BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
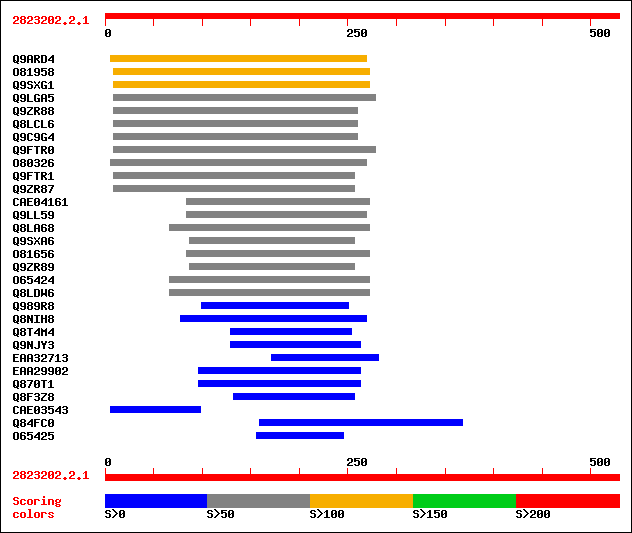
Query= 2823202.2.1
(529 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9ARD4|Q9ARD4 Putative nuclease. 122 3e-27
sptr|O81958|O81958 Endonuclease. 118 5e-26
sptr|Q9SXG1|Q9SXG1 Nuclease I. 108 3e-23
sptr|Q9LGA5|Q9LGA5 ESTs D48949(S15541). 86 2e-16
sptr|Q9ZR88|Q9ZR88 Bifunctional nuclease (Fragment). 84 1e-15
sptr|Q8LCL6|Q8LCL6 Putative bifunctional nuclease. 81 6e-15
sptr|Q9C9G4|Q9C9G4 Putative bifunctional nuclease. 81 6e-15
sptr|Q9FTR0|Q9FTR0 Putative bifunctional nuclease. 80 1e-14
sptr|O80326|O80326 Endonuclease precursor. 80 1e-14
sptr|Q9FTR1|Q9FTR1 Putative bifunctional nuclease. 79 4e-14
sptr|Q9ZR87|Q9ZR87 Bifunctional nuclease. 78 7e-14
sptrnew|CAE04161|CAE04161 OSJNBb0034I13.4 protein. 74 7e-13
sptr|Q9LL59|Q9LL59 CEL I mismatch endonuclease. 74 1e-12
sptr|Q8LA68|Q8LA68 Endonuclease, putative. 73 2e-12
sptr|Q9SXA6|Q9SXA6 Bifunctional nuclease bfn1 (Putative bifuncti... 71 8e-12
sptr|O81656|O81656 Senescence-associated protein 6. 70 2e-11
sptr|Q9ZR89|Q9ZR89 Bifunctional nuclease bfn1. 69 4e-11
sptr|O65424|O65424 Hypothetical 41.8 kDa protein (Putative bifun... 67 9e-11
sptr|Q8LDW6|Q8LDW6 Putative bifunctional nuclease. 67 9e-11
sptr|Q989R8|Q989R8 Endonuclease. 45 4e-04
sptr|Q8NIH8|Q8NIH8 Nuclease Le3. 41 0.009
sptr|Q8T4M4|Q8T4M4 P1/S1 nuclease. 37 0.17
sptr|Q9NJY3|Q9NJY3 Single strand-specific nuclease. 36 0.29
sptrnew|EAA32713|EAA32713 Hypothetical protein. 35 0.38
sptrnew|EAA29902|EAA29902 Hypothetical protein. 35 0.38
sptr|Q870T1|Q870T1 Probable nuclease S1. 35 0.38
sptr|Q8F3Z8|Q8F3Z8 Nuclease S1 (EC 3.1.30.1). 35 0.50
sptrnew|CAE03543|CAE03543 OSJNBa0060D06.9 protein. 34 0.86
sptr|Q84FC0|Q84FC0 MsrA-like protein. 34 0.86
sptr|O65425|O65425 Hypothetical 51.9 kDa protein (Putative bifun... 33 1.5
>sptr|Q9ARD4|Q9ARD4 Putative nuclease.
