BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
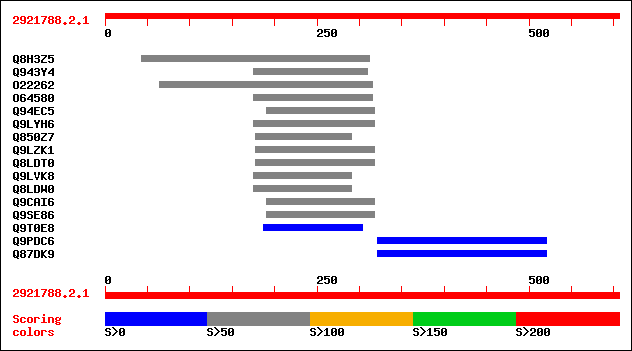
Query= 2921788.2.1
(606 letters)
Database: /db/trembl-ebi/tmp/swall
1,352,907 sequences; 430,432,536 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8H3Z5|Q8H3Z5 P0597G07.3 protein. 81 9e-15
sptr|Q943Y4|Q943Y4 P0552C05.17 protein (OSJNBb0022N24.1 protein). 67 2e-10
sptr|O22262|O22262 At2g47480 protein (Hypothetical protein). 65 5e-10
sptr|O64580|O64580 At2g19460 protein (Hypothetical protein) (Exp... 64 1e-09
sptr|Q94EC5|Q94EC5 P0002B05.14 protein. 63 3e-09
sptr|Q9LYH6|Q9LYH6 Hypothetical protein. 63 3e-09
sptr|Q850Z7|Q850Z7 Hypothetical protein OSJNBa0078D06.20. 61 1e-08
sptr|Q9LZK1|Q9LZK1 Hypothetical protein (At3g62640). 59 4e-08
sptr|Q8LDT0|Q8LDT0 Hypothetical protein. 59 4e-08
sptr|Q9LVK8|Q9LVK8 Gb|AAD10145.1 (Hypothetical protein). 55 9e-07
sptr|Q8LDW0|Q8LDW0 Hypothetical protein. 55 9e-07
sptr|Q9CAI6|Q9CAI6 Hypothetical protein. 51 1e-05
sptr|Q9SE86|Q9SE86 Hypothetical protein. 50 2e-05
sptr|Q9T0E8|Q9T0E8 Hypothetical protein. 48 8e-05
sptr|Q9PDC6|Q9PDC6 Hypothetical protein Xf1453. 32 8.0
sptr|Q87DK9|Q87DK9 ATPase. 32 8.0
>sptr|Q8H3Z5|Q8H3Z5 P0597G07.3 protein.
Length = 88
Score = 81.3 bits (199), Expect = 9e-15
Identities = 45/93 (48%), Positives = 53/93 (56%), Gaps = 3/93 (3%)
Frame = +1
Query: 43 GEYDRPYRNEVVPYG---DRRIDIIVKPPAXXXXXXXXXXXXXXXXXAWCFSDPEMKRRR 213
GEY N VVPYG +RR+ + + AW SDPEMKRRR
Sbjct: 10 GEYQYGNGNAVVPYGGGGERRMKAVCR----------------WVPGAWWLSDPEMKRRR 53
Query: 214 RVASYKAYSAEGKVKASFPRGFRWIKAKCSELI 312
RVA YK+Y+ EGKVKAS RG RWIKAKCS ++
Sbjct: 54 RVAGYKSYAVEGKVKASIRRGLRWIKAKCSHIV 86
>sptr|Q943Y4|Q943Y4 P0552C05.17 protein (OSJNBb0022N24.1 protein).
Length = 241
Score = 67.0 bits (162), Expect = 2e-10
Identities = 31/45 (68%), Positives = 33/45 (73%)
Frame = +1
Query: 175 AWCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSEL 309
A CF D E KRRRRVA YKAY+ EGKVKAS RG RW K KCS +
Sbjct: 194 AGCFGDAESKRRRRVAVYKAYAVEGKVKASLRRGIRWFKRKCSAI 238
>sptr|O22262|O22262 At2g47480 protein (Hypothetical protein).
