BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
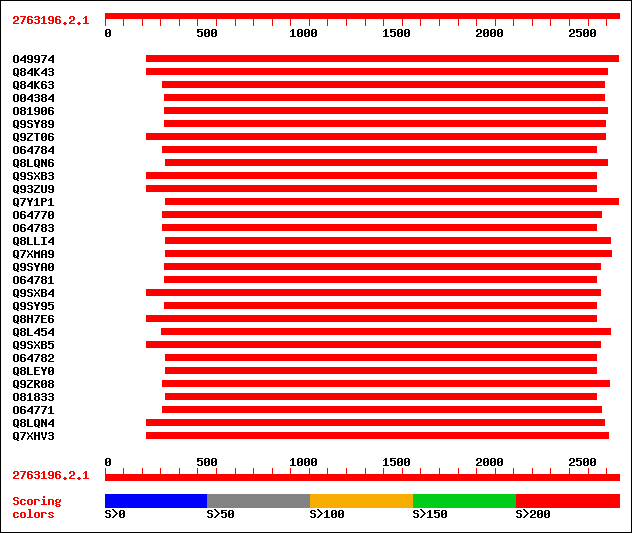
Query= 2763196.2.1
(2766 letters)
Database: /db/trembl-ebi/tmp/swall
1,302,759 sequences; 415,658,970 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O49974|O49974 KI domain interacting kinase 1. 1710 0.0
sptr|Q84K43|Q84K43 Putative S-receptor kinase. 1413 0.0
sptr|Q84K63|Q84K63 Putative S-receptor kinase. 838 0.0
sptr|O04384|O04384 Serine/threonine kinase. 681 0.0
sptr|O81906|O81906 Serine/threonine kinase - like protein (Serin... 681 0.0
sptr|Q9SY89|Q9SY89 T25B24.4 protein. 644 0.0
sptr|Q9ZT06|Q9ZT06 Receptor-like protein kinase. 598 e-169
sptr|O64784|O64784 T1F9.15. 594 e-168
sptr|Q8LQN6|Q8LQN6 Putative receptor kinase. 594 e-168
sptr|Q9SXB3|Q9SXB3 T28P6.7 protein (Very similar to receptor pro... 593 e-168
sptr|Q93ZU9|Q93ZU9 Putative receptor kinase. 593 e-168
sptr|Q7Y1P1|Q7Y1P1 Putative receptor-like kinase. 593 e-168
sptr|O64770|O64770 T1F9.1. 588 e-166
sptr|O64783|O64783 T1F9.14. 588 e-166
sptr|Q8LLI4|Q8LLI4 S-locus receptor-like kinase RLK14. 587 e-166
sptr|Q7XMA9|Q7XMA9 OSJNBb0015D13.18 protein. 586 e-166
sptr|Q9SYA0|Q9SYA0 T25B24.15 protein. 585 e-166
sptr|O64781|O64781 T1F9.12. 585 e-166
sptr|Q9SXB4|Q9SXB4 T28P6.6 protein. 584 e-165
sptr|Q9SY95|Q9SY95 T25B24.10 protein. 582 e-165
sptr|Q8H7E6|Q8H7E6 Hypothetical protein. 582 e-164
sptr|Q8L454|Q8L454 Putative serine/threonine kinase. 580 e-164
sptr|Q9SXB5|Q9SXB5 T28P6.5 protein. 579 e-164
sptr|O64782|O64782 T1F9.13 (At1g61380/T1F9_13). 575 e-162
sptr|Q8LEY0|Q8LEY0 Receptor kinase, putative. 573 e-162
sptr|Q9ZR08|Q9ZR08 Putative receptor kinase. 571 e-161
sptr|O81833|O81833 Putative receptor protein kinase. 571 e-161
sptr|O64771|O64771 T1F9.2. 571 e-161
sptr|Q8LQN4|Q8LQN4 Putative receptor kinase. 568 e-160
sptr|Q7XHV3|Q7XHV3 Putative receptor protein kinase. 566 e-160
>sptr|O49974|O49974 KI domain interacting kinase 1.
Length = 848
Score = 1710 bits (4429), Expect = 0.0
Identities = 827/848 (97%), Positives = 827/848 (97%)
Frame = -2
Query: 2765 MVSSPRXXXXXLAGASLCCVAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQP 2586
MVSSPR LAGASLCCVAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQP
Sbjct: 1 MVSSPRLLFLLLAGASLCCVAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQP 60
Query: 2585 SRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLW 2406
SRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLW
Sbjct: 61 SRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLW 120
Query: 2405 SSNTTSRAGPRGGYSAVLQDTGSLEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPK 2226
SSNTTSRAGPRGGYSAVLQDTGSLEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPK
Sbjct: 121 SSNTTSRAGPRGGYSAVLQDTGSLEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPK 180
Query: 2225 ERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL 2046
ERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL
Sbjct: 181 ERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL 240
Query: 2045 YRSGFTPAIDPVLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNEC 1866
YRSGFTPAIDPVLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNEC
Sbjct: 241 YRSGFTPAIDPVLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNEC 300
Query: 1865 EYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGD 1686
EYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGD
Sbjct: 301 EYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGD 360
Query: 1685 GFLPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHEL 1506
GFLPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHEL
Sbjct: 361 GFLPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHEL 420
Query: 1505 QTGAYTLNLKLPASELRGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWR 1326
QTGAYTLNLKLPASELRGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWR
Sbjct: 421 QTGAYTLNLKLPASELRGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWR 480
Query: 1325 SRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGG 1146
SRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGG
Sbjct: 481 SRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGG 540
Query: 1145 FGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKI 966
FGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKI
Sbjct: 541 FGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKI 600
Query: 965 LVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASN 786
LVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASN
Sbjct: 601 LVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASN 660
Query: 785 ILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGV 606
ILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGV
Sbjct: 661 ILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGV 720
Query: 605 LILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIA 426
LILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIA
Sbjct: 721 LILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIA 780
Query: 425 LLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTV 246
LLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLRG IGTV
Sbjct: 781 LLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLRGREIESSKSSEKDRSHSIGTV 840
Query: 245 TMTQLHGR 222
TMTQLHGR
Sbjct: 841 TMTQLHGR 848
>sptr|Q84K43|Q84K43 Putative S-receptor kinase.
Length = 853
Score = 1413 bits (3658), Expect = 0.0
Identities = 672/828 (81%), Positives = 723/828 (87%)
Frame = -2
Query: 2705 AAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWV 2526
AA TDTLRQGESL+GAATLVSSP GVFE GFFAPDPK PSR YLGIWY SISPRTVVWV
Sbjct: 28 AAAATDTLRQGESLTGAATLVSSPSGVFEVGFFAPDPKLPSRLYLGIWYRSISPRTVVWV 87
Query: 2525 ANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQD 2346
ANR APAT+ SPSLTL G+LRVLDG+AA+ ADAPLLW SN ++++ PRGGY AV+QD
Sbjct: 88 ANRAAPATAPSPSLTLAANGELRVLDGSAAD--ADAPLLWRSNASTQSAPRGGYKAVIQD 145
Query: 2345 TGSLEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYA 2166
TGSLEVRS+DG LWDSFWHP+DT+LSGMRIT++ PGRGP E M FTSW SETDPSPGRYA
Sbjct: 146 TGSLEVRSDDGTLWDSFWHPSDTMLSGMRITVRTPGRGPSEPMRFTSWTSETDPSPGRYA 205
Query: 2165 LGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPVLGNYYTYT 1986
LGLDP NSGQAYIW+DGNVT WRSGQW G NF+GIPWRPLY GF PA D LG YYTYT
Sbjct: 206 LGLDPANSGQAYIWRDGNVTIWRSGQWTGQNFVGIPWRPLYLYGFKPANDANLGAYYTYT 265
Query: 1985 ATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKA 1806
A+NTSLQRFVV+PNGTDICYMV+KS+Q+WE VW QPSNECEYYATCG NAKCTA QDGKA
Sbjct: 266 ASNTSLQRFVVMPNGTDICYMVKKSAQEWETVWMQPSNECEYYATCGANAKCTAMQDGKA 325
Query: 1805 KCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPDFSYWVSTV 1626
KCTCLKGF PKL +QWN GNWSQGC+RSPPLGC+ NQ+GDGFL + NIKWPDFSYW STV
Sbjct: 326 KCTCLKGFQPKLLDQWNMGNWSQGCVRSPPLGCQVNQTGDGFLSIPNIKWPDFSYWPSTV 385
Query: 1625 GDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHH 1446
DE GC CL+NCSCGAYVY T GCL WG++LIDM++ Q+G YTLNLKLPASELR HH
Sbjct: 386 QDENGCMNACLSNCSCGAYVYMTTIGCLLWGSDLIDMYQFQSGGYTLNLKLPASELRSHH 445
Query: 1445 PIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQS 1266
+WKIATI+SA+VLFVL ACL LWWK GRNIKD +H SWRS H+ST+SQQNS MLDISQS
Sbjct: 446 AVWKIATIVSAVVLFVLLACLFLWWKRGRNIKDVMHKSWRSMHTSTRSQQNSGMLDISQS 505
Query: 1265 IRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRL 1086
I F+DD EDGKSHELKVYS DRI+ AT NFSDSNKLG GGFGPVYMG LPGGEEVAVKRL
Sbjct: 506 IPFEDDTEDGKSHELKVYSFDRIKAATCNFSDSNKLGAGGFGPVYMGKLPGGEEVAVKRL 565
Query: 1085 CRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEK 906
CR SGQGLEEFKNEVILIAKLQHRNLVRLLGCCI EEKILVYEYMPNKSLDAFLFNPEK
Sbjct: 566 CRKSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIQGEEKILVYEYMPNKSLDAFLFNPEK 625
Query: 905 QRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMF 726
Q LLDW+KRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLD DM PKISDFGMARMF
Sbjct: 626 QGLLDWRKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDKDMNPKISDFGMARMF 685
Query: 725 GGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDS 546
GGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSD+Y FGVL+LEIITGKRA+SFH +DS
Sbjct: 686 GGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDIYSFGVLMLEIITGKRALSFHGQQDS 745
Query: 545 LNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILM 366
LNIAG+AWRQWNED ELIDP+IRASCS+RQVLRCIHIALLCVQDHA ERPDIP VILM
Sbjct: 746 LNIAGFAWRQWNEDKGEELIDPLIRASCSLRQVLRCIHIALLCVQDHAQERPDIPAVILM 805
Query: 365 LSNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
LS+DSSSLP PRPPTLML G IGTV+MTQLHGR
Sbjct: 806 LSSDSSSLPMPRPPTLMLHGRSAETSKSSEKDQSHSIGTVSMTQLHGR 853
>sptr|Q84K63|Q84K63 Putative S-receptor kinase.
