BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= 3024030.2.1
(412 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
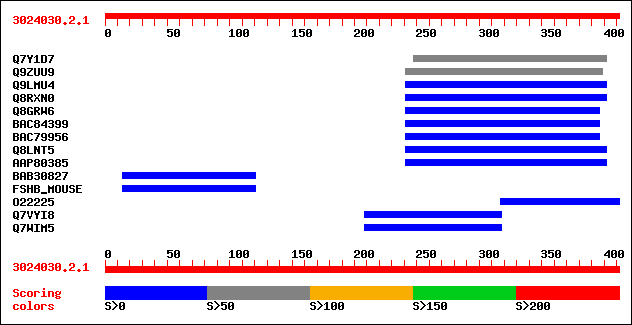
sptr|Q7Y1D7|Q7Y1D7 Putative ABC transporter. 94 5e-19
sptr|Q9ZUU9|Q9ZUU9 Putative ABC transporter. 75 2e-13
sptr|Q9LMU4|Q9LMU4 F2H15.7 protein. 36 0.13
sptr|Q8RXN0|Q8RXN0 Putative ABC transporter protein. 36 0.13
sptr|Q8GRW6|Q8GRW6 Putative ABC transporter family protein. 33 0.82
sptrnew|BAC84399|BAC84399 Putative ABC transporter family protein. 33 0.82
sptrnew|BAC79956|BAC79956 Putative ABC transporter family protein. 33 0.82
sptr|Q8LNT5|Q8LNT5 Putative ABC transporter. 32 1.8
sptrnew|AAP80385|AAP80385 ABC transporter. 31 3.1
sptrnew|BAB30827|BAB30827 8 days embryo whole body cDNA, RIKEN f... 30 5.3
sw|Q60687|FSHB_MOUSE Follitropin beta chain precursor (Follicle-... 30 5.3
sptr|O22225|O22225 At2g41640 protein (Unknown protein) (At2g4164... 30 7.0
sptr|Q7VYI8|Q7VYI8 Putative citrate lyase beta chain (EC 4.1.3.6). 30 9.1
sptr|Q7WIM5|Q7WIM5 Putative citrate lyase beta chain (EC 4.1.3.6). 30 9.1
>sptr|Q7Y1D7|Q7Y1D7 Putative ABC transporter.
Length = 687
Score = 93.6 bits (231), Expect = 5e-19
Identities = 44/52 (84%), Positives = 48/52 (92%)
Frame = -2
Query: 402 LSFISFHTYAVQGLVENEYVGTSFAVGQIRTIPGVQAVRGSYDISSSPNAKW 247
LS++SFH YAV+GL+ENEYVGTSFAVG IRTIPGVQAV GSYDISSS NAKW
Sbjct: 618 LSYVSFHVYAVEGLLENEYVGTSFAVGAIRTIPGVQAVGGSYDISSSANAKW 669
>sptr|Q9ZUU9|Q9ZUU9 Putative ABC transporter.
Length = 705
Score = 75.1 bits (183), Expect = 2e-13
Identities = 31/53 (58%), Positives = 44/53 (83%)
Frame = -2
Query: 399 SFISFHTYAVQGLVENEYVGTSFAVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
++ISFHTY+++GL+ENEY+G FAVG++R+I G QA++G+Y IS NAKW N
Sbjct: 607 AYISFHTYSIEGLLENEYLGEVFAVGEVRSISGYQAIQGNYQISPDTNAKWRN 659
>sptr|Q9LMU4|Q9LMU4 F2H15.7 protein.
Length = 659
Score = 35.8 bits (81), Expect = 0.13
Identities = 19/55 (34%), Positives = 31/55 (56%), Gaps = 1/55 (1%)
Frame = -2
Query: 402 LSFISFHTYAVQGLVENEYVGTSF-AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
+S+ISFH +A+QG +N+ G +F + G IPG + + I +KW+N
Sbjct: 532 MSYISFHFWALQGQYQNDLRGLTFDSQGSAFKIPGEYVLENVFQIDLH-RSKWIN 585
>sptr|Q8RXN0|Q8RXN0 Putative ABC transporter protein.
Length = 703
Score = 35.8 bits (81), Expect = 0.13
Identities = 19/55 (34%), Positives = 31/55 (56%), Gaps = 1/55 (1%)
Frame = -2
Query: 402 LSFISFHTYAVQGLVENEYVGTSF-AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
+S+ISFH +A+QG +N+ G +F + G IPG + + I +KW+N
Sbjct: 576 MSYISFHFWALQGQYQNDLRGLTFDSQGSAFKIPGEYVLENVFQIDLH-RSKWIN 629
>sptr|Q8GRW6|Q8GRW6 Putative ABC transporter family protein.
Length = 724
Score = 33.1 bits (74), Expect = 0.82
Identities = 21/54 (38%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Frame = -2
Query: 396 FISFHTYAVQGLVENEYVGTSF--AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
FIS+H YA QGL +NE +G F G TI G ++ + S +KWV+
Sbjct: 634 FISYHKYATQGLYKNELLGLVFEDIGGGGLTISGEYILKNYLQVELS-YSKWVD 686
>sptrnew|BAC84399|BAC84399 Putative ABC transporter family protein.
