BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
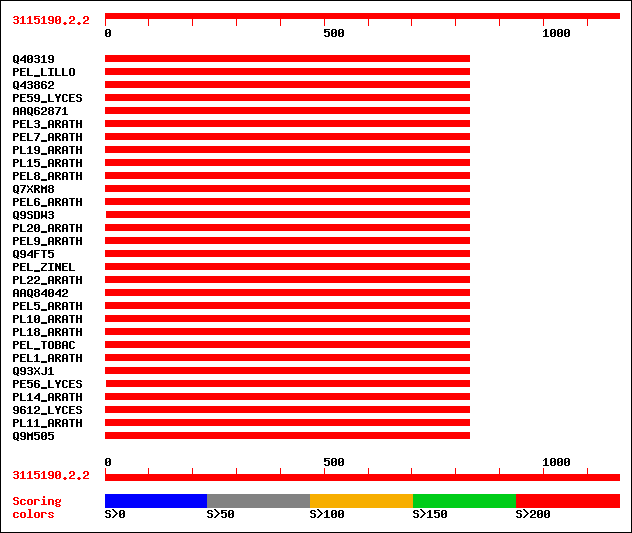
Query= 3115190.2.2
(1172 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q40319|Q40319 Pectate lyase homolog. 385 e-106
sw|P40973|PEL_LILLO Pectate lyase precursor (EC 4.2.2.2). 372 e-102
sptr|Q43862|Q43862 Pectate lyase homolog (EC 4.2.2.2). 365 e-100
sw|P15722|PE59_LYCES Probable pectate lyase P59 precursor (EC 4.... 360 2e-98
sptrnew|AAQ62871|AAQ62871 At1g14420. 350 2e-95
sw|Q9M9S2|PEL3_ARATH Probable pectate lyase 3 precursor (EC 4.2.... 350 2e-95
sw|Q9SRH4|PEL7_ARATH Probable pectate lyase 7 precursor (EC 4.2.... 349 5e-95
sw|Q9LFP5|PL19_ARATH Putative pectate lyase 19 precursor (EC 4.2... 347 2e-94
sw|Q944R1|PL15_ARATH Probable pectate lyase 15 precursor (EC 4.2... 336 5e-91
sw|Q9M8Z8|PEL8_ARATH Probable pectate lyase 8 precursor (EC 4.2.... 335 6e-91
sptr|Q7XRM8|Q7XRM8 OSJNBa0095E20.8 protein. 335 8e-91
sw|O64510|PEL6_ARATH Probable pectate lyase 6 precursor (EC 4.2.... 335 8e-91
sptr|Q9SDW3|Q9SDW3 Pectate lyase 2. 333 4e-90
sw|Q93WF1|PL20_ARATH Probable pectate lyase 20 precursor (EC 4.2... 330 3e-89
sw|Q9LRM5|PEL9_ARATH Putative pectate lyase 9 precursor (EC 4.2.... 330 3e-89
sptr|Q94FT5|Q94FT5 Pectate lyase (Fragment). 328 8e-89
sw|O24554|PEL_ZINEL Pectate lyase precursor (EC 4.2.2.2) (ZePel). 328 1e-88
sw|Q93Z25|PL22_ARATH Probable pectate lyase 22 precursor (EC 4.2... 327 3e-88
sptrnew|AAQ84042|AAQ84042 Pectate lyase. 325 8e-88
sw|Q9FXD8|PEL5_ARATH Probable pectate lyase 5 precursor (EC 4.2.... 324 1e-87
sw|Q9LJ42|PL10_ARATH Probable pectate lyase 10 precursor (EC 4.2... 323 4e-87
sw|Q9C5M8|PL18_ARATH Probable pectate lyase 18 precursor (EC 4.2... 323 4e-87
sw|P40972|PEL_TOBAC Pectate lyase precursor (EC 4.2.2.2). 319 5e-86
sw|Q940Q1|PEL1_ARATH Probable pectate lyase 1 precursor (EC 4.2.... 319 6e-86
sptr|Q93XJ1|Q93XJ1 Pectate lyase. 318 1e-85
sw|P15721|PE56_LYCES Probable pectate lyase P56 precursor (EC 4.... 318 1e-85
sw|Q9SVQ6|PL14_ARATH Putative pectate lyase 14 precursor (EC 4.2... 315 7e-85
sw|P24396|9612_LYCES Style development-specific protein 9612 pre... 314 1e-84
sw|Q9LTZ0|PL11_ARATH Putative pectate lyase 11 precursor (EC 4.2... 313 3e-84
sptr|Q9M505|Q9M505 Pectate lyase. 313 3e-84
>sptr|Q40319|Q40319 Pectate lyase homolog.
Length = 450
Score = 385 bits (990), Expect = e-106
Identities = 181/280 (64%), Positives = 220/280 (78%), Gaps = 3/280 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF RSM+I L+QEL++SSDKTIDGRGA V I +GAGIT+Q NVIIH L + ++K G
Sbjct: 171 IFQRSMVITLTQELMVSSDKTIDGRGANVQIRDGAGITMQFVNNVIIHGLRIKNIKARNG 230
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+RDS HIG RTR+DGD IS+F ++N+WIDHIS+S+CEDGL+DV+Q ST +TISNCH
Sbjct: 231 GLIRDSFDHIGVRTRSDGDAISVFGSSNIWIDHISLSDCEDGLVDVIQGSTAVTISNCHM 290
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T HNDVMLFGASD++ D+IMQITVAFNHFG+GL+QRMPRCRWGFFHV+NNDYTHW+MYA
Sbjct: 291 TKHNDVMLFGASDTYQDDKIMQITVAFNHFGQGLIQRMPRCRWGFFHVLNNDYTHWIMYA 350
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVIT-KHYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG APTI+SQGNR+IAP N AAK +T + YA E VW W W +E D FMNGA F SG
Sbjct: 351 IGGSSAPTILSQGNRFIAPHNNAAKTVTHRDYAPESVWSKWQWRSEGDHFMNGATFIQSG 410
Query: 718 GAPKQVDTNE--WVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
K + + +KP+ G+ RLTRFSG L+C G+PC
Sbjct: 411 PPIKNLPFKKGFLMKPRHGSQANRLTRFSGALNCVVGRPC 450
>sw|P40973|PEL_LILLO Pectate lyase precursor (EC 4.2.2.2).