Length = 865
Score = 838 bits (2166), Expect = 0.0
Identities = 445/823 (54%), Positives = 549/823 (66%), Gaps = 29/823 (3%)
Frame = -2
Query: 2690 DTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVA 2511
DTL QG+SL GA ++ S G F+ GFF P P + YLG+ Y + + +TV+WVANR A
Sbjct: 30 DTLSQGQSL-GANDMLVSANGTFKVGFFTPAGGDPGKVYLGVMYATSNVQTVMWVANRDA 88
Query: 2510 PATSASPSLTLTVTG--DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGS 2337
P +A+ + + TVTG +L V +G + W +N + A R ++ ++D G+
Sbjct: 89 PVRTAAGAASATVTGSGELLVKEGDR--------VAWRTNAS--AAGRSKHTLTIRDDGN 138
Query: 2336 LEVRSEDG----VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRY 2169
L + D V W+SF HPTDT + GM I L+ +R L+TSW S+ DP+ G +
Sbjct: 139 LVISGSDAAGTDVEWESFHHPTDTFVPGMEIALRQTNG---DRTLYTSWRSDADPATGDF 195
Query: 2168 ALGLDPGNSGQAYIWKDG---NVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDP--VLG 2004
LGLD S Q YIW+ N TYWRSGQW NF+GIPWR LY GF DP + G
Sbjct: 196 TLGLDA--SAQLYIWRSQGGKNSTYWRSGQWASGNFVGIPWRALYVYGFKLNGDPPPIAG 253
Query: 2003 NY-YTYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCT 1827
+ +T N+SL RFV+ PNG + CYM+ S DWELVW QP+ C Y CG NA+CT
Sbjct: 254 DMSIAFTPFNSSLYRFVLRPNGVETCYMLLGSG-DWELVWSQPTIPCHRYNLCGDNAECT 312
Query: 1826 ASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQS-----------GDGF 1680
A D + CTC GF PK +++N GNW+QGC+RS PL C + ++ GDGF
Sbjct: 313 AD-DNEPICTCFTGFEPKSPQEYNNGNWTQGCVRSVPLTCSSERNNTTAGGAGAGGGDGF 371
Query: 1679 LPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQT 1500
+ +K PDF+ W S VGD C CL NCSCGAY Y+ T CL WG EL+D+ + QT
Sbjct: 372 TVIRGVKLPDFAVWGSLVGDANSCEKACLGNCSCGAYSYS-TGSCLTWGQELVDIFQFQT 430
Query: 1499 GA----YTLNLKLPASELRGHHPIWK-IATIISAIVLFVLAACLLLWWKHGRNIKDAVHG 1335
G Y L +K+P+S L WK + ++ +V+ VL A LL WK R IK+ +
Sbjct: 431 GTEGAKYDLYVKVPSSLLDKSSGRWKTVVVVVVVVVVVVLLASGLLMWKCRRRIKEKLGI 490
Query: 1334 SWRSRHSSTQSQQNSAMLDISQSIRFDDDV-EDGKSHELKVYSLDRIRTATSNFSDSNKL 1158
+ A D S + + + E+GK+ EL +++ + + TAT NFS SNKL
Sbjct: 491 GRKKAQLPLLRPARDAKQDFSGPAQSEHEKSEEGKNCELPLFAFETLATATDNFSISNKL 550
Query: 1157 GEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPR 978
GEGGFG VY G LPGGEE+AVKRL R+SGQGLEEFKNEVILIAKLQHRNLVRLLGCCI
Sbjct: 551 GEGGFGHVYKGRLPGGEEIAVKRLSRSSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIQG 610
Query: 977 EEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDL 798
EEKILVYEYMPNKSLDAFLF+PE++ LLDW+ RF IIEG+ARGLLYLHRDSRLRVVHRDL
Sbjct: 611 EEKILVYEYMPNKSLDAFLFDPERRGLLDWRTRFQIIEGVARGLLYLHRDSRLRVVHRDL 670
Query: 797 KASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVY 618
KASNILLD DM PKISDFGMAR+FGGDQNQ NTNRVVGT GYMSPEYAMEG+FSV+SDVY
Sbjct: 671 KASNILLDRDMNPKISDFGMARIFGGDQNQVNTNRVVGTLGYMSPEYAMEGLFSVRSDVY 730
Query: 617 GFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRC 438
FG+LILEIITG++ SFH E SLNI GYAW+ WN D ELIDP IR +C ++ LRC
Sbjct: 731 SFGILILEIITGQKNSSFHHMEGSLNIVGYAWQLWNGDRGQELIDPAIRGTCPAKEALRC 790
Query: 437 IHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLR 309
+H+ALLCVQDHA +RPDIP V+L L +DSS LP PRPPT L+
Sbjct: 791 VHMALLCVQDHAHDRPDIPYVVLTLGSDSSVLPTPRPPTFTLQ 833
>sptr|O04384|O04384 Serine/threonine kinase.
Length = 850
Score = 681 bits (1756), Expect = 0.0
Identities = 383/810 (47%), Positives = 503/810 (62%), Gaps = 20/810 (2%)
Frame = -2
Query: 2690 DTLRQGESLSGAATL--VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANR 2517
DT+R+G L +T + SP+ FE GFF+P P R YLGIWY +I + VVWVANR
Sbjct: 27 DTIRRGGFLRDGSTHKPLVSPQKTFELGFFSPG-SSPGR-YLGIWYGNIEDKAVVWVANR 84
Query: 2516 VAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGS 2337
P + S LT++ G+L +L+G +WSSN TS ++L DTG+
Sbjct: 85 ENPISDRSGVLTISNDGNLVLLNGQNIT-------VWSSNITSTNNDNNRVGSIL-DTGN 136
Query: 2336 LEVR--SEDGVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYAL 2163
E+ S + V+W+SF HPTDT L MR+ + P G + + F SW SE DPSPG ++L
Sbjct: 137 FELIEVSSERVIWESFNHPTDTFLPHMRVRVN-PQTG--DNLAFVSWRSENDPSPGNFSL 193
Query: 2162 GLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL---YRSGFTPAIDP--VLGNY 1998
G+DP + + +W N WRSGQWN F GIP L Y GF + P Y
Sbjct: 194 GVDPSGAPEIVLWGRNNTRRWRSGQWNSAIFTGIPNMALLTNYLYGFKLSSPPDETGSVY 253
Query: 1997 YTYTATNTS-LQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTAS 1821
+TY ++ S L RF VL NGT+ ++S+ W P +EC+ Y CG C
Sbjct: 254 FTYVPSDPSVLLRFKVLHNGTEEELRWNETSKRWTKFQAAPESECDKYNRCGSFGICDMR 313
Query: 1820 QDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSG---DGFLPMGNIKWPD 1650
D C+C+KG+ P + GNWS+GC R PL CE N S D FL + ++K PD
Sbjct: 314 GDNGI-CSCVKGYEPV-----SLGNWSRGCRRRTPLRCERNVSNVGEDEFLTLKSVKLPD 367
Query: 1649 FSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLP 1470
F ++ D C+ CL NCSC A+ + GC+ W +L+D+ + + G +L+++L
Sbjct: 368 FETPEHSLADPEDCKDRCLKNCSCTAFTFVNGIGCMIWNQDLVDLQQFEAGGSSLHVRLA 427
Query: 1469 ASELRGHHPIWKIATIISAIV-LFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQN 1293
SE+ G KI I++ +V + +L LL W+ R K V G++ + T
Sbjct: 428 DSEI-GESKKTKIVVIVAVLVGVLLLGIFALLLWRFKR--KKDVSGTYCGHDADTSVVVV 484
Query: 1292 SAMLDISQSIRFDDDVE---DGKS---HELKVYSLDRIRTATSNFSDSNKLGEGGFGPVY 1131
+ F V+ +GK+ EL V+ L I AT++FS N+LG GGFGPVY
Sbjct: 485 DMTKAKDTTTAFTGSVDIMIEGKAVNTSELPVFCLKVIVKATNDFSRENELGRGGFGPVY 544
Query: 1130 MGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEY 951
G L G+E+AVKRL SGQG++EFKNE+ILIAKLQHRNLVRLLGCC EEK+LVYEY
Sbjct: 545 KGVLEDGQEIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEY 604
Query: 950 MPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDA 771
MPNKSLD F+F+ KQ L+DWK RF IIEGIARGLLYLHRDSRLR++HRDLK SN+LLD
Sbjct: 605 MPNKSLDFFIFDEMKQELVDWKLRFAIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDG 664
Query: 770 DMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEI 591
+M PKISDFGMAR+FGG+QN+ NT RVVGT+GYMSPEYAMEG+FSVKSDVY FGVL+LEI
Sbjct: 665 EMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEI 724
Query: 590 ITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQ 411
I+GKR S E ++ GYAW + + EL+DP IRA+C+ R+ LRCIH+A+LCVQ
Sbjct: 725 ISGKRNTSLRASEHG-SLIGYAWFLYTHGRSEELVDPKIRATCNKREALRCIHVAMLCVQ 783
Query: 410 DHADERPDIPTVILMLSNDSSSLPNPRPPT 321
D A ERP++ V+LML +D+++LP PR PT
Sbjct: 784 DSAAERPNMAAVLLMLESDTATLPVPRQPT 813
>sptr|O81906|O81906 Serine/threonine kinase - like protein
(Serine/threonine kinase-like protein).