Length = 289
Score = 122 bits (305), Expect = 3e-27
Identities = 62/88 (70%), Positives = 68/88 (77%)
Frame = +3
Query: 6 RAIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACTAYEGVDQDSTLEDDYFFA 185
+AIQ+NIT CRSRTKTCADKYA ESA LAC AY+GV QDSTL D+Y+F
Sbjct: 196 KAIQRNITEDWSSEEKQWEACRSRTKTCADKYAEESAVLACDAYKGVKQDSTLGDEYYFK 255
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFGGNS 269
ALPVVQKRIAQGGVRLAAILNRIF G +
Sbjct: 256 ALPVVQKRIAQGGVRLAAILNRIFSGKN 283
>sptr|O81958|O81958 Endonuclease.
Length = 288
Score = 118 bits (295), Expect = 5e-26
Identities = 58/88 (65%), Positives = 67/88 (76%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACTAYEGVDQDSTLEDDYFFAA 188
+IQ+NIT CRS+T TCA+KYA ESA LAC AYEGV+QD TL D+Y+F A
Sbjct: 197 SIQRNITDDWSSEEKQWETCRSKTTTCAEKYAQESAVLACDAYEGVEQDDTLGDEYYFKA 256
Query: 189 LPVVQKRIAQGGVRLAAILNRIFGGNSR 272
LPVVQKR+AQGG+RLAAILNRIF GN R
Sbjct: 257 LPVVQKRLAQGGLRLAAILNRIFSGNGR 284
>sptr|Q9SXG1|Q9SXG1 Nuclease I.
Length = 290
Score = 108 bits (271), Expect = 3e-23
Identities = 60/90 (66%), Positives = 66/90 (73%), Gaps = 2/90 (2%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTK-TCADKYAAESAKLACTAYEGVDQDSTLEDDYFFA 185
AIQ+NIT C SRTK TCA+KYA ESA LAC AYEGV+Q TL DDY+F
Sbjct: 197 AIQRNITDDWSSEEKQWEACGSRTKITCAEKYAKESALLACDAYEGVEQGDTLGDDYYFR 256
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFGG-NSR 272
ALPVV+KRIAQGGVRLA ILN+IF G NSR
Sbjct: 257 ALPVVEKRIAQGGVRLAVILNQIFSGKNSR 286
>sptr|Q9LGA5|Q9LGA5 ESTs D48949(S15541).
Length = 308
Score = 85.9 bits (211), Expect = 2e-16
Identities = 42/91 (46%), Positives = 59/91 (64%), Gaps = 1/91 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
A++ N+T C ++ +TCA+ YA ES L+C AY+ V+QD TL DDYF++
Sbjct: 210 ALKMNLTDGWSEDISHWENCGNKKETCANDYAIESIHLSCNYAYKDVEQDITLGDDYFYS 269
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFGGNSRSG 278
P+V+KR+AQ G+RLA ILNRIFG + G
Sbjct: 270 RYPIVEKRLAQAGIRLALILNRIFGEDKPDG 300
>sptr|Q9ZR88|Q9ZR88 Bifunctional nuclease (Fragment).
Length = 280
Score = 83.6 bits (205), Expect = 1e-15
Identities = 45/85 (52%), Positives = 52/85 (61%), Gaps = 1/85 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
AI+ NIT C + KTC + YA E K AC AY+GV S LEDDYF +
Sbjct: 196 AIETNITNVWGDQVKAWENCSANQKTCPNIYATEGIKAACNWAYKGVTNGSVLEDDYFLS 255
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFG 260
LP+V R+AQGGVRLAA LNRIFG
Sbjct: 256 RLPIVNWRLAQGGVRLAANLNRIFG 280
>sptr|Q8LCL6|Q8LCL6 Putative bifunctional nuclease.