Length = 434
Score = 372 bits (956), Expect = e-102
Identities = 174/279 (62%), Positives = 213/279 (76%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF +SM+I+L QEL++++DKTIDGRGA V IA GA +TVQ NVIIH +H+HD+K G
Sbjct: 156 IFGKSMVIRLKQELIINNDKTIDGRGANVQIAGGAQLTVQFVHNVIIHGIHIHDIKPGEG 215
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+RDS H G RTR+DGDGIS+ ++N+WIDH+S++ C DGLIDV+ ST ITISNCH
Sbjct: 216 GLIRDSEKHSGIRTRSDGDGISIIGSSNIWIDHVSLARCSDGLIDVILGSTAITISNCHL 275
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T H+DVML GASD++ QD+IMQ+TVAFNHFGRGLVQRMPRCR+GF HVVNNDYTHW+MYA
Sbjct: 276 TEHDDVMLLGASDTYTQDEIMQVTVAFNHFGRGLVQRMPRCRYGFVHVVNNDYTHWIMYA 335
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKH-YAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
+GG PTIISQGNRYIAP AAK +TK YAE W W W ++ DLF++GA F SG
Sbjct: 336 VGGSQHPTIISQGNRYIAPHIEAAKEVTKRDYAEPAEWSKWTWKSQGDLFVSGAFFVESG 395
Query: 718 GA-PKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G + + +K KPGT+V RLTRFSG L+C C
Sbjct: 396 GPFENKYSKKDLIKAKPGTFVQRLTRFSGALNCKENMEC 434
>sptr|Q43862|Q43862 Pectate lyase homolog (EC 4.2.2.2).
Length = 438
Score = 365 bits (936), Expect = e-100
Identities = 175/279 (62%), Positives = 217/279 (77%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+FAR M+I+L QEL+++ +KTIDGRGAQVHI A IT+Q QNVI+HNLH+HD K G
Sbjct: 161 VFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITLQNVQNVILHNLHIHDSKAHSG 219
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G++RDS H G RTR+DGDG+S+ S++NVWIDH+SMS+C DGLIDVV ST IT+SN HF
Sbjct: 220 GMIRDSKRHYGLRTRSDGDGVSVLSSSNVWIDHVSMSSCSDGLIDVVNGSTAITVSNSHF 279
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VMLFGAS+ PQD +MQ+TVAFNHFGRGLVQRMPRCR+GFFHVVNNDYTHW+MYA
Sbjct: 280 TDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHWIMYA 339
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTIISQGNR+IAP + AK +TK Y +K WVW ++ D+ MNGA FN SG
Sbjct: 340 IGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAFFNESG 399
Query: 718 GA-PKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G ++ D +++ K G YV +LTRF+G L C G+PC
Sbjct: 400 GQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 438
>sw|P15722|PE59_LYCES Probable pectate lyase P59 precursor (EC
4.2.2.2).
Length = 449
Score = 360 bits (925), Expect = 2e-98
Identities = 172/281 (61%), Positives = 206/281 (73%), Gaps = 4/281 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M I+L QE++M SDKTID RG VHI GAGIT+Q +NVIIH LH+HD+ G
Sbjct: 169 IFKRGMNIRLHQEMIMQSDKTIDARGVNVHITKGAGITLQYIKNVIIHGLHIHDIVEGNG 228
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G++RD+ HIG RT++DGDGIS+F A+ +WIDH+SM C DGLID V+ STGITISN HF
Sbjct: 229 GMVRDAVDHIGIRTKSDGDGISIFGASYIWIDHVSMQRCYDGLIDAVEGSTGITISNGHF 288
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VMLFGASDS DQ+MQIT+AFNHFG+ L+QRMPRCRWG+ HVVNNDYTHW MYA
Sbjct: 289 TDHNEVMLFGASDSSSIDQVMQITLAFNHFGKRLIQRMPRCRWGYIHVVNNDYTHWNMYA 348
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTII QGNR+IAPP+I K +TK Y E VW W W +E +LFMNGA F SG
Sbjct: 349 IGGSMHPTIIHQGNRFIAPPDIFKKQVTKREYNPESVWMQWTWRSEGNLFMNGAYFTESG 408
Query: 718 G---APKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
+ K D + + P VT +TRF+G L C GKPC
Sbjct: 409 DPEWSSKHKDLYDGISAAPAEDVTWMTRFAGVLGCKPGKPC 449
>sptrnew|AAQ62871|AAQ62871 At1g14420.
Length = 459
Score = 350 bits (898), Expect = 2e-95
Identities = 174/285 (61%), Positives = 210/285 (73%), Gaps = 8/285 (2%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IFARSMII+L QEL+++SDKTIDGRGA+V+I GAG+T+Q NVIIHN++V + G
Sbjct: 175 IFARSMIIKLQQELMITSDKTIDGRGARVYIMEGAGLTLQFVNNVIIHNIYVKHIVPGNG 234
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+RDS HIG RT++DGDGISLF ATN+WIDH+SM+ C DG+ID + ST +TISN HF
Sbjct: 235 GLIRDSEAHIGLRTKSDGDGISLFGATNIWIDHVSMTRCADGMIDAIDGSTAVTISNSHF 294
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H +VMLFGA D D+ MQITVAFNHFG+ L QRMPRCR+G HVVNNDYTHW MYA
Sbjct: 295 TDHQEVMLFGARDEHVIDKKMQITVAFNHFGKRLEQRMPRCRYGTIHVVNNDYTHWEMYA 354
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTIISQGNR+IAPPN AK ITK Y G WK+W W +E D F+NGA F SG
Sbjct: 355 IGGNMNPTIISQGNRFIAPPNEEAKQITKREYTPYGEWKSWNWQSEGDYFLNGAYFVQSG 414
Query: 718 GA------PKQVDTNEW-VKPKPGTYVTRLTRFSGTLSCCTGKPC 831
A PK N++ ++PKPGT V +LT +G L C G+ C
Sbjct: 415 KANAWSSKPKTPLPNKFTIRPKPGTMVRKLTMDAGVLGCKLGEAC 459
>sw|Q9M9S2|PEL3_ARATH Probable pectate lyase 3 precursor (EC 4.2.2.2)
(Pectate lyase A2).
