2e-09
Identities = 33/86 (38%), Positives = 51/86 (59%)
Frame = +1
Query: 7 GNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFK 186
G FE +L ++ P Y K L KAM G+GTD+K+LI ++ +T ++ I Y +
Sbjct: 64 GKFEDVILGLML---RPEAYLCKQLHKAMDGIGTDEKSLIEIICPQTNDQIRAIVDCYEE 120
Query: 187 KYKKPLAEAIHSETSGNYRTFLLSLV 264
Y +PLAE + SETSG++R L ++
Sbjct: 121 MYSRPLAEHLCSETSGSFRRLLTMII 146
Score = 38.9 bits (89), Expect = 0.062
Identities = 25/65 (38%), Positives = 35/65 (53%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A LRKAMKG GTD++ +I ++ R+ Q I AE FK+ E SE G +
Sbjct: 17 ANALRKAMKGFGTDEQAIIDILCARSNGQRQEI-AEAFKR------ELGRSELGGKFEDV 69
Query: 250 LLSLV 264
+L L+
Sbjct: 70 ILGLM 74
>sptr|Q9FUG5|Q9FUG5 Annexin.
Length = 334
Score = 91.7 bits (226), Expect = 8e-18
Identities = 41/89 (46%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
T G F ++ ++ CA+N Y+AK L ++MKG+GTDD TL R++VT E++M+ IKA +
Sbjct: 225 THGGFRVSVRVVMHCAKNLINYYAKTLYESMKGMGTDDSTLTRIIVTCAELNMKDIKAHF 284
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+KY++PL E I +T G+++TFL+ LVG
Sbjct: 285 SRKYQRPLHEMISLDTMGHFQTFLMLLVG 313
>sptr|Q7QG24|Q7QG24 AgCP13874 (Fragment).
Length = 323
Score = 91.3 bits (225), Expect = 1e-17
Identities = 45/88 (51%), Positives = 61/88 (69%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG AL AI+ C + +FAK L KAM G+GTDD TLIR++V+R+EID+Q IK E+
Sbjct: 234 SGELYDALSAIVECVQMAPHFFAKRLHKAMDGVGTDDATLIRIIVSRSEIDLQNIKDEFE 293
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L A+ SETSG+Y+ L +L+G
Sbjct: 294 QMYNKTLVSAVRSETSGDYKRALCALIG 321
Score = 63.5 bits (153), Expect = 2e-09
Identities = 33/86 (38%), Positives = 51/86 (59%)
Frame = +1
Query: 7 GNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFK 186
G FE +L ++ P Y K L KAM G+GTD+K+LI ++ +T ++ I Y +
Sbjct: 79 GKFEDVILGLML---RPEAYLCKQLHKAMDGIGTDEKSLIEIICPQTNDQIRAIVDCYEE 135
Query: 187 KYKKPLAEAIHSETSGNYRTFLLSLV 264
Y +PLAE + SETSG++R L ++
Sbjct: 136 MYSRPLAEHLCSETSGSFRRLLTMII 161
Score = 43.1 bits (100), Expect = 0.003
Identities = 21/65 (32%), Positives = 36/65 (55%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A LRKAMKG GTD++ +I ++ R+ Q I + ++ + L + + SE G +
Sbjct: 25 ANALRKAMKGFGTDEQAIIDILCARSNGQRQEIAEAFKRELGRDLIDDLKSELGGKFEDV 84
Query: 250 LLSLV 264
+L L+
Sbjct: 85 ILGLM 89
>sw|Q29471|ANXD_CANFA Annexin A13 (Annexin XIII) (Annexin,
intestine-specific) (ISA).