Length = 290
Score = 81.3 bits (199), Expect = 6e-15
Identities = 42/85 (49%), Positives = 56/85 (65%), Gaps = 1/85 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
A+++NIT C +T C D YA+E + AC AY+GV + TLED+YF++
Sbjct: 207 ALKKNITTEWADQVKRWDTCTKKT-ACPDIYASEGIQAACDWAYKGVTEGDTLEDEYFYS 265
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFG 260
LP+V +R+AQGGVRLAA LNRIFG
Sbjct: 266 RLPIVYQRLAQGGVRLAATLNRIFG 290
>sptr|Q9C9G4|Q9C9G4 Putative bifunctional nuclease.
Length = 290
Score = 81.3 bits (199), Expect = 6e-15
Identities = 42/85 (49%), Positives = 56/85 (65%), Gaps = 1/85 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
A+++NIT C +T C D YA+E + AC AY+GV + TLED+YF++
Sbjct: 207 ALKKNITTEWADQVKRWETCTKKT-ACPDIYASEGIQAACDWAYKGVTEGDTLEDEYFYS 265
Query: 186 ALPVVQKRIAQGGVRLAAILNRIFG 260
LP+V +R+AQGGVRLAA LNRIFG
Sbjct: 266 RLPIVYQRLAQGGVRLAATLNRIFG 290
>sptr|Q9FTR0|Q9FTR0 Putative bifunctional nuclease.
Length = 310
Score = 80.1 bits (196), Expect = 1e-14
Identities = 42/93 (45%), Positives = 59/93 (63%), Gaps = 3/93 (3%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTL--EDDYF 179
A++ N+T C ++ +TCA+ YA ES L+C AY+ V+QD TL E DYF
Sbjct: 210 ALKMNLTDGWSEDISHWENCGNKKETCANDYAIESIHLSCNYAYKDVEQDITLGGEYDYF 269
Query: 180 FAALPVVQKRIAQGGVRLAAILNRIFGGNSRSG 278
++ P+V+KR+AQ G+RLA ILNRIFG + G
Sbjct: 270 YSRYPIVEKRLAQAGIRLALILNRIFGEDKPDG 302
>sptr|O80326|O80326 Endonuclease precursor.
Length = 303
Score = 80.1 bits (196), Expect = 1e-14
Identities = 43/89 (48%), Positives = 52/89 (58%), Gaps = 1/89 (1%)
Frame = +3
Query: 6 RAIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFF 182
+AIQ N T C +KYA ES LAC YEGV+ TL DDYF
Sbjct: 206 KAIQANFTHGLWSDDVNSWKDCDDISNCVNKYAKESIALACKWGYEGVEAGETLSDDYFD 265
Query: 183 AALPVVQKRIAQGGVRLAAILNRIFGGNS 269
+ +P+V KRIAQGGVRL+ ILNR+FG +S
Sbjct: 266 SRMPIVMKRIAQGGVRLSMILNRVFGSSS 294
>sptr|Q9FTR1|Q9FTR1 Putative bifunctional nuclease.
Length = 311
Score = 78.6 bits (192), Expect = 4e-14
Identities = 44/84 (52%), Positives = 50/84 (59%), Gaps = 1/84 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
A+ QNIT C TC D YA+ES AC AY+ V +DS LED YF +
Sbjct: 227 ALMQNITGEWSQRVPGWEECSKNQTTCPDTYASESIAAACDWAYKDVTEDSLLEDAYFGS 286
Query: 186 ALPVVQKRIAQGGVRLAAILNRIF 257
LPVV R+AQGGVRLAA LNRIF
Sbjct: 287 RLPVVNLRLAQGGVRLAATLNRIF 310
>sptr|Q9ZR87|Q9ZR87 Bifunctional nuclease.
Length = 328
Score = 77.8 bits (190), Expect = 7e-14
Identities = 40/84 (47%), Positives = 52/84 (61%), Gaps = 1/84 (1%)
Frame = +3
Query: 9 AIQQNITXXXXXXXXXXXXCRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFA 185
AI++NIT C S + C D +A+ES K +C AY STL D+YF++
Sbjct: 209 AIEKNITDRWSNDISSWVNCTSGEEVCPDPWASESIKYSCNYAYRNATPGSTLGDEYFYS 268
Query: 186 ALPVVQKRIAQGGVRLAAILNRIF 257
LP+V+ R+AQGGVRLAA LNRIF
Sbjct: 269 RLPIVEMRLAQGGVRLAATLNRIF 292
>sptrnew|CAE04161|CAE04161 OSJNBb0034I13.4 protein.