Length = 110
Score = 65.5 bits (158), Expect = 5e-10
Identities = 33/84 (39%), Positives = 45/84 (53%)
Frame = +1
Query: 64 RNEVVPYGDRRIDIIVKPPAXXXXXXXXXXXXXXXXXAWCFSDPEMKRRRRVASYKAYSA 243
RN++ YG R + PP SD EMKR++R+A YKAY+
Sbjct: 29 RNQI--YGTRDYPPSLPPPPGKVAAGRSNASSSLRIGG--LSDAEMKRKKRIARYKAYTV 84
Query: 244 EGKVKASFPRGFRWIKAKCSELIH 315
EGKVK++ GFRWIK KC +++H
Sbjct: 85 EGKVKSTLKNGFRWIKNKCCQIVH 108
>sptr|O64580|O64580 At2g19460 protein (Hypothetical protein)
(Expressed protein).
Length = 116
Score = 64.3 bits (155), Expect = 1e-09
Identities = 24/47 (51%), Positives = 39/47 (82%)
Frame = +1
Query: 175 AWCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIH 315
+W F DP+++R++RV SY+AY+ EGK+K SF + F+WIK KC++L++
Sbjct: 70 SWGFVDPDLQRKKRVVSYRAYTVEGKLKGSFRKSFKWIKDKCNKLLN 116
>sptr|Q94EC5|Q94EC5 P0002B05.14 protein.
Length = 195
Score = 62.8 bits (151), Expect = 3e-09
Identities = 27/43 (62%), Positives = 35/43 (81%)
Frame = +1
Query: 190 DPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
DP+M+R+RRVASYKAYS EGKVK SF + F+WIK + L++G
Sbjct: 151 DPDMERKRRVASYKAYSVEGKVKGSFRKSFKWIKDRYLHLVYG 193
>sptr|Q9LYH6|Q9LYH6 Hypothetical protein.
Length = 105
Score = 62.8 bits (151), Expect = 3e-09
Identities = 24/48 (50%), Positives = 37/48 (77%)
Frame = +1
Query: 175 AWCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
+W +DPE++R++RVASYK Y EGKVK SF FRW+K + +++++G
Sbjct: 56 SWGITDPELQRKKRVASYKMYGVEGKVKGSFRNSFRWLKQRYTQVVYG 103
>sptr|Q850Z7|Q850Z7 Hypothetical protein OSJNBa0078D06.20.
Length = 74
Score = 60.8 bits (146), Expect = 1e-08
Identities = 27/38 (71%), Positives = 30/38 (78%)
Frame = +1
Query: 178 WCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIK 291
W DPE KRRRRVA+YKAY+ E +VKAS RGFRWIK
Sbjct: 30 WWSGDPEAKRRRRVAAYKAYAVEARVKASLRRGFRWIK 67
>sptr|Q9LZK1|Q9LZK1 Hypothetical protein (At3g62640).
Length = 110
Score = 59.3 bits (142), Expect = 4e-08
Identities = 24/47 (51%), Positives = 34/47 (72%)
Frame = +1
Query: 178 WCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
W D E KR++R+A+YK Y+ EGKVK++ +GF WIK + S +IHG
Sbjct: 64 WRLIDAETKRKKRIATYKTYALEGKVKSTVKKGFHWIKDRYSHIIHG 110
>sptr|Q8LDT0|Q8LDT0 Hypothetical protein.
Length = 110
Score = 59.3 bits (142), Expect = 4e-08
Identities = 24/47 (51%), Positives = 34/47 (72%)
Frame = +1
Query: 178 WCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
W D E KR++R+A+YK Y+ EGKVK++ +GF WIK + S +IHG
Sbjct: 64 WRLIDAETKRKKRIATYKTYALEGKVKSTVKKGFHWIKNRYSHIIHG 110
>sptr|Q9LVK8|Q9LVK8 Gb|AAD10145.1 (Hypothetical protein).
Length = 102
Score = 54.7 bits (130), Expect = 9e-07
Identities = 22/39 (56%), Positives = 28/39 (71%)
Frame = +1
Query: 175 AWCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIK 291
+W FSDPE +R+RRVA YK YS E K+K S + F+W K
Sbjct: 58 SWSFSDPESRRKRRVAGYKVYSVEQKMKGSIRKSFKWFK 96
>sptr|Q8LDW0|Q8LDW0 Hypothetical protein.