Length = 849
Score = 681 bits (1756), Expect = 0.0
Identities = 379/814 (46%), Positives = 499/814 (61%), Gaps = 19/814 (2%)
Frame = -2
Query: 2705 AAQKTDTLRQGESLSGAATL--VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVV 2532
++ +T+R+GESL + SP+ FE GFF+P + ++LGIWY +I + VV
Sbjct: 22 SSMAANTIRRGESLRDGINHKPLVSPQKTFELGFFSPGSS--THRFLGIWYGNIEDKAVV 79
Query: 2531 WVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVL 2352
WVANR P + S L ++ G+L +LDG +WSSN S +
Sbjct: 80 WVANRATPISDQSGVLMISNDGNLVLLDGKNIT-------VWSSNIESSTTNNNNRVVSI 132
Query: 2351 QDTGS--LEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSP 2178
DTG+ L D +W+SF HPTDT L MR+ + P G + F SW SETDPSP
Sbjct: 133 HDTGNFVLSETDTDRPIWESFNHPTDTFLPQMRVRVN-PQTG--DNHAFVSWRSETDPSP 189
Query: 2177 GRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL---YRSGFTPAIDP-- 2013
G Y+LG+DP + + +W+ WRSGQWN F GIP L Y GF + P
Sbjct: 190 GNYSLGVDPSGAPEIVLWEGNKTRKWRSGQWNSAIFTGIPNMSLLTNYLYGFKLSSPPDE 249
Query: 2012 VLGNYYTYTATNTS-LQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNA 1836
Y+TY ++ S L RF VL NGT+ ++ + W +P +EC+ Y CG
Sbjct: 250 TGSVYFTYVPSDPSVLLRFKVLYNGTEEELRWNETLKKWTKFQSEPDSECDQYNRCGKFG 309
Query: 1835 KCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQS--GDGFLPMGNI 1662
C + C+C+ G+ EQ + GNWS+GC R PL CE N S D FL + ++
Sbjct: 310 ICDM-KGSNGICSCIHGY-----EQVSVGNWSRGCRRRTPLKCERNISVGEDEFLTLKSV 363
Query: 1661 KWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLN 1482
K PDF + D CR CL NCSC AY GC+ W +L+D+ + + G +L+
Sbjct: 364 KLPDFEIPEHNLVDPEDCRERCLRNCSCNAYSLVGGIGCMIWNQDLVDLQQFEAGGSSLH 423
Query: 1481 LKLPASELRGHHPIWKIATIISAIVLFVLAACL-LLWWKHGRNIKDAVHGSWRSRHSSTQ 1305
++L SE+ G + KIA I++ +V +L LL W+ R K V G++ +++ T
Sbjct: 424 IRLADSEV-GENRKTKIAVIVAVLVGVILIGIFALLLWRFKR--KKDVSGAYCGKNTDTS 480
Query: 1304 SQQNSAMLDISQSIRFDDDVE---DGKS---HELKVYSLDRIRTATSNFSDSNKLGEGGF 1143
+ F V+ +GK+ EL V+SL+ I AT++F N+LG GGF
Sbjct: 481 VVVADLTKSKETTSAFSGSVDIMIEGKAVNTSELPVFSLNAIAIATNDFCKENELGRGGF 540
Query: 1142 GPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKIL 963
GPVY G L G E+AVKRL SGQG++EFKNE+ILIAKLQHRNLVRLLGCC EEK+L
Sbjct: 541 GPVYKGVLEDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKML 600
Query: 962 VYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNI 783
VYEYMPNKSLD FLF+ KQ L+DWK RF IIEGIARGLLYLHRDSRLR++HRDLK SN+
Sbjct: 601 VYEYMPNKSLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHRDSRLRIIHRDLKVSNV 660
Query: 782 LLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVL 603
LLDA+M PKISDFGMAR+FGG+QN+ NT RVVGT+GYMSPEYAMEG+FSVKSDVY FGVL
Sbjct: 661 LLDAEMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVL 720
Query: 602 ILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIAL 423
+LEI++GKR S E ++ GYAW + + EL+DP IR +CS R+ LRCIH+A+
Sbjct: 721 LLEIVSGKRNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIRVTCSKREALRCIHVAM 779
Query: 422 LCVQDHADERPDIPTVILMLSNDSSSLPNPRPPT 321
LCVQD A ERP++ +V+LML +D+++L PR PT
Sbjct: 780 LCVQDSAAERPNMASVLLMLESDTATLAAPRQPT 813
>sptr|Q9SY89|Q9SY89 T25B24.4 protein.
Length = 842
Score = 644 bits (1661), Expect = 0.0
Identities = 361/811 (44%), Positives = 496/811 (61%), Gaps = 19/811 (2%)
Frame = -2
Query: 2696 KTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANR 2517
+ T+R+G+SL +S E FE GFF P K + +Y+GIWY +I P+TVVWVANR
Sbjct: 34 RNHTIREGDSL------ISEDES-FELGFFTP--KNSTLRYVGIWYKNIEPQTVVWVANR 84
Query: 2516 VAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGS 2337
P +L + G+L +++G N T +WS+N + AVL TG
Sbjct: 85 EKPLLDHKGALKIADDGNLVIVNGQ--NET-----IWSTNVEPESN---NTVAVLFKTGD 134
Query: 2336 LEVRSEDGV---LWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYA 2166
L + S+ W+SF +PTDT L GMR+ + P G E F W SE+DPSPG+Y+
Sbjct: 135 LVLCSDSDRRKWYWESFNNPTDTFLPGMRVRVN-PSLG--ENRAFIPWKSESDPSPGKYS 191
Query: 2165 LGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRS---GFTPAIDPVLGN-- 2001
+G+DP + + IW +G WRSG WN F GIP + + GF + P
Sbjct: 192 MGIDPVGALEIVIW-EGEKRKWRSGPWNSAIFTGIPDMLRFTNYIYGFKLSSPPDRDGSV 250
Query: 2000 YYTYTATNTS-LQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTA 1824
Y+TY A+++S RF + P+G + + K ++W L+ ++PS ECE Y CG + C
Sbjct: 251 YFTYVASDSSDFLRFWIRPDGVEEQFRWNKDIRNWNLLQWKPSTECEKYNRCGNYSVCDD 310
Query: 1823 SQD-GKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQS-----GDGFLPMGNI 1662
S++ KC+C+ GF P Q+QWN ++S GC R PL C NQS DGF + I
Sbjct: 311 SKEFDSGKCSCIDGFEPVHQDQWNNRDFSGGCQRRVPLNC--NQSLVAGQEDGFTVLKGI 368
Query: 1661 KWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLN 1482
K PDF V E C+ VC +CSC AY GC+ W +LIDM + G ++N
Sbjct: 369 KVPDFGSVVLHNNSET-CKDVCARDCSCKAYALVVGIGCMIWTRDLIDMEHFERGGNSIN 427
Query: 1481 LKLPASELRG---HHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSS 1311
++L S+L G + +W I + S I F+L C+ + WK +++K + W+ + +
Sbjct: 428 IRLAGSKLGGGKENSTLWII--VFSVIGAFLLGLCIWILWKFKKSLKAFL---WKKKDIT 482
Query: 1310 TQSQ-QNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPV 1134
+N + D V+ + +L ++S D + +AT +F++ NKLG+GGFG V
Sbjct: 483 VSDIIENRDYSSSPIKVLVGDQVD---TPDLPIFSFDSVASATGDFAEENKLGQGGFGTV 539
Query: 1133 YMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYE 954
Y G G E+AVKRL S QGLEEFKNE++LIAKLQHRNLVRLLGCCI EK+L+YE
Sbjct: 540 YKGNFSEGREIAVKRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKMLLYE 599
Query: 953 YMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLD 774
YMPNKSLD FLF+ KQ LDW+KR+++I GIARGLLYLHRDSRL+++HRDLKASNILLD
Sbjct: 600 YMPNKSLDRFLFDESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNILLD 659
Query: 773 ADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILE 594
+M PKISDFGMAR+F Q+ NT RVVGT+GYM+PEYAMEGIFS KSDVY FGVLILE
Sbjct: 660 TEMNPKISDFGMARIFNYRQDHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVLILE 719
Query: 593 IITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCV 414
I++G++ VSF D ++ GYAW W++ E+IDP+++ + V + +RCIH+ +LC
Sbjct: 720 IVSGRKNVSFR-GTDHGSLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGMLCT 778
Query: 413 QDHADERPDIPTVILMLSNDSSSLPNPRPPT 321
QD RP++ +V+LML + +S LP PR PT
Sbjct: 779 QDSVIHRPNMGSVLLMLESQTSQLPPPRQPT 809
>sptr|Q9ZT06|Q9ZT06 Receptor-like protein kinase.