Length = 252
Score = 74.3 bits (181), Expect = 7e-13
Identities = 37/64 (57%), Positives = 46/64 (71%), Gaps = 1/64 (1%)
Frame = +3
Query: 84 TCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFG 260
+C+ KYA ES LAC AY V + TL DDYF + LP+V +RIAQGGVRLA LNR+FG
Sbjct: 183 SCSTKYATESINLACKWAYNDVREGETLSDDYFGSRLPIVTRRIAQGGVRLAMFLNRLFG 242
Query: 261 GNSR 272
++R
Sbjct: 243 EHNR 246
>sptr|Q9LL59|Q9LL59 CEL I mismatch endonuclease.
Length = 296
Score = 73.9 bits (180), Expect = 1e-12
Identities = 35/63 (55%), Positives = 45/63 (71%), Gaps = 1/63 (1%)
Frame = +3
Query: 84 TCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFG 260
TCA+KYA ES KLAC Y+ V+ TL D YF +P+V KRIAQGG+RL+ ILNR+ G
Sbjct: 229 TCANKYAKESIKLACNWGYKDVESGETLSDKYFNTRMPIVMKRIAQGGIRLSMILNRVLG 288
Query: 261 GNS 269
++
Sbjct: 289 SSA 291
>sptr|Q8LA68|Q8LA68 Endonuclease, putative.
Length = 296
Score = 72.8 bits (177), Expect = 2e-12
Identities = 36/70 (51%), Positives = 45/70 (64%), Gaps = 1/70 (1%)
Frame = +3
Query: 66 CRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAI 242
C K C + YA+ES LAC AY +TL D+YF + LPVV+KR+AQGG+RLAA
Sbjct: 223 CHFHQKACPNLYASESIDLACKYAYRNATPGTTLGDEYFLSRLPVVEKRLAQGGIRLAAT 282
Query: 243 LNRIFGGNSR 272
LNRIF +
Sbjct: 283 LNRIFSAKPK 292
>sptr|Q9SXA6|Q9SXA6 Bifunctional nuclease bfn1 (Putative
bifunctional nuclease bfn1).
Length = 305
Score = 70.9 bits (172), Expect = 8e-12
Identities = 35/58 (60%), Positives = 41/58 (70%), Gaps = 1/58 (1%)
Frame = +3
Query: 87 CADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIF 257
C KYA+ES KLAC Y+GV TL ++YF LP+V KRI QGGVRLA ILNR+F
Sbjct: 236 CPHKYASESIKLACKWGYKGVKSGETLSEEYFNTRLPIVMKRIVQGGVRLAMILNRVF 293
>sptr|O81656|O81656 Senescence-associated protein 6.
Length = 298
Score = 69.7 bits (169), Expect = 2e-11
Identities = 33/64 (51%), Positives = 45/64 (70%), Gaps = 1/64 (1%)
Frame = +3
Query: 84 TCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFG 260
+C K+A+ES LAC Y+GV TL D+YF + +P+V KRIAQGGVRLA +LNR+F
Sbjct: 229 SCPKKWASESISLACKWGYKGVTPGETLSDEYFNSRMPIVMKRIAQGGVRLAMVLNRVFS 288
Query: 261 GNSR 272
+ +
Sbjct: 289 DHKQ 292
>sptr|Q9ZR89|Q9ZR89 Bifunctional nuclease bfn1.
Length = 305
Score = 68.6 bits (166), Expect = 4e-11
Identities = 35/58 (60%), Positives = 40/58 (68%), Gaps = 1/58 (1%)
Frame = +3
Query: 87 CADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIF 257
C KYA+ES KLAC Y+GV TL ++YF LP+V KRI QGGVRLA ILNR F
Sbjct: 236 CPHKYASESIKLACKWGYKGVKSGETLSEEYFNTRLPIVMKRIVQGGVRLAMILNRDF 293
>sptr|O65424|O65424 Hypothetical 41.8 kDa protein (Putative
bifunctional nuclease).