Length = 459
Score = 350 bits (898), Expect = 2e-95
Identities = 174/285 (61%), Positives = 210/285 (73%), Gaps = 8/285 (2%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IFARSMII+L QEL+++SDKTIDGRGA+V+I GAG+T+Q NVIIHN++V + G
Sbjct: 175 IFARSMIIKLQQELMITSDKTIDGRGARVYIMEGAGLTLQFVNNVIIHNIYVKHIVPGNG 234
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+RDS HIG RT++DGDGISLF ATN+WIDH+SM+ C DG+ID + ST +TISN HF
Sbjct: 235 GLIRDSEAHIGLRTKSDGDGISLFGATNIWIDHVSMTRCADGMIDAIDGSTAVTISNSHF 294
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H +VMLFGA D D+ MQITVAFNHFG+ L QRMPRCR+G HVVNNDYTHW MYA
Sbjct: 295 TDHQEVMLFGARDEHVIDKKMQITVAFNHFGKRLEQRMPRCRYGTIHVVNNDYTHWEMYA 354
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTIISQGNR+IAPPN AK ITK Y G WK+W W +E D F+NGA F SG
Sbjct: 355 IGGNMNPTIISQGNRFIAPPNEEAKQITKREYTPYGEWKSWNWQSEGDYFLNGAYFVQSG 414
Query: 718 GA------PKQVDTNEW-VKPKPGTYVTRLTRFSGTLSCCTGKPC 831
A PK N++ ++PKPGT V +LT +G L C G+ C
Sbjct: 415 KANAWSSKPKTPLPNKFTIRPKPGTMVRKLTMDAGVLGCKLGEAC 459
>sw|Q9SRH4|PEL7_ARATH Probable pectate lyase 7 precursor (EC 4.2.2.2).
Length = 475
Score = 349 bits (895), Expect = 5e-95
Identities = 165/281 (58%), Positives = 203/281 (72%), Gaps = 4/281 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F M I+LSQEL+++SDKTID RGA VHIA GAGIT+Q N+IIH LHVH + + G
Sbjct: 195 VFKHDMSIRLSQELMITSDKTIDARGANVHIAYGAGITMQYVHNIIIHGLHVHHIVKSSG 254
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+RDS H G R ADGDGIS+F ATN+W+DHISMS C+DGLID + ST ITISN HF
Sbjct: 255 GLIRDSINHFGHRGEADGDGISIFGATNIWLDHISMSKCQDGLIDAIMGSTAITISNSHF 314
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HNDVML GA ++ D+ MQ+TVA+NHFG+GLVQRMPR RWGF HVVNNDYTHW +YA
Sbjct: 315 THHNDVMLLGAQNNNMDDKKMQVTVAYNHFGKGLVQRMPRVRWGFVHVVNNDYTHWELYA 374
Query: 541 IGGGDAPTIISQGNRYIAPPNIA--AKVITKHYAEEGVWKNWVWHTEDDLFMNGAIFNPS 714
IGG PTI+S GNR+IAPP+ +V + YA E WKNW W +E D+FMN A F S
Sbjct: 375 IGGSQGPTILSHGNRFIAPPHKQHYREVTKRDYASESEWKNWNWRSEKDVFMNNAYFRQS 434
Query: 715 GGAPKQV--DTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G + + +KPK G V++LT+++G L C GK C
Sbjct: 435 GNPHFKCSHSRQQMIKPKNGMAVSKLTKYAGALDCRVGKAC 475
>sw|Q9LFP5|PL19_ARATH Putative pectate lyase 19 precursor (EC
4.2.2.2).
Length = 472
Score = 347 bits (891), Expect = 2e-94
Identities = 160/281 (56%), Positives = 208/281 (74%), Gaps = 4/281 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF M I+L+QELL++S KTID RGA VH+A+GAGIT+Q +NVIIH LH+H + + G
Sbjct: 192 IFKNDMSIRLNQELLINSHKTIDARGANVHVAHGAGITMQFVKNVIIHGLHIHHISESSG 251
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G++RDS H G RTRADGDG+S++ ++N+W+DHISMS C+DGLID + STGITISN HF
Sbjct: 252 GMIRDSVDHFGMRTRADGDGLSIYGSSNIWLDHISMSKCQDGLIDAIVGSTGITISNSHF 311
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HNDVML GA ++ D+ MQ+TVA+NHFG+GLVQRMPR RWGF HVVNNDYTHW +YA
Sbjct: 312 THHNDVMLLGAQNTNEADKHMQVTVAYNHFGKGLVQRMPRIRWGFVHVVNNDYTHWELYA 371
Query: 541 IGGGDAPTIISQGNRYIAPPNIA--AKVITKHYAEEGVWKNWVWHTEDDLFMNGAIFNPS 714
IGG PTI+S GNR+IAPP+ +V + YA E WK+W W ++ D+FMNGA F S
Sbjct: 372 IGGSQGPTILSHGNRFIAPPHKPHYREVTKRDYASEDEWKHWNWRSDKDVFMNGAYFRQS 431
Query: 715 GGAPKQV--DTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G + + +KPK G V++LT+++G L C G+ C
Sbjct: 432 GNPQYKCAHTRQQMIKPKNGLAVSKLTKYAGALDCRVGRRC 472
>sw|Q944R1|PL15_ARATH Probable pectate lyase 15 precursor (EC 4.2.2.2)
(Pectate lyase A11).
Length = 470
Score = 336 bits (861), Expect = 5e-91
Identities = 158/279 (56%), Positives = 199/279 (71%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+I+L QEL+M+S KTID RG+ VHIANGA IT+Q NVIIH LH+HD K T
Sbjct: 192 VFKRDMVIELKQELIMNSFKTIDARGSNVHIANGACITIQFITNVIIHGLHIHDCKPTGN 251
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G RT ADGD +S+F ++++WIDH S+S+C DGL+D V ST IT+SN HF
Sbjct: 252 AMVRSSPSHFGWRTMADGDAVSIFGSSHIWIDHNSLSHCADGLVDAVMGSTAITVSNNHF 311
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D++MQ+T+A+NHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 312 THHNEVMLLGHSDSYTKDKLMQVTIAYNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 371
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNRY AP + AK +TK + WK W W +E DL +NGA F PSG
Sbjct: 372 IGGSAEPTINSQGNRYAAPMDRFAKEVTKRVETDASEWKKWNWRSEGDLLLNGAFFRPSG 431
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + KP + V +T +G L C G+PC
Sbjct: 432 AGASASYGRASSLAAKPSSMVDTITSTAGALGCRKGRPC 470
>sw|Q9M8Z8|PEL8_ARATH Probable pectate lyase 8 precursor (EC
4.2.2.2).