Length = 830
Score = 598 bits (1541), Expect = e-169
Identities = 340/838 (40%), Positives = 475/838 (56%), Gaps = 14/838 (1%)
Frame = -2
Query: 2693 TDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRV 2514
TD + + T+VS+ F GFF+P + +Y GIW+++I +TVVWVAN
Sbjct: 22 TDVITFSSEFRDSETVVSN-HSTFRFGFFSP--VNSTGRYAGIWFNNIPVQTVVWVANSN 78
Query: 2513 APATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSL 2334
+P +S ++++ G+L V+DG + WS+N Y+ +L +TG+L
Sbjct: 79 SPINDSSGMVSISKEGNLVVMDGRGQ-------VHWSTNVLVPVAANTFYARLL-NTGNL 130
Query: 2333 EV----RSEDGVLWDSFWHPTDTILSGMRITLQAP-GRGPKERMLFTSWASETDPSPGRY 2169
+ + D +LW+SF HP + L M + GR K R SW S DPSPGRY
Sbjct: 131 VLLGTTNTGDEILWESFEHPQNIYLPTMSLATDTKTGRSLKLR----SWKSPFDPSPGRY 186
Query: 2168 ALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPV-LGNYYT 1992
+ GL P + +WKD ++ WRSG WNG FIG+P + F + G+
Sbjct: 187 SAGLIPLPFPELVVWKD-DLLMWRSGPWNGQYFIGLPNMDYRINLFELTLSSDNRGSVSM 245
Query: 1991 YTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDG 1812
A NT L F++ G+ + Q+W+ PS +C+ YATCG A C +
Sbjct: 246 SYAGNTLLYHFLLDSEGSVFQRDWNVAIQEWKTWLKVPSTKCDTYATCGQFASCRFNPGS 305
Query: 1811 KAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCET------NQSGDGFLPMGNIKWPD 1650
C C+K F P+ +WN GNW+QGC+R PL CE+ ++ DGF+ + +K P
Sbjct: 306 TPPCMCIKRFKPQSYAEWNNGNWTQGCVRKAPLQCESRDNNDGSRKSDGFVRVQKMKVPH 365
Query: 1649 FSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLP 1470
+ +E C CL NCSC A + GCL W L+DM E ++L
Sbjct: 366 NPQ--RSGANEQDCPESCLKNCSCTANSFDRGIGCLLWSGNLMDMQEFSGTGVVFYIRLA 423
Query: 1469 ASEL--RGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQ 1296
SE R + I T++ LF L LW K+ R + S
Sbjct: 424 DSEFKKRTNRSIVITVTLLVGAFLFAGTVVLALWKIAKHREKNRNTRLLNERMEALSSND 483
Query: 1295 NSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLP 1116
A+L V K EL ++ + AT+NFS +NKLG+GGFG VY G L
Sbjct: 484 VGAIL-----------VNQYKLKELPLFEFQVLAVATNNFSITNKLGQGGFGAVYKGRLQ 532
Query: 1115 GGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKS 936
G ++AVKRL R SGQG+EEF NEV +I+KLQHRNLVRLLG CI EE++LVYE+MP
Sbjct: 533 EGLDIAVKRLSRTSGQGVEEFVNEVFVISKLQHRNLVRLLGFCIEGEERMLVYEFMPENC 592
Query: 935 LDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPK 756
LDA+LF+P KQRLLDWK RF+II+GI RGL+YLHRDSRL+++HRDLKASNILLD ++ PK
Sbjct: 593 LDAYLFDPVKQRLLDWKTRFNIIDGICRGLMYLHRDSRLKIIHRDLKASNILLDENLNPK 652
Query: 755 ISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKR 576
ISDFG+AR+F G++++ +T RVVGT+GYM+PEYAM G+FS KSDV+ GV++LEI++G+R
Sbjct: 653 ISDFGLARIFQGNEDEVSTVRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIVSGRR 712
Query: 575 AVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADE 396
SF+ + N++ YAW+ WN L+DPVI C ++ RC+H+ LLCVQDHA++
Sbjct: 713 NSSFYNDGQNPNLSAYAWKLWNTGEDIALVDPVIFEECFENEIRRCVHVGLLCVQDHAND 772
Query: 395 RPDIPTVILMLSNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
RP + TVI MLS+++S+LP P+ P + R I V++T++ GR
Sbjct: 773 RPSVATVIWMLSSENSNLPEPKQPAFIPRRGTSEVESSGQSDPRASINNVSLTKITGR 830
>sptr|O64784|O64784 T1F9.15.
Length = 821
Score = 594 bits (1531), Expect = e-168
Identities = 357/801 (44%), Positives = 459/801 (57%), Gaps = 23/801 (2%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+ + QY+GIW+ ++PR +VWVANR P +S +LT++ G
Sbjct: 34 LSSPGGSYELGFFSSN--NSGNQYVGIWFKKVTPRVIVWVANREKPVSSTMANLTISSNG 91
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LD + L+WSS + A L DTG+L V LW SF
Sbjct: 92 SLILLD-------SKKDLVWSSGGDPTSNK---CRAELLDTGNLVVVDNVTGNYLWQSFE 141
Query: 2291 HPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGN 2112
H DT+L + P K+R+L TSW SETDPSPG + + P Q I K G+
Sbjct: 142 HLGDTMLPLTSLMYDIPNN--KKRVL-TSWKSETDPSPGEFVAEITPQVPSQGLIRK-GS 197
Query: 2111 VTYWRSGQWNGVNFIGIPWRPL-YRSGFTPAIDPVLGN--YYTYTATNTSLQRFVVLPNG 1941
YWRSG W G F GIP Y + D V G + N +L + P G
Sbjct: 198 SPYWRSGPWAGTRFTGIPEMDASYVNPLGMVQDEVNGTGVFAFCVLRNFNLSYIKLTPEG 257
Query: 1940 TDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQ 1761
+ + R + DW + P C+ Y CGP C S G C CLKGF PK E+
Sbjct: 258 S--LRITRNNGTDWIKHFEGPLTSCDLYGRCGPFGLCVRS--GTPMCQCLKGFEPKSDEE 313
Query: 1760 WNAGNWSQGCIRSPPLGCETNQS-------GDGFLPMGNIKWPDFSYWVSTVGDEPGCRT 1602
W +GNWS+GC+R L C+ N S D F + NIK PD SY +++ +E C
Sbjct: 314 WRSGNWSRGCVRRTNLSCQGNSSVETQGKDRDVFYHVSNIKPPD-SYELASFSNEEQCHQ 372
Query: 1601 VCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIATI 1422
CL NCSC A+ Y + GCL W EL+D + G TL+L+L SEL G I KI T+
Sbjct: 373 GCLRNCSCTAFSYVSGIGCLVWNQELLDTVKFIGGGETLSLRLAHSELTGRKRI-KIITV 431
Query: 1421 ----ISAIVLFVLAACLLLWWKHGRN-----IKDAVHGSWRSRHSSTQSQQNSAMLDISQ 1269
+S ++ VL AC ++ +N KD V G+W+S QSQ S
Sbjct: 432 ATLSLSVCLILVLVACGCWRYRVKQNGSSLVSKDNVEGAWKS---DLQSQDVSG------ 482
Query: 1268 SIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKR 1089
L + + ++TAT+NFS NKLG+GGFG VY G L G+E+AVKR
Sbjct: 483 ---------------LNFFEIHDLQTATNNFSVLNKLGQGGFGTVYKGKLQDGKEIAVKR 527
Query: 1088 LCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPE 909
L +S QG EEF NE+ LI+KLQHRNL+RLLGCCI EEK+LVYEYM NKSLD F+F+ +
Sbjct: 528 LTSSSVQGTEEFMNEIKLISKLQHRNLLRLLGCCIDGEEKLLVYEYMVNKSLDIFIFDLK 587
Query: 908 KQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARM 729
K+ +DW RF+II+GIARGLLYLHRDS LRVVHRDLK SNILLD M PKISDFG+AR+
Sbjct: 588 KKLEIDWATRFNIIQGIARGLLYLHRDSFLRVVHRDLKVSNILLDEKMNPKISDFGLARL 647
Query: 728 FGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHED 549
F G+Q+Q +T VVGT GYMSPEYA G FS KSD+Y FGVL+LEIITGK SF +D
Sbjct: 648 FHGNQHQDSTGSVVGTLGYMSPEYAWTGTFSEKSDIYSFGVLMLEIITGKEISSFSYGKD 707
Query: 548 SLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVL--RCIHIALLCVQDHADERPDIPTV 375
+ N+ YAW W+E+ L+D + S SV V RC+HI LLCVQ A +RP+I V
Sbjct: 708 NKNLLSYAWDSWSENGGVNLLDQDLDDSDSVNSVEAGRCVHIGLLCVQHQAIDRPNIKQV 767
Query: 374 ILMLSNDSSSLPNPRPPTLML 312
+ ML++ ++ LP P P +L
Sbjct: 768 MSMLTS-TTDLPKPTQPMFVL 787
>sptr|Q8LQN6|Q8LQN6 Putative receptor kinase.