Length = 362
Score = 67.4 bits (163), Expect = 9e-11
Identities = 34/70 (48%), Positives = 43/70 (61%), Gaps = 1/70 (1%)
Frame = +3
Query: 66 CRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAI 242
C+ C + YA+ES LAC AY +TL D YF + LPVV+KR+AQGG+RLA
Sbjct: 289 CQLNQTACPNPYASESIDLACKYAYRNATAGTTLGDYYFVSRLPVVEKRLAQGGIRLAGT 348
Query: 243 LNRIFGGNSR 272
LNRIF +
Sbjct: 349 LNRIFSAKRK 358
>sptr|Q8LDW6|Q8LDW6 Putative bifunctional nuclease.
Length = 294
Score = 67.4 bits (163), Expect = 9e-11
Identities = 34/70 (48%), Positives = 43/70 (61%), Gaps = 1/70 (1%)
Frame = +3
Query: 66 CRSRTKTCADKYAAESAKLACT-AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAI 242
C+ C + YA+ES LAC AY +TL D YF + LPVV+KR+AQGG+RLA
Sbjct: 221 CQLNQTACPNPYASESIDLACKYAYRNATAGTTLGDYYFVSRLPVVEKRLAQGGIRLAGT 280
Query: 243 LNRIFGGNSR 272
LNRIF +
Sbjct: 281 LNRIFSAKRK 290
>sptr|Q989R8|Q989R8 Endonuclease.
Length = 278
Score = 45.4 bits (106), Expect = 4e-04
Identities = 21/51 (41%), Positives = 34/51 (66%)
Frame = +3
Query: 99 YAAESAKLACTAYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNR 251
+A E+ LA G+ + L++DY+ ALPVV +++ + G+RLAA+LNR
Sbjct: 217 WALEAHTLAQEMAAGITNGANLDNDYYAKALPVVDEQLGRAGLRLAAVLNR 267
>sptr|Q8NIH8|Q8NIH8 Nuclease Le3.
Length = 298
Score = 40.8 bits (94), Expect = 0.009
Identities = 22/67 (32%), Positives = 38/67 (56%), Gaps = 3/67 (4%)
Frame = +3
Query: 78 TKTCADKYAAESAKLACT---AYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILN 248
T C +A ES C+ Y+ + + ++DY+ A+P+++ +IA+ G RLAA LN
Sbjct: 229 TLNCPLVWAKESNAYDCSFVWTYDSYEDLCSDDNDYYSGAVPIIELQIAKQGYRLAAWLN 288
Query: 249 RIFGGNS 269
+F G +
Sbjct: 289 VLFDGKT 295
>sptr|Q8T4M4|Q8T4M4 P1/S1 nuclease.
Length = 316
Score = 36.6 bits (83), Expect = 0.17
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +3
Query: 129 TAYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRI 254
T+Y GV +TL DDY V + R+ GG RL +LN +
Sbjct: 252 TSYPGVTPGATLSDDYLARCKRVAEARLTLGGYRLGYLLNEL 293
>sptr|Q9NJY3|Q9NJY3 Single strand-specific nuclease.
Length = 315
Score = 35.8 bits (81), Expect = 0.29
Identities = 17/45 (37%), Positives = 24/45 (53%)
Frame = +3
Query: 129 TAYEGVDQDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFGG 263
T+Y GV +TL D Y V + R+ GG RL +LN++ G
Sbjct: 251 TSYPGVTPGATLSDAYLDKCKRVAEARLTLGGYRLGYLLNQLLSG 295
>sptrnew|EAA32713|EAA32713 Hypothetical protein.
Length = 306
Score = 35.4 bits (80), Expect = 0.38
Identities = 18/37 (48%), Positives = 26/37 (70%)
Frame = +3
Query: 171 DYFFAALPVVQKRIAQGGVRLAAILNRIFGGNSRSGF 281
DY+ A VV++ I +GG+RLA LN IF ++R+GF
Sbjct: 268 DYYKGATEVVERSIIKGGIRLAGWLNLIF--DNRTGF 302
>sptrnew|EAA29902|EAA29902 Hypothetical protein.