Length = 729
Score = 33.1 bits (74), Expect = 0.82
Identities = 21/54 (38%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Frame = -2
Query: 396 FISFHTYAVQGLVENEYVGTSF--AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
FIS+H YA QGL +NE +G F G TI G ++ + S +KWV+
Sbjct: 639 FISYHKYATQGLYKNELLGLVFEDIGGGGLTISGEYILKNYLQVELS-YSKWVD 691
>sptrnew|BAC79956|BAC79956 Putative ABC transporter family protein.
Length = 729
Score = 33.1 bits (74), Expect = 0.82
Identities = 21/54 (38%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Frame = -2
Query: 396 FISFHTYAVQGLVENEYVGTSF--AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
FIS+H YA QGL +NE +G F G TI G ++ + S +KWV+
Sbjct: 639 FISYHKYATQGLYKNELLGLVFEDIGGGGLTISGEYILKNYLQVELS-YSKWVD 691
>sptr|Q8LNT5|Q8LNT5 Putative ABC transporter.
Length = 723
Score = 32.0 bits (71), Expect = 1.8
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 2/56 (3%)
Frame = -2
Query: 402 LSFISFHTYAVQGLVENEYVGTSF--AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
+S+ISFH +A+QG +N+ G F ++ IPG + + I S +KW++
Sbjct: 597 MSYISFHYWALQGQYQNDLKGLVFDNQDDELPKIPGEYILENVFQIDVS-RSKWLD 651
>sptrnew|AAP80385|AAP80385 ABC transporter.
Length = 705
Score = 31.2 bits (69), Expect = 3.1
Identities = 17/56 (30%), Positives = 30/56 (53%), Gaps = 2/56 (3%)
Frame = -2
Query: 402 LSFISFHTYAVQGLVENEYVGTSF--AVGQIRTIPGVQAVRGSYDISSSPNAKWVN 241
+S+ISFH +A+QG +N+ G F ++ IPG + + I +KW++
Sbjct: 578 MSYISFHFWALQGQYQNDLKGLLFDNQPPELPKIPGEYILENVFQIDVG-RSKWID 632
>sptrnew|BAB30827|BAB30827 8 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:5730420N04
product:follicle stimulating hormone beta, full insert
sequence.
Length = 130
Score = 30.4 bits (67), Expect = 5.3
Identities = 12/36 (33%), Positives = 17/36 (47%)
Frame = -1
Query: 121 AFCCHWCKVIFELLSNRVSECRRCLVSSLVWFGLVC 14
A CCH C++ +S ECR C+ + W C
Sbjct: 16 AICCHSCELTNITISVEKEECRFCISINTTWCAGYC 51
>sw|Q60687|FSHB_MOUSE Follitropin beta chain precursor
(Follicle-stimulating hormone beta subunit) (FSH-beta)
(FSH-B).
Length = 130
Score = 30.4 bits (67), Expect = 5.3
Identities = 12/36 (33%), Positives = 17/36 (47%)
Frame = -1
Query: 121 AFCCHWCKVIFELLSNRVSECRRCLVSSLVWFGLVC 14
A CCH C++ +S ECR C+ + W C
Sbjct: 16 AICCHSCELTNITISVEKEECRFCISINTTWCAGYC 51
>sptr|O22225|O22225 At2g41640 protein (Unknown protein)
(At2g41640/T32G6.16).
Length = 500
Score = 30.0 bits (66), Expect = 7.0
Identities = 12/32 (37%), Positives = 18/32 (56%)
Frame = +2
Query: 317 ICPTANDVPTYSFSTKPCTAYVWNEMKDSLVP 412
+C +DVP FST T V++E D ++P
Sbjct: 174 VCDVYHDVPAVFFSTGGYTGNVYHEFNDGIIP 205
>sptr|Q7VYI8|Q7VYI8 Putative citrate lyase beta chain (EC 4.1.3.6).
Length = 303
Score = 29.6 bits (65), Expect = 9.1
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 2/39 (5%)
Frame = +1
Query: 208 ISERHQ*QNQEVHPLRV--WGRRNVVRASNGLDTRYGPD 318
+S + + + Q H LR +GRR+V+ NGLDT +G D
Sbjct: 58 VSAKEEARVQVTHALREGGFGRRSVIVRVNGLDTPWGED 96
>sptr|Q7WIM5|Q7WIM5 Putative citrate lyase beta chain (EC 4.1.3.6).
Length = 303
Score = 29.6 bits (65), Expect = 9.1
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 2/39 (5%)
Frame = +1
Query: 208 ISERHQ*QNQEVHPLRV--WGRRNVVRASNGLDTRYGPD 318
+S + + + Q H LR +GRR+V+ NGLDT +G D
Sbjct: 58 VSAKEEARVQVTHALREGGFGRRSVIVRVNGLDTPWGED 96
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 333,277,815
Number of Sequences: 1395590
Number of extensions: 5826504
Number of successful extensions: 20306
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 19936
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20304
length of database: 442,889,342
effective HSP length: 112
effective length of database: 286,583,262
effective search space used: 6877998288
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