Length = 856
Score = 594 bits (1531), Expect = e-168
Identities = 353/826 (42%), Positives = 472/826 (57%), Gaps = 32/826 (3%)
Frame = -2
Query: 2705 AAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWV 2526
AA D + Q ++G TLVSS GVFE GFF P+ R YLGIWY SI +TVVWV
Sbjct: 25 AATAADVIGQAGFITGNQTLVSSG-GVFELGFFVPNGATDGRTYLGIWYASIPGQTVVWV 83
Query: 2525 ANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQD 2346
ANR P + L+ G L + D A N T +WSS +R G +A LQD
Sbjct: 84 ANRQDPVVNVPAVARLSADGRLVIAD--AKNTT-----VWSSPAPARNVTAAGATARLQD 136
Query: 2345 TGSLEVRSED--GVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGR 2172
G+L V S V W SF +PTDT+L GM++ + G M TSW S +DPSPG
Sbjct: 137 DGNLVVSSGSPGSVAWQSFDYPTDTLLPGMKLGVDVKN-GITRNM--TSWTSSSDPSPGS 193
Query: 2171 YALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPVLGNYYT 1992
Y L PG + ++++ G + SG WNG G+P FT P YY+
Sbjct: 194 YTFKLVPGGLPEFFLFR-GPAMIYGSGPWNGAELTGVPDLKSQDFAFTVVSSPD-ETYYS 251
Query: 1991 YTATNTSL-QRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQD 1815
Y+ N SL RFV + V + W WY P++ C+ YA CG C S
Sbjct: 252 YSILNPSLLSRFVADATAGQVQRFVWINGA-WSSFWYYPTDPCDGYAKCGAFGYCDTSTP 310
Query: 1814 GKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWP---DFS 1644
C+CL GF P+ +QW + S GC+ + L C+ +GDGF + +K P + +
Sbjct: 311 --TLCSCLPGFQPRSPQQWGLRDASGGCVLTANLTCDG--AGDGFWTVNRMKLPAATNAT 366
Query: 1643 YWVSTVGDEPGCRTVCLNNCSCGAYVYT-----ATTGCLAWGNELIDMHELQTGAYTLNL 1479
+ D+ CR VCL NCSC AY + GC+ W +L+DM + + +
Sbjct: 367 VYAGMTLDQ--CRQVCLGNCSCRAYAAANASGGVSRGCVIWAVDLLDMRQYSGVVQDVYI 424
Query: 1478 KLPASEL-------RGHHPIWKIATIISAIVLFVLAACLLL-----WW--------KHGR 1359
+L SE+ HP + + A+V+ ++ LLL WW +
Sbjct: 425 RLAQSEVDALNAAANSEHPS---NSAVIAVVVATISGVLLLGAVGGWWFWRNRVRTRRNE 481
Query: 1358 NIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDVE-DGKSHELKVYSLDRIRTATS 1182
A G ++QQ+ A + + R D E D K +L + L I AT
Sbjct: 482 TAAAAAGGGDDVLPFRVRNQQHPAS-SVKRDQRLDVKRECDEKDLDLPLLDLKAIVAATD 540
Query: 1181 NFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVR 1002
+F+ SNK+GEGGFGPVYMG L G+EVAVKRL R S QG+ EFKNEV LIAKLQHRNLVR
Sbjct: 541 DFAASNKIGEGGFGPVYMGKLEDGQEVAVKRLSRRSVQGVVEFKNEVKLIAKLQHRNLVR 600
Query: 1001 LLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSR 822
LLGCCI +E++LVYEYM N+SLD F+F+ K++LL W KRF+II G+ARGLLYLH DSR
Sbjct: 601 LLGCCIDDDERMLVYEYMHNQSLDTFIFDEGKRKLLRWSKRFEIIVGVARGLLYLHEDSR 660
Query: 821 LRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGI 642
R++HRDLKASN+LLD +M PKISDFG+ARMFGGDQ T +V+GT+GYMSPEYAM+G+
Sbjct: 661 FRIIHRDLKASNVLLDRNMVPKISDFGIARMFGGDQTTAYTRKVIGTYGYMSPEYAMDGV 720
Query: 641 FSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASC 462
FS+KSDVY FGVL+LEI+TG+R F+ E LN+ Y+W W E + +L+D ++ S
Sbjct: 721 FSMKSDVYSFGVLVLEIVTGRRNRGFYEAELDLNLLRYSWLLWKEGRSVDLLDQLLGGSF 780
Query: 461 SVRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPP 324
+VLRCI +ALLCV+ RP + +V++ML++++++LP P P
Sbjct: 781 DYSEVLRCIQVALLCVEVQPRNRPLMSSVVMMLASENATLPEPNEP 826
>sptr|Q9SXB3|Q9SXB3 T28P6.7 protein (Very similar to receptor protein
kinases).
Length = 820
Score = 593 bits (1529), Expect = e-168
Identities = 346/823 (42%), Positives = 472/823 (57%), Gaps = 15/823 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ Q QY+GIW+ I+PR VVWVANR P T+ +LT++ G
Sbjct: 42 LSSPGGFYELGFFSPNNSQ--NQYVGIWFKKITPRVVVWVANREKPITTPVANLTISRNG 99
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LD + ++WS T R A L DTG+L + + + +LW SF
Sbjct: 100 SLILLDSSKN-------VVWS---TRRPSISNKCHAKLLDTGNLVIVDDVSENLLWQSFE 149
Query: 2291 HPTDTIL--SGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKD 2118
+P DT+L S + L A G E+ + +SW S TDPSPG + + L P Q +
Sbjct: 150 NPGDTMLPYSSLMYNL-ATG----EKRVLSSWKSHTDPSPGDFVVRLTPQVPAQIVTMR- 203
Query: 2117 GNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGN-YYTYTATNTSLQRFVVLPN 1944
G+ Y RSG W F G+P Y S F+ + D G ++Y ++ L R ++
Sbjct: 204 GSSVYKRSGPWAKTGFTGVPLMDESYTSPFSLSQDVGNGTGLFSYLQRSSELTRVIITSE 263
Query: 1943 GTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQE 1764
G R + W L + P+N C+ Y CGP C S KC C+KGF PK +E
Sbjct: 264 G--YLKTFRYNGTGWVLDFITPANLCDLYGACGPFGLCVTSNP--TKCKCMKGFVPKYKE 319
Query: 1763 QWNAGNWSQGCIRSPPLGCETNQSG-------DGFLPMGNIKWPDFSYWVSTVGDEPGCR 1605
+W GN + GC+R L C+ N S D F + N+K PD + S V D C
Sbjct: 320 EWKRGNMTSGCMRRTELSCQANLSTKTQGKGVDVFYRLANVKPPDLYEYASFV-DADQCH 378
Query: 1604 TVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIAT 1425
CL+NCSC A+ Y GCL W +ELID G L+++L +SEL G I
Sbjct: 379 QGCLSNCSCSAFAYITGIGCLLWNHELIDTIRYSVGGEFLSIRLASSELAGSRRTKIIVG 438
Query: 1424 IISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDV 1245
IS + +LA +W++ K V +W ++S S +N +
Sbjct: 439 SISLSIFVILAFGSYKYWRY--RAKQNVGPTWAFFNNSQDSWKNG--------------L 482
Query: 1244 EDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQG 1065
E + L + ++ IR AT+NF+ SNKLG+GGFGPVY GTL +++AVKRL +SGQG
Sbjct: 483 EPQEISGLTFFEMNTIRAATNNFNVSNKLGQGGFGPVYKGTLSDKKDIAVKRLSSSSGQG 542
Query: 1064 LEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWK 885
EEF NE+ LI+KLQHRNLVRLLGCCI EEK+L+YE++ NKSLD FLF+ + +DW
Sbjct: 543 TEEFMNEIKLISKLQHRNLVRLLGCCIDGEEKLLIYEFLVNKSLDTFLFDLTLKLQIDWP 602
Query: 884 KRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQF 705
KRF+II+G++RGLLYLHRDS +RV+HRDLK SNILLD M PKISDFG+ARMF G Q+Q
Sbjct: 603 KRFNIIQGVSRGLLYLHRDSCMRVIHRDLKVSNILLDDKMNPKISDFGLARMFQGTQHQD 662
Query: 704 NTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYA 525
NT +VVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+GK+ SF C E+ + G+A
Sbjct: 663 NTRKVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIISGKKISSFCCGEEGKTLLGHA 722
Query: 524 WRQWNEDNAAELIDPVIRASCS--VRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDS 351
W W E +L+D I +SCS +V RC+ I LLC+Q A +RP+I V+ M+++ +
Sbjct: 723 WECWLETGGVDLLDEDISSSCSPVEVEVARCVQIGLLCIQQQAVDRPNIAQVVTMMTS-A 781
Query: 350 SSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
+ LP P+ P L+ + VT T+++GR
Sbjct: 782 TDLPRPKQPLFALQ----IQDQESVVSVSKSVNHVTQTEIYGR 820
>sptr|Q93ZU9|Q93ZU9 Putative receptor kinase.