Length = 102
Score = 54.7 bits (130), Expect = 9e-07
Identities = 22/39 (56%), Positives = 28/39 (71%)
Frame = +1
Query: 175 AWCFSDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIK 291
+W FSDPE +R+RRVA YK YS E K+K S + F+W K
Sbjct: 58 SWSFSDPESRRKRRVAGYKVYSVEQKMKGSIRKSFKWFK 96
>sptr|Q9CAI6|Q9CAI6 Hypothetical protein.
Length = 127
Score = 50.8 bits (120), Expect = 1e-05
Identities = 19/43 (44%), Positives = 29/43 (67%)
Frame = +1
Query: 190 DPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
+ E++R++RVASY Y EG+VK S + F+W K CS ++G
Sbjct: 83 EAEIQRKKRVASYNVYGVEGRVKGSMKKSFKWFKETCSNAVYG 125
>sptr|Q9SE86|Q9SE86 Hypothetical protein.
Length = 125
Score = 50.1 bits (118), Expect = 2e-05
Identities = 19/43 (44%), Positives = 29/43 (67%)
Frame = +1
Query: 190 DPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCSELIHG 318
+ E++R++RVA+Y Y EGKVK S + F+W K CS ++G
Sbjct: 81 EAEIQRKKRVAAYNVYGVEGKVKGSIRKNFKWFKETCSNAVNG 123
>sptr|Q9T0E8|Q9T0E8 Hypothetical protein.
Length = 87
Score = 48.1 bits (113), Expect = 8e-05
Identities = 20/39 (51%), Positives = 29/39 (74%)
Frame = +1
Query: 187 SDPEMKRRRRVASYKAYSAEGKVKASFPRGFRWIKAKCS 303
+DPEMKR++RVASY ++ E K+K++ F+WIK K S
Sbjct: 39 TDPEMKRKKRVASYNLFATEEKLKSTLKNSFKWIKNKFS 77
>sptr|Q9PDC6|Q9PDC6 Hypothetical protein Xf1453.
Length = 429
Score = 31.6 bits (70), Expect = 8.0
Identities = 23/69 (33%), Positives = 37/69 (53%), Gaps = 2/69 (2%)
Frame = +3
Query: 321 NPASR*NSIVLAWCKMRLQSARDGLSVVISLIRS-QNVEENLF*DCVENFVAPLV-IGKE 494
NP+ NS +L+ C++ + A +V++L R+ Q+ E L +E A L+ I K
Sbjct: 125 NPSFELNSALLSRCRVHVLEAVSSQDIVVALQRALQDTERGLGEQKIEVSEASLLEIAKA 184
Query: 495 LDGDLLRLL 521
DGD+ R L
Sbjct: 185 ADGDVRRAL 193
>sptr|Q87DK9|Q87DK9 ATPase.
Length = 453
Score = 31.6 bits (70), Expect = 8.0
Identities = 23/69 (33%), Positives = 37/69 (53%), Gaps = 2/69 (2%)
Frame = +3
Query: 321 NPASR*NSIVLAWCKMRLQSARDGLSVVISLIRS-QNVEENLF*DCVENFVAPLV-IGKE 494
NP+ NS +L+ C++ + A +V++L R+ Q+ E L +E A L+ I K
Sbjct: 151 NPSFELNSALLSRCRVHVLEAVSSQDIVVALQRALQDTERGLGGQKIEVSQASLLEIAKA 210
Query: 495 LDGDLLRLL 521
DGD+ R L
Sbjct: 211 ADGDVRRAL 219
Database: /db/trembl-ebi/tmp/swall
Posted date: Jan 23, 2004 7:20 PM
Number of letters in database: 430,432,536
Number of sequences in database: 1,352,907
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 377,032,372
Number of Sequences: 1352907
Number of extensions: 7162366
Number of successful extensions: 15835
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 15563
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 15834
length of database: 430,432,536
effective HSP length: 117
effective length of database: 272,142,417
effective search space used: 22859963028
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