Length = 416
Score = 335 bits (860), Expect = 6e-91
Identities = 158/279 (56%), Positives = 201/279 (72%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+I L QEL+M+S KTIDGRG VHIANGA +T+Q N+I+H +HVHD K T
Sbjct: 138 IFKRDMVITLKQELIMNSFKTIDGRGVNVHIANGACLTIQYVTNIIVHGIHVHDCKPTGN 197
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G R+ ADGD IS+F ++++WIDH S+SNC DGL+D V SST IT+SN F
Sbjct: 198 AMVRSSPSHYGFRSMADGDAISIFGSSHIWIDHNSLSNCADGLVDAVMSSTAITVSNNFF 257
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D++MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 258 THHNEVMLLGHSDSYTRDKVMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYA 317
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP N AK +TK Y E WK+W W +E DLF+NGA F SG
Sbjct: 318 IGGSAGPTINSQGNRFLAPVNPFAKEVTKREYTGESKWKHWNWRSEGDLFLNGAFFTRSG 377
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + K + V +T +SG L+C G+ C
Sbjct: 378 AGAGANYARASSLSAKSSSLVGTMTSYSGALNCRAGRRC 416
>sptr|Q7XRM8|Q7XRM8 OSJNBa0095E20.8 protein.
Length = 472
Score = 335 bits (859), Expect = 8e-91
Identities = 159/279 (56%), Positives = 198/279 (70%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+I L QEL+M+S KTIDGRGA VHIANGA IT+Q NVIIH LH+HD + T
Sbjct: 194 VFKRDMVITLKQELIMNSFKTIDGRGANVHIANGACITIQYVTNVIIHGLHIHDCRPTGN 253
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G RT ADGD +S+F A+++W+DH S+SNC DGLID + ST IT+SN +F
Sbjct: 254 AMVRSSPSHYGWRTMADGDAVSIFGASHIWVDHCSLSNCADGLIDAIMGSTAITVSNNYF 313
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D+ MQ+T+AFNHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 314 THHNEVMLLGHSDSYVKDKAMQVTIAFNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYA 373
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNRY+AP N AK +TK + +WK W W +E DL +NGA F PSG
Sbjct: 374 IGGSAEPTINSQGNRYLAPTNPFAKEVTKRVETAQTIWKGWNWRSEGDLLLNGAFFTPSG 433
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + K + V +T +G LSC G C
Sbjct: 434 AGASASYSRASSLGAKSSSMVGTITSGAGALSCRGGSAC 472
>sw|O64510|PEL6_ARATH Probable pectate lyase 6 precursor (EC 4.2.2.2).
Length = 455
Score = 335 bits (859), Expect = 8e-91
Identities = 165/285 (57%), Positives = 209/285 (73%), Gaps = 8/285 (2%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IFARSMII+L QEL++++DKTIDGRGA+++I GAG+T+Q +NVIIHN+H+ +K G
Sbjct: 171 IFARSMIIKLQQELIITNDKTIDGRGAKIYITGGAGLTLQFVRNVIIHNIHIKQIKRGAG 230
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+ DS H G RT +DGDGI++F ATNVWIDH+SM++C DG+ID + ST ITISN HF
Sbjct: 231 GLIIDSEQHFGLRTVSDGDGINIFGATNVWIDHVSMTDCSDGMIDAIMGSTAITISNSHF 290
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H++VMLFG ++ D+ MQITVAFNHFG+ L QRMPR R+G HVVNNDYTHW MYA
Sbjct: 291 TDHDEVMLFGGTNKDVIDKKMQITVAFNHFGKRLKQRMPRVRFGLVHVVNNDYTHWEMYA 350
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITK-HYAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTIISQGNR+IAPP +K +TK Y WK+W W +E D F+NGA F SG
Sbjct: 351 IGGNMNPTIISQGNRFIAPPIEDSKQVTKREYTPYPEWKSWNWQSEKDYFLNGAYFVQSG 410
Query: 718 GA------PKQVDTNEW-VKPKPGTYVTRLTRFSGTLSCCTGKPC 831
A PK ++ ++P+PGT V RLT+ +GTL C GK C
Sbjct: 411 KANAWSATPKNPIPRKFAIRPQPGTKVRRLTKDAGTLGCKPGKSC 455
>sptr|Q9SDW3|Q9SDW3 Pectate lyase 2.
Length = 454
Score = 333 bits (853), Expect = 4e-90
Identities = 159/278 (57%), Positives = 195/278 (70%), Gaps = 2/278 (0%)
Frame = +1
Query: 4 FARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMGG 183
F M I L +EL+M+S KTIDGRG VHIANGA IT+Q NVIIH LH+HD K T
Sbjct: 177 FKHDMEITLKEELIMNSFKTIDGRGVNVHIANGACITIQYITNVIIHGLHIHDCKPTGNA 236
Query: 184 LMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHFT 363
++R SP+H G RT ADGD +S+F ++++W+DH S+SNC DGL+D V ST IT+SN +FT
Sbjct: 237 MVRSSPSHYGWRTMADGDAVSIFGSSHIWVDHCSLSNCADGLVDAVMGSTAITVSNNYFT 296
Query: 364 NHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYAI 543
+HN+VML G +DS+ +D IMQ+T+AFNHFG GL+QRMPRCR G+FHVVNNDYTHW MYAI
Sbjct: 297 HHNEVMLLGHTDSYARDSIMQVTIAFNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYAI 356
Query: 544 GGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG- 717
GG PTI SQGNRY+AP N AK +TK ++ WKNW W +E DL +NGA F PSG
Sbjct: 357 GGSANPTINSQGNRYLAPTNPFAKEVTKRVDTDQSTWKNWNWRSEGDLLLNGAFFTPSGA 416
Query: 718 GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA KP + V LT +G LSC G C
Sbjct: 417 GASASYARASSFGAKPSSLVDTLTSDAGVLSCQVGTRC 454
>sw|Q93WF1|PL20_ARATH Probable pectate lyase 20 precursor (EC
4.2.2.2).