Length = 830
Score = 593 bits (1529), Expect = e-168
Identities = 346/823 (42%), Positives = 472/823 (57%), Gaps = 15/823 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ Q QY+GIW+ I+PR VVWVANR P T+ +LT++ G
Sbjct: 52 LSSPGGFYELGFFSPNNSQ--NQYVGIWFKKITPRVVVWVANREKPITTPVANLTISRNG 109
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LD + ++WS T R A L DTG+L + + + +LW SF
Sbjct: 110 SLILLDSSKN-------VVWS---TRRPSISNKCHAKLLDTGNLVIVDDVSENLLWQSFE 159
Query: 2291 HPTDTIL--SGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKD 2118
+P DT+L S + L A G E+ + +SW S TDPSPG + + L P Q +
Sbjct: 160 NPGDTMLPYSSLMYNL-ATG----EKRVLSSWKSHTDPSPGDFVVRLTPQVPAQIVTMR- 213
Query: 2117 GNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGN-YYTYTATNTSLQRFVVLPN 1944
G+ Y RSG W F G+P Y S F+ + D G ++Y ++ L R ++
Sbjct: 214 GSSVYKRSGPWAKTGFTGVPLMDESYTSPFSLSQDVGNGTGLFSYLQRSSELTRVIITSE 273
Query: 1943 GTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQE 1764
G R + W L + P+N C+ Y CGP C S KC C+KGF PK +E
Sbjct: 274 G--YLKTFRYNGTGWVLDFITPANLCDLYGACGPFGLCVTSNP--TKCKCMKGFVPKYKE 329
Query: 1763 QWNAGNWSQGCIRSPPLGCETNQSG-------DGFLPMGNIKWPDFSYWVSTVGDEPGCR 1605
+W GN + GC+R L C+ N S D F + N+K PD + S V D C
Sbjct: 330 EWKRGNMTSGCMRRTELSCQANLSTKTQGKGVDVFYRLANVKPPDLYEYASFV-DADQCH 388
Query: 1604 TVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIAT 1425
CL+NCSC A+ Y GCL W +ELID G L+++L +SEL G I
Sbjct: 389 QGCLSNCSCSAFAYITGIGCLLWNHELIDTIRYSVGGEFLSIRLASSELAGSRRTKIIVG 448
Query: 1424 IISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDV 1245
IS + +LA +W++ K V +W ++S S +N +
Sbjct: 449 SISLSIFVILAFGSYKYWRY--RAKQNVGPTWAFFNNSQDSWKNG--------------L 492
Query: 1244 EDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQG 1065
E + L + ++ IR AT+NF+ SNKLG+GGFGPVY GTL +++AVKRL +SGQG
Sbjct: 493 EPQEISGLTFFEMNTIRAATNNFNVSNKLGQGGFGPVYKGTLSDKKDIAVKRLSSSSGQG 552
Query: 1064 LEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWK 885
EEF NE+ LI+KLQHRNLVRLLGCCI EEK+L+YE++ NKSLD FLF+ + +DW
Sbjct: 553 TEEFMNEIKLISKLQHRNLVRLLGCCIDGEEKLLIYEFLVNKSLDTFLFDLALKLQIDWP 612
Query: 884 KRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQF 705
KRF+II+G++RGLLYLHRDS +RV+HRDLK SNILLD M PKISDFG+ARMF G Q+Q
Sbjct: 613 KRFNIIQGVSRGLLYLHRDSCMRVIHRDLKVSNILLDDKMNPKISDFGLARMFQGTQHQD 672
Query: 704 NTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYA 525
NT +VVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+GK+ SF C E+ + G+A
Sbjct: 673 NTRKVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIISGKKISSFCCGEEGKTLLGHA 732
Query: 524 WRQWNEDNAAELIDPVIRASCS--VRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDS 351
W W E +L+D I +SCS +V RC+ I LLC+Q A +RP+I V+ M+++ +
Sbjct: 733 WECWLETGGVDLLDEDISSSCSPVEVEVARCVQIGLLCIQQQAVDRPNIAQVVTMMTS-A 791
Query: 350 SSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
+ LP P+ P L+ + VT T+++GR
Sbjct: 792 TDLPRPKQPLFALQ----IQDQESVVSVSKSVNHVTQTEIYGR 830
>sptr|Q7Y1P1|Q7Y1P1 Putative receptor-like kinase.
Length = 868
Score = 593 bits (1528), Expect = e-168
Identities = 361/866 (41%), Positives = 481/866 (55%), Gaps = 52/866 (6%)
Frame = -2
Query: 2765 MVSSPRXXXXXLAGASLCCVAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQP 2586
MV+ P A L A DT+ L+G T+VS+ G F GFF PD
Sbjct: 2 MVTWPWRALPLAAVLFLFLSPAASVDTVTMEAPLAGNRTIVSAG-GTFTLGFFTPDVAPA 60
Query: 2585 SRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLW 2406
R+YLGIWY +I RTVVWVANR +P SP+L + G L ++DG ++W
Sbjct: 61 GRRYLGIWYSNILARTVVWVANRQSPVVGGSPTLKINGNGSLAIVDGQGR-------VVW 113
Query: 2405 SSNTTSRAG-PRGGYSAVLQDTGSLEVR-SEDGVLWDSFWHPTDTILSGMRITLQAPGRG 2232
+S S + G A L D G+ +R + GV W SF +PTDT+L GM++ + R
Sbjct: 114 ASPVMSASVLSAGSAKAQLLDNGNFVLRFASAGVAWQSFDYPTDTLLPGMKLGIDF--RT 171
Query: 2231 PKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIP-- 2058
+R + SW + DPSPG Y+ +DP S + ++++ TY SG WNG F G+P
Sbjct: 172 GLDRYM-NSWRAADDPSPGEYSFRIDPSGSPEFFLYRWSTRTYG-SGPWNGYQFSGVPNL 229
Query: 2057 -WRPLYRSGFTPAIDPVLGNYYTYTA--TNTSLQRFVVLPNGTDICYMVRKSSQDWELVW 1887
L + D YY Y + T L RFV+ +G M +++ W +
Sbjct: 230 RTNTLLSYQYVSTADEA---YYRYEVDDSTTILTRFVMNSSGQIQRLMWIDTTRSWSVFS 286
Query: 1886 YQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGC 1707
P +ECE Y CG C Q C C +GF P+ + W + S GCIR L C
Sbjct: 287 SYPMDECEAYRACGAYGVCNVEQS--PMCGCAEGFEPRYPKAWALRDGSGGCIRRTALNC 344
Query: 1706 ETNQSGDGFLPMGNIKWPDFSYWVSTVGDEPG---CRTVCLNNCSCGAYVYTATT----- 1551
GDGF N+K P+ + +TV G CR CL+NC+C AY T
Sbjct: 345 T---GGDGFAVTRNMKLPESAN--ATVDMALGLEECRLSCLSNCACRAYASANVTSADAK 399
Query: 1550 GCLAWGNELIDMHELQTGAYTLNLKLPASEL-----RGHHPIWKIATII--SAIVLFVLA 1392
GC W +L+DM + G L ++L AS+L + K+ II S + L +L
Sbjct: 400 GCFMWTADLLDMRQFDNGGQDLFVRLAASDLPTNSVSDNSQTAKLVEIIVPSVVALLLLL 459
Query: 1391 A----CLLLWWKHGRNIKDAVHGS-----------------------WRSRH--SSTQSQ 1299
A C++ K+ + I A++ W+ H +S +Q
Sbjct: 460 AGLVICVIKAKKNRKAIPSALNNGQVTPFGQRNHTASALNNWEITPFWQRNHVAASNDAQ 519
Query: 1298 QNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTL 1119
N++M Q D D L + ++ I AT+NFS NKLG+GGFGPVYMG L
Sbjct: 520 DNNSMRPAGQGNHQDLD--------LPSFVIETILYATNNFSADNKLGQGGFGPVYMGRL 571
Query: 1118 PGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNK 939
G+++AVKRL R S QGL EFKNEV LIAKLQHRNLVRLLGCCI E++L+YEYM N+
Sbjct: 572 DNGQDIAVKRLSRRSTQGLREFKNEVKLIAKLQHRNLVRLLGCCIDGSERMLIYEYMHNR 631
Query: 938 SLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKP 759
SL+ FLFN EKQ +L+W KRF+II GIARG+LYLH+DS LR++HRDLKASNILLD DM P
Sbjct: 632 SLNTFLFNEEKQSILNWSKRFNIINGIARGILYLHQDSALRIIHRDLKASNILLDRDMNP 691
Query: 758 KISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGK 579
KISDFG+AR+FG DQ T +VVGT+GYMSPEYAM+G+FS+KSDV+ FGVL+LEI++GK
Sbjct: 692 KISDFGVARIFGTDQTSAYTKKVVGTYGYMSPEYAMDGVFSMKSDVFSFGVLVLEIVSGK 751
Query: 578 RAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIR-ASCSVRQVLRCIHIALLCVQDHA 402
+ F+ +E LN+ YAWR W E + E +D I S +V +VLRCI I LLCVQ+
Sbjct: 752 KNRGFYHNELDLNLLRYAWRLWKEGRSLEFLDQSIAGTSSNVTEVLRCIQIGLLCVQEQP 811
Query: 401 DERPDIPTVILMLSNDSSSLPNPRPP 324
RP + V +MLS++S +L P P
Sbjct: 812 RHRPTMSAVTMMLSSESPALLEPCEP 837
>sptr|O64770|O64770 T1F9.1.