Length = 315
Score = 91.3 bits (225), Expect = 1e-17
Identities = 41/88 (46%), Positives = 67/88 (76%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+ + A L ++RCA + YFA L K+MKG GTD++TLI ++VTR E+D+Q IKA++
Sbjct: 227 TSGDLQKAYLTLVRCARDQEGYFADRLYKSMKGTGTDEETLIHIIVTRAEVDLQGIKAKF 286
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLV 264
+KY+K L++ + S+TSG+++ L++L+
Sbjct: 287 QEKYQKSLSDMVRSDTSGDFQKLLVALL 314
Score = 71.6 bits (174), Expect = 9e-12
Identities = 37/87 (42%), Positives = 57/87 (65%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SGNFE LA+L + P++Y A+ L+KAMKGLGTD+ LI ++ TRT ++ IK Y
Sbjct: 72 SGNFEKTALALL---DRPSEYDARQLQKAMKGLGTDEAVLIEILCTRTNKEIMAIKEAYQ 128
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLV 264
+ + + L + ++TSGN + L+SL+
Sbjct: 129 RLFDRSLESDVKADTSGNLKAILVSLL 155
Score = 53.9 bits (128), Expect = 2e-06
Identities = 27/65 (41%), Positives = 39/65 (60%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
AK L KA KG+GTD+ +I ++ +RT + Q IK +Y Y K L E S+ SGN+
Sbjct: 19 AKKLNKACKGMGTDEAAIIEILSSRTSDERQQIKQKYKATYGKDLEEVFKSDLSGNFEKT 78
Query: 250 LLSLV 264
L+L+
Sbjct: 79 ALALL 83
>sptrnew|AAH60389|AAH60389 MGC68504 protein.
Length = 321
Score = 91.3 bits (225), Expect = 1e-17
Identities = 48/88 (54%), Positives = 62/88 (70%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG+ E +LLAI++C ++ YFA+ L K+MKGLGTDDKTLIRV+V+R EIDM I+ E+
Sbjct: 231 SGHLEDSLLAIVKCIKSRPAYFAERLYKSMKGLGTDDKTLIRVMVSRCEIDMLEIRCEFK 290
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K Y K L I + SG+YR LL L G
Sbjct: 291 KMYGKSLHSFIKGDCSGDYRKVLLKLCG 318
Score = 54.3 bits (129), Expect = 1e-06
Identities = 32/90 (35%), Positives = 52/90 (57%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
+GNFE +L ++ + Y + L+KAMKG GTD+ LI ++ +R+ +++ I Y
Sbjct: 75 TGNFEKVILGLITSS---TLYDVEELKKAMKGAGTDEGCLIEILASRSAEEIKNINITYK 131
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
KY K L + I S+TS ++ L+SL G
Sbjct: 132 IKYGKSLEDDICSDTSFMFQRVLVSLAAGG 161
Score = 47.8 bits (112), Expect = 1e-04
Identities = 25/62 (40%), Positives = 34/62 (54%)
Frame = +1
Query: 79 LRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLS 258
LR AMKG GTD+ +I V+ RT Q IK Y K L + + SE +GN+ +L
Sbjct: 25 LRNAMKGAGTDEDAVIDVIANRTLSQRQEIKTAYKTTVGKDLDDDLKSELTGNFEKVILG 84
Query: 259 LV 264
L+
Sbjct: 85 LI 86
>sptr|Q90X16|Q90X16 Annexin 4.
Length = 321
Score = 91.3 bits (225), Expect = 1e-17
Identities = 48/88 (54%), Positives = 62/88 (70%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG+ E +LLAI++C ++ YFA+ L K+MKGLGTDDKTLIRV+V+R EIDM I+ E+
Sbjct: 231 SGHLEDSLLAIVKCIKSRPAYFAERLYKSMKGLGTDDKTLIRVMVSRCEIDMLEIRCEFK 290
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K Y K L I + SG+YR LL L G
Sbjct: 291 KMYGKSLHSFIKGDCSGDYRKVLLKLCG 318
Score = 54.3 bits (129), Expect = 1e-06
Identities = 32/90 (35%), Positives = 52/90 (57%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
+GNFE +L ++ + Y + L+KAMKG GTD+ LI ++ +R+ +++ I Y
Sbjct: 75 TGNFEKVILGLITSS---TLYDVEELKKAMKGAGTDEGCLIEILASRSAEEIKNINITYK 131
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVGPG 273
KY K L + I S+TS ++ L+SL G
Sbjct: 132 IKYGKSLEDDICSDTSFMFQRVLVSLAAGG 161
Score = 47.8 bits (112), Expect = 1e-04
Identities = 25/62 (40%), Positives = 34/62 (54%)
Frame = +1
Query: 79 LRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTFLLS 258
LR AMKG GTD+ +I V+ RT Q IK Y K L + + SE +GN+ +L
Sbjct: 25 LRNAMKGAGTDEDAVIDVIANRTLSQRQEIKTAYKTTVGKDLDDDLKSELTGNFEKVILG 84
Query: 259 LV 264
L+
Sbjct: 85 LI 86
>sptr|Q804H0|Q804H0 Annexin 1c.