Length = 417
Score = 330 bits (845), Expect = 3e-89
Identities = 156/279 (55%), Positives = 203/279 (72%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+I L +EL+M+S KTIDGRG VHIANGA IT+Q N+IIH +H+HD + T
Sbjct: 139 VFKRDMVITLKEELIMNSFKTIDGRGVNVHIANGACITIQFVTNIIIHGIHIHDCRPTGN 198
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G RT ADGDGIS+F ++++WIDH S+SNC DGLID V +ST ITISN +F
Sbjct: 199 AMVRSSPSHYGWRTMADGDGISIFGSSHIWIDHNSLSNCADGLIDAVMASTAITISNNYF 258
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SD++ +D++MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 259 THHNEVMLLGHSDTYTRDKVMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYA 318
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKH-YAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG +PTI SQGNRY+AP N AK +TK YA + W++W W +E DLF+NGA F SG
Sbjct: 319 IGGSASPTINSQGNRYLAPRNRFAKEVTKRDYAGQWQWRHWNWRSEGDLFLNGAFFTRSG 378
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G + K + V +T +G L+C G+ C
Sbjct: 379 SGLGASYARASSLAAKSSSLVGVITYNAGALNCRGGRRC 417
>sw|Q9LRM5|PEL9_ARATH Putative pectate lyase 9 precursor (EC 4.2.2.2).
Length = 452
Score = 330 bits (845), Expect = 3e-89
Identities = 158/278 (56%), Positives = 197/278 (70%), Gaps = 1/278 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+I+L QEL+M+S KTID RGA VHIANGA IT+Q NVI+H LH+HD K T
Sbjct: 174 IFKRDMVIKLKQELIMNSFKTIDARGANVHIANGACITIQNITNVIVHGLHIHDCKRTGN 233
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
+R SP+ G R ADGD I++F ++++WIDH S+SNC DGL+DVV ST ITISN HF
Sbjct: 234 VTVRSSPSQAGFRGTADGDAINIFGSSHIWIDHNSLSNCTDGLVDVVNGSTAITISNNHF 293
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H++VML G +DS+ +D++MQ+TVA+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 294 THHDEVMLLGHNDSYTRDKMMQVTVAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWKMYA 353
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHYAEEG-VWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR+ AP N +AK +TK +G W W W +E DL +NGA F PSG
Sbjct: 354 IGGSANPTINSQGNRFAAPKNHSAKEVTKRLDTKGNEWMEWNWRSEKDLLVNGAFFTPSG 413
Query: 718 GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
+ KP + V +T +G LSC GKPC
Sbjct: 414 EGASGDSQTLSLPAKPASMVDAITASAGALSCRRGKPC 451
>sptr|Q94FT5|Q94FT5 Pectate lyase (Fragment).
Length = 368
Score = 328 bits (842), Expect = 8e-89
Identities = 157/279 (56%), Positives = 194/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+IQL QEL+M+S KTIDGRG VHIANGA IT+Q NVI+H LH+HD K T
Sbjct: 90 VFKRDMVIQLKQELIMNSFKTIDGRGVNVHIANGACITIQFVTNVIVHGLHIHDCKPTGN 149
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G RT ADGD IS+F ++++W+DH S+SNC DGL+D V ST ITISN H
Sbjct: 150 AMVRSSPSHFGWRTMADGDAISIFGSSHIWVDHNSLSNCADGLVDAVMGSTAITISNNHL 209
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D+ MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 210 THHNEVMLLGHSDSYTRDKQMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYA 269
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNRY AP N AK +TK + W+ W W +E DL +NGA F PSG
Sbjct: 270 IGGSADPTINSQGNRYAAPTNPFAKEVTKRVETSQTQWRGWNWRSEGDLLLNGAFFTPSG 329
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + K V +T +G L C G+ C
Sbjct: 330 AGASAVYARASSLGAKSSAMVGTITASAGALGCRRGRTC 368
>sw|O24554|PEL_ZINEL Pectate lyase precursor (EC 4.2.2.2) (ZePel).
Length = 401
Score = 328 bits (840), Expect = 1e-88
Identities = 159/279 (56%), Positives = 195/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+IQL QEL+M+S KTIDGRG VHI NG IT+ A N+IIH +H+HD K
Sbjct: 123 IFKRDMVIQLRQELVMNSHKTIDGRGVNVHIGNGPCITIHYASNIIIHGIHIHDCKQAGN 182
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G +R+SP H G T++DGDGIS+F++ ++WIDH S+SNC DGLID + ST ITISN +
Sbjct: 183 GNIRNSPHHSGWWTQSDGDGISIFASKDIWIDHNSLSNCHDGLIDAIHGSTAITISNNYM 242
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SDS+ QD+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 243 THHDKVMLLGHSDSYTQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 302
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKH-YAEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG +PTI SQGNR++AP K +TKH A E WKNW W +E DL +NGA F SG
Sbjct: 303 IGGSASPTIYSQGNRFLAPNTRFDKEVTKHENAPESEWKNWNWRSEGDLMLNGAYFRESG 362
Query: 718 G-APKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G A + +P T V +TR +G L C G C
Sbjct: 363 GRAASSFARASSLSGRPSTLVASMTRSAGALVCRKGSRC 401
>sw|Q93Z25|PL22_ARATH Probable pectate lyase 22 precursor (EC
4.2.2.2).
Length = 432
Score = 327 bits (837), Expect = 3e-88
Identities = 156/281 (55%), Positives = 198/281 (70%), Gaps = 4/281 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M IQL +EL+M+S KT+DGRGA VHI+ G IT+Q N+IIH LH+HD K
Sbjct: 152 IFKRDMTIQLKEELIMNSFKTLDGRGASVHISGGPCITIQYVTNIIIHGLHIHDCKQGGN 211
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
+RDSP H G RT +DGDG+S+F ++VW+DH S+SNC DGLID ++ ST ITISN +
Sbjct: 212 TYVRDSPEHYGYRTVSDGDGVSIFGGSHVWVDHCSLSNCNDGLIDAIRGSTAITISNNYL 271
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN VML G SD++ QD+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 272 THHNKVMLLGHSDTYEQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 331
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP + ++K +TKH A E W+NW W +E DL +NGA F SG
Sbjct: 332 IGGSANPTINSQGNRFLAPDDSSSKEVTKHEDAPEDEWRNWNWRSEGDLLLNGAFFTYSG 391
Query: 718 GAPKQVDT---NEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
P + + + +P ++V +T SG LSC G C
Sbjct: 392 AGPAKSSSYSKASSLAARPSSHVGEITIASGALSCKRGSHC 432
>sptrnew|AAQ84042|AAQ84042 Pectate lyase.