Length = 324
Score = 35.4 bits (80), Expect = 0.38
Identities = 24/60 (40%), Positives = 34/60 (56%), Gaps = 4/60 (6%)
Frame = +3
Query: 96 KYAAESAKLACTAY--EGVD--QDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFGG 263
++A E + CT +G + +D L YF AA PVV+ +IA+ G RLAA L+ I G
Sbjct: 234 QWAIEGNEHVCTVVLPDGPEAIRDQELGGAYFEAAAPVVELQIAKAGYRLAAWLDLIVSG 293
>sptr|Q870T1|Q870T1 Probable nuclease S1.
Length = 324
Score = 35.4 bits (80), Expect = 0.38
Identities = 24/60 (40%), Positives = 34/60 (56%), Gaps = 4/60 (6%)
Frame = +3
Query: 96 KYAAESAKLACTAY--EGVD--QDSTLEDDYFFAALPVVQKRIAQGGVRLAAILNRIFGG 263
++A E + CT +G + +D L YF AA PVV+ +IA+ G RLAA L+ I G
Sbjct: 234 QWAIEGNEHVCTVVLPDGPEAIRDQELGGAYFEAAAPVVELQIAKAGYRLAAWLDLIVSG 293
>sptr|Q8F3Z8|Q8F3Z8 Nuclease S1 (EC 3.1.30.1).
Length = 306
Score = 35.0 bits (79), Expect = 0.50
Identities = 18/45 (40%), Positives = 27/45 (60%), Gaps = 3/45 (6%)
Frame = +3
Query: 132 AYEGVDQDSTL---EDDYFFAALPVVQKRIAQGGVRLAAILNRIF 257
AY+G+ ++ D Y ALPVV+ ++A GVRLA L ++F
Sbjct: 256 AYDGIPSGRSITRISDSYIQNALPVVKHQLANAGVRLARHLEKLF 300
>sptrnew|CAE03543|CAE03543 OSJNBa0060D06.9 protein.
Length = 234
Score = 34.3 bits (77), Expect = 0.86
Identities = 16/31 (51%), Positives = 18/31 (58%)
Frame = +3
Query: 6 RAIQQNITXXXXXXXXXXXXCRSRTKTCADK 98
+AI+ NIT CRSRTKTCADK
Sbjct: 204 KAIKMNITDEWSTEEKQWETCRSRTKTCADK 234
>sptr|Q84FC0|Q84FC0 MsrA-like protein.
Length = 407
Score = 34.3 bits (77), Expect = 0.86
Identities = 19/70 (27%), Positives = 31/70 (44%)
Frame = -3
Query: 368 PAPPHPLLHMSNRLISTTGMDMDRSINCSEA*SAVPTKYPVEDRGQPDAPLRDPLLNHRQ 189
PA P P + + R + D+ R+++ P Y V +G + R+PL NH +
Sbjct: 72 PAAPGPTIQDTRRYEKPSDADLRRTLS--------PLAYEVTQKGATEPAFRNPLWNHHE 123
Query: 188 RGEEVVIFQG 159
G V + G
Sbjct: 124 EGLYVDVVSG 133
>sptr|O65425|O65425 Hypothetical 51.9 kDa protein (Putative
bifunctional nuclease).
Length = 454
Score = 33.5 bits (75), Expect = 1.5
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +3
Query: 156 STLEDDYFFAALPVVQKRIAQGGVRLAAIL 245
S +D+YF + LPVV+KR+AQ R +IL
Sbjct: 157 SQTQDEYFLSRLPVVEKRLAQVSKRFRSIL 186
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 373,802,033
Number of Sequences: 1272877
Number of extensions: 7775052
Number of successful extensions: 21734
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 20878
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 21696
length of database: 406,886,216
effective HSP length: 114
effective length of database: 261,778,238
effective search space used: 15968472518
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