Length = 804
Score = 588 bits (1516), Expect = e-166
Identities = 336/795 (42%), Positives = 452/795 (56%), Gaps = 8/795 (1%)
Frame = -2
Query: 2672 ESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSAS 2493
ES +SS G++E GFF+P+ Q Y+GIW+ I PR VVWVANR P T S
Sbjct: 29 ESPLSVEQTLSSSNGIYELGFFSPNNSQ--NLYVGIWFKGIIPRVVVWVANRETPTTDTS 86
Query: 2492 PSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEV--RSE 2319
+L ++ G L + +G ++WS + G A L D G+L V +
Sbjct: 87 ANLAISSNGSLLLFNGKHG-------VVWSIGENFASN---GSRAELTDNGNLVVIDNAS 136
Query: 2318 DGVLWDSFWHPTDTIL--SGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGN 2145
LW+SF H DT+L S + L A G E+ + TSW ++TDPSPG + + P
Sbjct: 137 GRTLWESFEHFGDTMLPFSSLMYNL-ATG----EKRVLTSWKTDTDPSPGVFVGQITPQV 191
Query: 2144 SGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGNYYTYTATNTSL 1968
Q I + G+ Y+R+G W F GIP Y S F+ D ++TY + L
Sbjct: 192 PSQVLIMR-GSTRYYRTGPWAKTRFTGIPLMDDTYASPFSLQQDANGSGFFTYFDRSFKL 250
Query: 1967 QRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLK 1788
R ++ G+ R + DWEL + P+N C+ Y CGP C S KC CLK
Sbjct: 251 SRIIISSEGS--MKRFRHNGTDWELSYMAPANSCDIYGVCGPFGLCIVSVP--LKCKCLK 306
Query: 1787 GFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDG---FLPMGNIKWPDFSYWVSTVGDE 1617
GF P E+W GNW+ GC R L C+ N +G F P+ N+K PDF + S+V D
Sbjct: 307 GFVPHSTEEWKRGNWTGGCARLTELHCQGNSTGKDVNIFHPVTNVKLPDFYEYESSV-DA 365
Query: 1616 PGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIW 1437
C CL+NCSC A+ Y GCL W L+D + G L+++L SEL G+
Sbjct: 366 EECHQSCLHNCSCLAFAYIHGIGCLIWNQNLMDAVQFSAGGEILSIRLAHSELGGNKRNK 425
Query: 1436 KIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRF 1257
I ++ LFV+ + A G WR R A +
Sbjct: 426 IIVASTVSLSLFVI-------------LTSAAFGFWRYRVKHKAYTLKDA---------W 463
Query: 1256 DDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRN 1077
+D++ + L+ + ++ I+TAT+NFS SNKLG+GGFG VY G L G+E+AVK+L +
Sbjct: 464 RNDLKSKEVPGLEFFEMNTIQTATNNFSLSNKLGQGGFGSVYKGKLQDGKEIAVKQLSSS 523
Query: 1076 SGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRL 897
SGQG EEF NE++LI+KLQHRNLVR+LGCCI EEK+L+YE+M NKSLD F+F+ K+
Sbjct: 524 SGQGKEEFMNEIVLISKLQHRNLVRVLGCCIEGEEKLLIYEFMLNKSLDTFVFDARKKLE 583
Query: 896 LDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGD 717
+DW KRFDI++GIARGLLYLHRDSRL+V+HRDLK SNILLD M PKISDFG+ARM+ G
Sbjct: 584 VDWPKRFDIVQGIARGLLYLHRDSRLKVIHRDLKVSNILLDEKMNPKISDFGLARMYEGT 643
Query: 716 QNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNI 537
Q Q T RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEII G++ F E+ +
Sbjct: 644 QCQDKTRRVVGTLGYMSPEYAWTGVFSEKSDIYSFGVLLLEIIIGEKISRFSYGEEGKTL 703
Query: 536 AGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILMLSN 357
YAW W E +L+D + SC +V RC+ I LLCVQ +RP+ ++ ML+
Sbjct: 704 LAYAWESWGETKGIDLLDQDLADSCRPLEVGRCVQIGLLCVQHQPADRPNTLELLAMLTT 763
Query: 356 DSSSLPNPRPPTLML 312
+S LP+P+ PT ++
Sbjct: 764 -TSDLPSPKQPTFVV 777
>sptr|O64783|O64783 T1F9.14.
Length = 828
Score = 588 bits (1516), Expect = e-166
Identities = 343/808 (42%), Positives = 463/808 (57%), Gaps = 29/808 (3%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ + QY+GIW+ +I+PR VVWVANR P T+ + +LT+ G
Sbjct: 39 LSSPNGTYELGFFSPNNSR--NQYVGIWFKNITPRVVVWVANRDKPVTNNAANLTINSNG 96
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSEDGV----LWDS 2298
L +++ + ++WS T + A L + G+L + DGV LW+S
Sbjct: 97 SLILVE-------REQNVVWSIGETFSSNE---LRAELLENGNLVLI--DGVSERNLWES 144
Query: 2297 FWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKD 2118
F H DT+L + P K+R+L +SW + TDPSPG + L Q +I +
Sbjct: 145 FEHLGDTMLLESSVMYDVPNN--KKRVL-SSWKNPTDPSPGEFVAELTTQVPPQGFIMR- 200
Query: 2117 GNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGN---YYTYTATNTSLQRFVVL 1950
G+ YWR G W V F GIP + S F + D G Y+ N++L +
Sbjct: 201 GSRPYWRGGPWARVRFTGIPEMDGSHVSKFDISQDVAAGTGSLTYSLERRNSNLSYTTLT 260
Query: 1949 PNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKL 1770
G+ ++ + W P + C+ Y TCGP C S KC CLKGF PK
Sbjct: 261 SAGS--LKIIWNNGSGWVTDLEAPVSSCDVYNTCGPFGLCIRSNP--PKCECLKGFVPKS 316
Query: 1769 QEQWNAGNWSQGCIRSPPLGCETNQS-------GDGFLPMGNIKWPDFSYWVSTVGDEPG 1611
E+WN NW+ GC+R L C+ N S GD F + N+K PDF ++S + +E
Sbjct: 317 DEEWNKRNWTGGCMRRTNLSCDVNSSATAQANNGDIFDIVANVKPPDFYEYLSLINEED- 375
Query: 1610 CRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKI 1431
C+ CL NCSC A+ Y GCL W EL+D+ + G TL+++L +SEL G + + I
Sbjct: 376 CQQRCLGNCSCTAFSYIEQIGCLVWNRELVDVMQFVAGGETLSIRLASSELAGSNRVKII 435
Query: 1430 ATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDD 1251
I +I +F++ W+ WR + + Q+ N L+ SQ D
Sbjct: 436 VASIVSISVFMILVFASYWY-------------WR--YKAKQNDSNPIPLETSQ----DA 476
Query: 1250 DVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSG 1071
E K ++ + + I T T+NFS NKLG+GGFGPVY G L G+E+A+KRL SG
Sbjct: 477 WREQLKPQDVNFFDMQTILTITNNFSMENKLGQGGFGPVYKGNLQDGKEIAIKRLSSTSG 536
Query: 1070 QGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPE------ 909
QGLEEF NE+ILI+KLQHRNLVRLLGCCI EEK+L+YE+M NKSL+ F+F
Sbjct: 537 QGLEEFMNEIILISKLQHRNLVRLLGCCIEGEEKLLIYEFMANKSLNTFIFGQSLILTNL 596
Query: 908 --------KQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKI 753
K+ LDW KRF+II+GIA GLLYLHRDS LRVVHRD+K SNILLD +M PKI
Sbjct: 597 FLIWLDSTKKLELDWPKRFEIIQGIACGLLYLHRDSCLRVVHRDMKVSNILLDEEMNPKI 656
Query: 752 SDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRA 573
SDFG+ARMF G Q+Q NT RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEIITGKR
Sbjct: 657 SDFGLARMFQGTQHQANTRRVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIITGKRI 716
Query: 572 VSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADER 393
SF E+ + +AW W E ++L+D I +S S +V RC+ I LLC+Q A +R
Sbjct: 717 SSFTIGEEGKTLLEFAWDSWCESGGSDLLDQDISSSGSESEVARCVQIGLLCIQQQAGDR 776
Query: 392 PDIPTVILMLSNDSSSLPNPRPPTLMLR 309
P+I V+ ML+ + LP P+ P ++
Sbjct: 777 PNIAQVMSMLTT-TMDLPKPKQPVFAMQ 803
>sptr|Q8LLI4|Q8LLI4 S-locus receptor-like kinase RLK14.