Length = 341
Score = 90.1 bits (222), Expect = 2e-17
Identities = 46/88 (52%), Positives = 60/88 (68%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
SG+ E L+ +++ A N YFA+ L+ AMKGLGTDD TLIR++V+R+EID+ I EY
Sbjct: 252 SGHLEDCLMTLVKAAWNKPAYFAEKLQHAMKGLGTDDNTLIRIIVSRSEIDLLKIMQEYK 311
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSLVG 267
+ Y K L EAI SET G+Y LL L G
Sbjct: 312 RMYGKTLQEAIQSETKGDYEKILLVLCG 339
Score = 50.4 bits (119), Expect = 2e-05
Identities = 29/86 (33%), Positives = 50/86 (58%)
Frame = +1
Query: 4 SGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYF 183
S + E +LA+L P++Y A ++ A+KGLGT + L ++ TR+ ++ +K +
Sbjct: 96 SSHLEDVVLALLM---TPSEYDAFEMKNALKGLGTSENVLSEILGTRSNKEITALKNSFK 152
Query: 184 KKYKKPLAEAIHSETSGNYRTFLLSL 261
+ Y + L E I+S+ GN T LL+L
Sbjct: 153 EVYGEMLEEDINSDVKGNLETALLAL 178
Score = 42.4 bits (98), Expect = 0.006
Identities = 22/65 (33%), Positives = 37/65 (56%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A L+KA++ G D+ T+I V+ ++ Q IKA Y + KPLA+A+ S +
Sbjct: 43 AAKLKKAIETKGVDEATIIEVLAKKSNAQRQQIKAAYQQSAGKPLADALKKALSSHLEDV 102
Query: 250 LLSLV 264
+L+L+
Sbjct: 103 VLALL 107
>sptrnew|AAH65430|AAH65430 Hypothetical protein.
Length = 317
Score = 89.7 bits (221), Expect = 3e-17
Identities = 45/89 (50%), Positives = 64/89 (71%)
Frame = +1
Query: 1 TSGNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEY 180
TSG+ + LLA+++CA + YFA L AMKG GTDD+TLIR++VTR+E+D+ I+AE+
Sbjct: 225 TSGSLQEILLAVVKCARSVPGYFADSLYAAMKGAGTDDQTLIRIMVTRSEVDLLDIRAEF 284
Query: 181 FKKYKKPLAEAIHSETSGNYRTFLLSLVG 267
K++ L + I S+TSG+YR LL L G
Sbjct: 285 RKRFATSLHKMIQSDTSGDYRKTLLLLCG 313
Score = 54.3 bits (129), Expect = 1e-06
Identities = 28/86 (32%), Positives = 50/86 (58%)
Frame = +1
Query: 7 GNFEFALLAILRCAENPAKYFAKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFK 186
G FE ++A++ P Y LR A+KG GTD+K LI ++ +R+ ++ IK+ Y +
Sbjct: 73 GKFEDLIVALMT---PPIIYEVTCLRNAIKGAGTDEKVLIEILASRSPNEVNEIKSSYKR 129
Query: 187 KYKKPLAEAIHSETSGNYRTFLLSLV 264
++ K L E + +T G++ L+ L+
Sbjct: 130 EHDKDLEEDVTGDTGGHFERMLVVLL 155
Score = 49.3 bits (116), Expect = 5e-05
Identities = 24/67 (35%), Positives = 41/67 (61%)
Frame = +1
Query: 70 AKLLRKAMKGLGTDDKTLIRVVVTRTEIDMQYIKAEYFKKYKKPLAEAIHSETSGNYRTF 249
A++L KAMKGLGTD+ ++++++ R+ Q IKA Y + K L + SE G +
Sbjct: 19 AEVLYKAMKGLGTDEDSILQLLTKRSNGQRQEIKAAYKTLHGKDLVNDLKSELGGKFEDL 78
Query: 250 LLSLVGP 270
+++L+ P
Sbjct: 79 IVALMTP 85
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 532,512,272
Number of Sequences: 1395590
Number of extensions: 10790422
Number of successful extensions: 24416
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 23452
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 24389
length of database: 442,889,342
effective HSP length: 119
effective length of database: 276,814,132
effective search space used: 28235041464
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