Length = 418
Score = 325 bits (833), Expect = 8e-88
Identities = 159/279 (56%), Positives = 193/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M IQL +EL+M+S KTIDGRGA VHIA G IT+Q N+IIH LH+HD K
Sbjct: 140 IFQRDMTIQLKEELIMNSFKTIDGRGASVHIAGGPCITIQFVTNIIIHGLHIHDCKQGGN 199
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP H G RT +DGDG+S+F ++VW+DH S+SNC+DGL+D + ST ITISN +
Sbjct: 200 AMVRSSPRHFGWRTVSDGDGVSIFGGSHVWVDHCSLSNCKDGLVDAIYGSTAITISNNYM 259
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SDS+ D+ MQIT+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 260 THHDKVMLLGHSDSYTNDKNMQITIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 319
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR+ AP ++K +TKH A E WKNW W +E DL +NGA F SG
Sbjct: 320 IGGSADPTINSQGNRFAAPDIRSSKEVTKHEDAPESEWKNWNWRSEGDLMLNGAFFTASG 379
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + KP + V +T SG LSC G C
Sbjct: 380 AGASSSYARASSLGAKPSSLVGAITTASGALSCRKGSRC 418
>sw|Q9FXD8|PEL5_ARATH Probable pectate lyase 5 precursor (EC
4.2.2.2).
Length = 408
Score = 324 bits (831), Expect = 1e-87
Identities = 156/279 (55%), Positives = 196/279 (70%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IFAR M+I+L +EL+M+S KTIDGRGA VHIA GA ITVQ N+IIH +++HD K
Sbjct: 130 IFARDMVIKLKEELIMNSFKTIDGRGASVHIAGGACITVQYVTNIIIHGVNIHDCKRKGN 189
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
+RDSP+H G RT +DGD +S+F ++VW+DH S+SNC DGLID + ST ITISN +
Sbjct: 190 AYVRDSPSHYGWRTASDGDAVSIFGGSHVWVDHCSLSNCADGLIDAIHGSTAITISNNYL 249
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
++HN VML G SDS+ +D+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 250 SHHNKVMLLGHSDSYTRDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWQMYA 309
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPS- 714
IGG APTI SQGNR++AP + K +TK+ A WK W W +E DLF+NGA F PS
Sbjct: 310 IGGSAAPTINSQGNRFLAPNDHVFKEVTKYEDAPRSKWKKWNWRSEGDLFLNGAFFTPSG 369
Query: 715 GGAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GGA + +P + V +T +G L C G C
Sbjct: 370 GGASSSYAKASSLSARPSSLVASVTSNAGALFCRKGSRC 408
>sw|Q9LJ42|PL10_ARATH Probable pectate lyase 10 precursor (EC
4.2.2.2).
Length = 440
Score = 323 bits (827), Expect = 4e-87
Identities = 154/279 (55%), Positives = 197/279 (70%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+I L+QEL+M+S KTIDGRG V IA GA IT+Q N+IIH ++VHD + T
Sbjct: 162 VFKRDMVITLTQELIMNSFKTIDGRGVNVAIAGGACITIQYVTNIIIHGINVHDCRRTGN 221
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP+H G RT ADGD IS+F ++++WIDH S+SNC DGLID + ST ITISN +
Sbjct: 222 AMVRSSPSHYGWRTMADGDAISIFGSSHIWIDHNSLSNCADGLIDAIMGSTAITISNNYM 281
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D++MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW+MYA
Sbjct: 282 THHNEVMLMGHSDSYTRDKLMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWVMYA 341
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHYAE-EGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP N AK +TK +G WK W W ++ DL +NGA F SG
Sbjct: 342 IGGSANPTINSQGNRFLAPGNPFAKEVTKRVGSWQGEWKQWNWRSQGDLMLNGAYFTKSG 401
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
AP + KP + V+ LT SG L C G C
Sbjct: 402 AAAPASYARASSLGAKPASVVSMLTYSSGALKCRIGMRC 440
>sw|Q9C5M8|PL18_ARATH Probable pectate lyase 18 precursor (EC
4.2.2.2) (Pectate lyase A10).
Length = 408
Score = 323 bits (827), Expect = 4e-87
Identities = 156/279 (55%), Positives = 192/279 (68%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M IQL +EL+M+S KTIDGRGA VHI+ G IT+Q N+IIH +H+HD K
Sbjct: 130 IFQRDMTIQLKEELIMNSFKTIDGRGASVHISGGPCITIQYVTNIIIHGIHIHDCKQGGN 189
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R SP H G RT +DGDG+S+F ++VW+DH S SNCEDGLID + ST IT+SN H
Sbjct: 190 AMVRSSPRHFGWRTISDGDGVSIFGGSHVWVDHCSFSNCEDGLIDAIMGSTAITLSNNHM 249
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SD++ +D+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 250 THHDKVMLLGHSDTYSRDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 309
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP +K +TKH A E WK W W + DL +NGA F PSG
Sbjct: 310 IGGSANPTINSQGNRFLAPNIRFSKEVTKHEDAPESEWKRWNWRSSGDLLLNGAFFTPSG 369
Query: 718 GAPKQVDTN-EWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + KP + V LT SG L+C G C
Sbjct: 370 GAASSSYAKASSLGAKPSSLVGPLTSTSGALNCRKGSRC 408
>sw|P40972|PEL_TOBAC Pectate lyase precursor (EC 4.2.2.2).
Length = 397
Score = 319 bits (818), Expect = 5e-86
Identities = 156/282 (55%), Positives = 200/282 (70%), Gaps = 5/282 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF ++M I+LS+EL+++S+KTIDGRG VHI NGAGI +Q A N+II NL +H++ T G
Sbjct: 116 IFGKNMKIKLSRELIVTSNKTIDGRGFNVHIQNGAGIKIQSASNIIISNLRIHNIVPTPG 175
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
GL+R+S H+G R +GDGIS+FS+ ++WIDHISMS DGLID V +ST ITISNCHF
Sbjct: 176 GLLRESEDHVGLRGSDEGDGISIFSSHDIWIDHISMSRATDGLIDAVAASTNITISNCHF 235
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H VMLFGA+D + D+ M+IT+A+NHFG+ L QRMPRCR+GFFH+VNNDYTHW YA
Sbjct: 236 TDHEKVMLFGANDHYVLDKDMKITLAYNHFGKRLDQRMPRCRFGFFHLVNNDYTHWERYA 295
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVIT---KHYAEEGVWKNWVWHTEDDLFMNGAIFNP 711
IGG TIISQGNR+IA + K +T K A W W W ++ D NGA F P
Sbjct: 296 IGGSSGATIISQGNRFIAEDELLVKEVTYREKLTASVAEWMKWTWISDGDDMENGATFTP 355
Query: 712 SG--GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
SG ++D +KP+P + V LT+FSG LSC G+PC
Sbjct: 356 SGDQNLLDKIDHLNLIKPEPSSKVGILTKFSGALSCVKGRPC 397
>sw|Q940Q1|PEL1_ARATH Probable pectate lyase 1 precursor (EC
4.2.2.2) (Pectate lyase A1).
Length = 431
Score = 319 bits (817), Expect = 6e-86
Identities = 156/279 (55%), Positives = 190/279 (68%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+IQL QEL+++S KTIDGRGA VHIANG IT+Q NVI+H LH+HD K T
Sbjct: 151 VFKRDMVIQLKQELIVNSFKTIDGRGANVHIANGGCITIQFVTNVIVHGLHIHDCKPTGN 210
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R S TH G RT ADGD IS+F +++VWIDH S+S+C DGL+D V ST ITISN H
Sbjct: 211 AMVRSSETHFGWRTMADGDAISIFGSSHVWIDHNSLSHCADGLVDAVMGSTAITISNNHL 270
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+VML G SDS+ +D+ MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 271 THHNEVMLLGHSDSYMRDKAMQVTIAYNHFGVGLIQRMPRCRHGYFHVVNNDYTHWEMYA 330
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNRY AP N AK +TK WK W W +E DL NGA F SG
Sbjct: 331 IGGSANPTINSQGNRYAAPKNPFAKEVTKRVDTPASHWKGWNWRSEGDLLQNGAYFTSSG 390
Query: 718 GAPK-QVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
A + K + V +T +G L C G+ C
Sbjct: 391 AAASGSYARASSLSAKSSSLVGHITSDAGALPCRRGRQC 429
>sptr|Q93XJ1|Q93XJ1 Pectate lyase.
Length = 409
Score = 318 bits (814), Expect = 1e-85
Identities = 153/279 (54%), Positives = 195/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+FAR M+I+L +EL+M+S KTIDGRGA VHIA G IT+Q N+IIH +++HD K
Sbjct: 131 VFARDMVIKLREELIMNSFKTIDGRGASVHIAGGPCITIQYVTNIIIHGVNIHDCKRGGN 190
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
+RDSP+H G RT +DGDG+S+F ++VW+DH S+SNC DGLID + ST ITISN +
Sbjct: 191 AHVRDSPSHYGWRTVSDGDGVSIFGGSHVWVDHCSLSNCNDGLIDAIHGSTAITISNNYL 250
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN VML G SDS+ QD+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 251 THHNKVMLLGHSDSYKQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWKMYA 310
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP + K +TKH A + WK W W +E DL +NGA F SG
Sbjct: 311 IGGSADPTINSQGNRFLAPNDRFNKEVTKHEDAPQSAWKGWNWRSEGDLLLNGAFFTASG 370
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + + + V+ +T +G+L C G C
Sbjct: 371 AGASSSYAKASSLGARSSSLVSSITAGAGSLVCKKGSRC 409
>sw|P15721|PE56_LYCES Probable pectate lyase P56 precursor (EC
4.2.2.2).
Length = 398
Score = 318 bits (814), Expect = 1e-85
Identities = 157/281 (55%), Positives = 199/281 (70%), Gaps = 5/281 (1%)
Frame = +1
Query: 4 FARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMGG 183
FARSM I+L++EL++SS+KTIDGRG VHIANGAGI +Q A NVII NL +H++ T GG
Sbjct: 118 FARSMRIRLTRELIVSSNKTIDGRGKYVHIANGAGIKIQSASNVIISNLRIHNIVPTAGG 177
Query: 184 LMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHFT 363
L+R+S H+G R +GD IS+F++ ++WIDHISMS DGLID V ST ITISNCHFT
Sbjct: 178 LLRESDDHLGLRGADEGDAISIFNSHDIWIDHISMSRATDGLIDAVAGSTNITISNCHFT 237
Query: 364 NHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYAI 543
+H VMLFGA+D +D+ M+IT+A+NHFG+ L QRMPRCR+GFFH+VNNDYTHW YAI
Sbjct: 238 DHEKVMLFGANDHAEEDRGMKITLAYNHFGKRLDQRMPRCRFGFFHLVNNDYTHWERYAI 297
Query: 544 GGGDAPTIISQGNRYIAPPNIAAKVIT---KHYAEEGVWKNWVWHTEDDLFMNGAIFNPS 714
GG TIISQGNR+IA + K +T K + W W W T+ D F NGA F PS
Sbjct: 298 GGSSGATIISQGNRFIAEDKLLVKEVTYREKSTSSVEEWMKWTWITDGDDFENGATFTPS 357
Query: 715 G--GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
G ++D ++P+P + V LT+FSG LSC +PC
Sbjct: 358 GDQNLLSKIDHLNLIQPEPSSKVGLLTKFSGALSCKIRRPC 398
>sw|Q9SVQ6|PL14_ARATH Putative pectate lyase 14 precursor (EC
4.2.2.2).
Length = 418
Score = 315 bits (808), Expect = 7e-85
Identities = 152/279 (54%), Positives = 195/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
+F R M+I LSQEL+M+S KTIDGRG VHIA GA +TVQ N+IIH +++HD K T
Sbjct: 140 VFKRDMVITLSQELIMNSFKTIDGRGVNVHIAGGACLTVQYVTNIIIHGINIHDCKRTGN 199
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
++R S +H G RT ADGDGIS+F ++++WIDH S+S+C DGLID + ST ITISN +
Sbjct: 200 AMVRSSESHYGWRTMADGDGISIFGSSHIWIDHNSLSSCADGLIDAIMGSTAITISNNYL 259
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+HN+ +L G +DS+ +D++MQ+T+A+NHFG GL+QRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 260 THHNEAILLGHTDSYTRDKMMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHWEMYA 319
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP N AK +TK A +G W NW W ++ DL +NGA F SG
Sbjct: 320 IGGSANPTINSQGNRFLAPGNRFAKEVTKRVGAGKGEWNNWNWRSQGDLMLNGAYFTSSG 379
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + K + V LT SG L C G C
Sbjct: 380 AGASANYARASSLAAKSSSLVGMLTSSSGALKCRIGTLC 418
>sw|P24396|9612_LYCES Style development-specific protein 9612
precursor.
Length = 404
Score = 314 bits (805), Expect = 1e-84
Identities = 153/281 (54%), Positives = 192/281 (68%), Gaps = 4/281 (1%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+IQL QEL+M+S KTIDGRGA VHI+ G IT+ N+IIH +++HD K +
Sbjct: 124 IFKRDMVIQLKQELVMNSYKTIDGRGASVHISGGPCITIHHTSNIIIHGINIHDCKQSGN 183
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G +RDSP H G +DGDGIS+F N+W+DH S+SNC DGLID + ST ITISN +F
Sbjct: 184 GNIRDSPNHSGWWDVSDGDGISIFGGKNIWVDHCSLSNCHDGLIDAIHGSTAITISNNYF 243
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SDS+ QD+ MQ+TVAFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 244 THHDKVMLLGHSDSFTQDKGMQVTVAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 303
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG APTI SQGNR++AP K +TKH A E W++W W +E DL +NGA F +G
Sbjct: 304 IGGSAAPTINSQGNRFLAPNEKYRKEVTKHEDAPESQWRSWNWRSEGDLMLNGAYFRQTG 363
Query: 718 GAPKQVDT---NEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
T + +P + V +T +G ++C G C
Sbjct: 364 AGASSSSTYARASSLSARPSSLVGSITTNAGPVNCKKGSRC 404
>sw|Q9LTZ0|PL11_ARATH Putative pectate lyase 11 precursor (EC
4.2.2.2).
Length = 409
Score = 313 bits (802), Expect = 3e-84
Identities = 147/279 (52%), Positives = 195/279 (69%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+I+L +EL+++S KTIDGRG+ VHI +G + + A N+IIH +++HD K G
Sbjct: 131 IFKRDMVIRLKKELIITSFKTIDGRGSSVHITDGPCLKIHYATNIIIHGINIHDCKPGSG 190
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
G+++D P H G ++DGD +++F +VWIDH S+SNC+DGLID + ST ITISN H
Sbjct: 191 GMIKDGPHHTGWWMQSDGDAVAIFGGKHVWIDHCSLSNCDDGLIDAIHGSTAITISNNHM 250
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SDS+ QD+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 251 THHDKVMLLGHSDSYTQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 310
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG +PTI SQGNR++AP K +TKH A E W++W W +E D+ +NGA F SG
Sbjct: 311 IGGSASPTIYSQGNRFLAPNTRFNKEVTKHEDAPESKWRDWNWRSEGDMLLNGAYFRESG 370
Query: 718 G-APKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
AP + +P + V +T +GTLSC G+ C
Sbjct: 371 AEAPSTYARASSLSARPSSLVGSITTTAGTLSCRRGRRC 409
>sptr|Q9M505|Q9M505 Pectate lyase.
Length = 398
Score = 313 bits (802), Expect = 3e-84
Identities = 153/279 (54%), Positives = 190/279 (68%), Gaps = 2/279 (0%)
Frame = +1
Query: 1 IFARSMIIQLSQELLMSSDKTIDGRGAQVHIANGAGITVQLAQNVIIHNLHVHDVKHTMG 180
IF R M+I+L QEL+M+S KTIDGRGA VHIA G IT+ A N+IIH LH+HD K
Sbjct: 120 IFKRDMVIKLKQELVMNSFKTIDGRGASVHIAGGPCITIHYASNIIIHGLHIHDCKQGGN 179
Query: 181 GLMRDSPTHIGSRTRADGDGISLFSATNVWIDHISMSNCEDGLIDVVQSSTGITISNCHF 360
+R+SP H G T +DGDG+S+F ++W+DH S+SNC DGLID + ST ITISN
Sbjct: 180 ANIRNSPHHSGWWTVSDGDGVSIFGGRHIWVDHCSLSNCHDGLIDAIHGSTAITISNNFM 239
Query: 361 TNHNDVMLFGASDSWPQDQIMQITVAFNHFGRGLVQRMPRCRWGFFHVVNNDYTHWLMYA 540
T+H+ VML G SDS+ +D+ MQ+T+AFNHFG GLVQRMPRCR G+FHVVNNDYTHW MYA
Sbjct: 240 THHDKVMLLGHSDSYTEDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYTHWEMYA 299
Query: 541 IGGGDAPTIISQGNRYIAPPNIAAKVITKHY-AEEGVWKNWVWHTEDDLFMNGAIFNPSG 717
IGG PTI SQGNR++AP + K +TKH A E W++W W +E DL +NGA F SG
Sbjct: 300 IGGSADPTINSQGNRFLAPNDRFKKAVTKHEDAPESEWRHWNWRSEGDLMLNGAFFLQSG 359
Query: 718 -GAPKQVDTNEWVKPKPGTYVTRLTRFSGTLSCCTGKPC 831
GA + +P + V +T SG L C G C
Sbjct: 360 AGASSSYARRSSLSARPSSLVGSITLGSGALGCRKGSRC 398
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 855,731,632
Number of Sequences: 1395590
Number of extensions: 19362511
Number of successful extensions: 58576
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 54520
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 58236
length of database: 442,889,342
effective HSP length: 125
effective length of database: 268,440,592
effective search space used: 71136756880
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