Length = 813
Score = 587 bits (1512), Expect = e-166
Identities = 343/812 (42%), Positives = 467/812 (57%), Gaps = 13/812 (1%)
Frame = -2
Query: 2720 SLCCVAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPR 2541
SLC + D L + L L+S GVF GFF+P K + Y+GIWYH I R
Sbjct: 16 SLC----KSDDQLTPAKPLHPGDMLISDG-GVFALGFFSPT-KSNATLYVGIWYHKIPNR 69
Query: 2540 TVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYS 2361
TVVWVANR P T+ S ++ VL + + LW + G G +
Sbjct: 70 TVVWVANRDNPITAPSSAMLFISNSSDLVLSESGGH------TLWEARNNITTGGSGA-T 122
Query: 2360 AVLQDTGSLEVRSEDG-VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDP 2184
VL ++G+L +RS + +LW SF H TDTIL GM++ L+ G+ + SW DP
Sbjct: 123 VVLLNSGNLVLRSPNHTILWQSFDHLTDTILPGMKLLLKYNGQVAQR---IVSWKGPDDP 179
Query: 2183 SPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPVLG 2004
S G ++L DP + Q +W +G YWRSG WNG + I+
Sbjct: 180 STGNFSLSGDPNSDFQVLVW-NGTSPYWRSGAWNGALVSATFQSNTSSVTYQTIINKGNE 238
Query: 2003 NYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQ-DWELVWYQPSNECEYYATCGPNAKCT 1827
Y Y+ ++ S ++L I ++ S+ W +++ PS CE YA+CGP C
Sbjct: 239 IYMMYSVSDDSPSMRLMLDYTGTIKMLIWNSNLFAWSVLFSNPSYTCERYASCGPFGYCD 298
Query: 1826 ASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPD- 1650
A++ C CL GF P + N S+GC+R + C GD FL + +K PD
Sbjct: 299 AAE-AFPTCKCLDGFKP------DGLNISRGCVRKEQMKCSY---GDSFLTLPGMKTPDK 348
Query: 1649 FSYWVSTVGDEPGCRTVCLNNCSCGAYVYTA---------TTGCLAWGNELIDMHELQTG 1497
F Y + DE C C +NCSC AY Y T+ CL W EL+D+ ++ G
Sbjct: 349 FLYIRNRSLDE--CMEECRHNCSCTAYAYANLSTASMMGDTSRCLVWMGELLDLAKVTGG 406
Query: 1496 AYTLNLKLPA-SELRGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSR 1320
L L+LP+ + ++ + KI + A +L + CL+ W R + R
Sbjct: 407 GENLYLRLPSPTAVKKETDVVKIVLPVVASLLILTCICLV-WICKSRG---------KQR 456
Query: 1319 HSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFG 1140
Q++ L S + +D + + + AT+NFS N LG+GGFG
Sbjct: 457 SKEIQNKIMVQYLSASNELGAEDV-------DFPFIGFEEVVIATNNFSSYNMLGKGGFG 509
Query: 1139 PVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILV 960
VY G L GG+EVAVKRL + SGQG+EEF+NEV+LIA+LQHRNLV+L+GCCI +EK+L+
Sbjct: 510 KVYKGILEGGKEVAVKRLSKGSGQGIEEFRNEVVLIARLQHRNLVKLVGCCIHEDEKLLI 569
Query: 959 YEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNIL 780
YEY+PNKSLDAFLF+ ++ +LDW RF II+G+ARGLLYLH+DSRL ++HRDLKA NIL
Sbjct: 570 YEYLPNKSLDAFLFDATRKTVLDWPNRFKIIKGVARGLLYLHQDSRLTIIHRDLKAGNIL 629
Query: 779 LDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLI 600
LDA+M PKISDFGMAR+FGG+Q Q NT RVVGT+GYMSPEYAMEGIFSVKSD+Y FG+L+
Sbjct: 630 LDAEMSPKISDFGMARIFGGNQQQANTTRVVGTYGYMSPEYAMEGIFSVKSDIYSFGILL 689
Query: 599 LEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALL 420
LEII+G R S H N+ Y+W W + NA +L+D + SC + +VLRCIHIALL
Sbjct: 690 LEIISGFRISSPHLIMGFPNLIAYSWSLWKDGNARDLVDSSVVESCPLHEVLRCIHIALL 749
Query: 419 CVQDHADERPDIPTVILMLSNDSSSLPNPRPP 324
C+QDH D+RP + +V+ ML N+++ LP P+ P
Sbjct: 750 CIQDHPDDRPLMSSVVFMLENNTAPLPQPKQP 781
>sptr|Q7XMA9|Q7XMA9 OSJNBb0015D13.18 protein.
Length = 3259
Score = 586 bits (1510), Expect = e-166
Identities = 342/818 (41%), Positives = 468/818 (57%), Gaps = 18/818 (2%)
Frame = -2
Query: 2723 ASLCCVAAQKTDTLRQGESLSGAATL-----VSSPEGVFEAGFFAPDPKQPSRQYLGIWY 2559
A+ AA T + L+ A L + S GVF GFF+P + Y+GIWY
Sbjct: 2451 AAAAATAATATAAISDESELTPAKPLYPGDMLISDGGVFALGFFSPTNSNATL-YVGIWY 2509
Query: 2558 HSISPRTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAG 2379
H I RTVVWVANR P T+ S ++ VL + + LW + G
Sbjct: 2510 HKIPNRTVVWVANRDNPITAPSSAMLFISNSSDLVLSESGGH------TLWEARNNITTG 2563
Query: 2378 PRGGYSAVLQDTGSLEVRSEDG-VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSW 2202
G + VL ++G+L +RS + +LW SF H TDTIL GM++ L+ G+ + SW
Sbjct: 2564 GSGA-TVVLLNSGNLVLRSPNHTILWQSFDHLTDTILPGMKLLLKYNGQVAQR---IVSW 2619
Query: 2201 ASETDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPA 2022
DPS G ++L DP + Q +W +G YWRSG WNG + +
Sbjct: 2620 KGPDDPSTGNFSLSGDPNSDFQVLVW-NGTSPYWRSGAWNGALVSAMFQSNTSSVTYQTI 2678
Query: 2021 IDPVLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQ-DWELVWYQPSNECEYYATCG 1845
I+ Y Y+ ++ S ++L I ++ S+ W +++ PS CE YA+CG
Sbjct: 2679 INKGNEIYMMYSVSDDSPSMRLMLDYTGTIKMLIWNSNLFAWSVLFSNPSYTCERYASCG 2738
Query: 1844 PNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGN 1665
P C A++ C CL GF P + N S+GC+R + C GD FL +
Sbjct: 2739 PFGYCDAAE-AFPTCKCLDGFKP------DGLNISRGCVRKEQMKCSY---GDSFLTLPG 2788
Query: 1664 IKWPD-FSYWVSTVGDEPGCRTVCLNNCSCGAYVYTA---------TTGCLAWGNELIDM 1515
+K PD F Y + DE C C +NCSC AY Y T+ CL W EL+D+
Sbjct: 2789 MKTPDKFLYIRNRSLDE--CMEECRHNCSCTAYAYANLSTASMMGDTSRCLVWMGELLDL 2846
Query: 1514 HELQTGAYTLNLKLPA-SELRGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVH 1338
++ G L L+LP+ + ++ + KI + A +L + CL+ W R
Sbjct: 2847 AKVTGGGENLYLRLPSPTAVKKETDVVKIVLPVVASLLILTCICLV-WICKSRG------ 2899
Query: 1337 GSWRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKL 1158
+ R Q++ L S + +D + + + AT+NFS N L
Sbjct: 2900 ---KQRSKEIQNKIMVQYLSASNELGAEDV-------DFPFIGFEEVVIATNNFSSYNML 2949
Query: 1157 GEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPR 978
G+GGFG VY G L GG+EVAVKRL + SGQG+EEF+NEV+LIA+LQHRNLV+L+GCCI
Sbjct: 2950 GKGGFGKVYKGILEGGKEVAVKRLSKGSGQGIEEFRNEVVLIARLQHRNLVKLVGCCIHE 3009
Query: 977 EEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDL 798
+EK+L+YEY+PNKSLDAFLF+ ++ +LDW RF II+G+ARGLLYLH+DSRL ++HRDL
Sbjct: 3010 DEKLLIYEYLPNKSLDAFLFDATRKTVLDWPNRFKIIKGVARGLLYLHQDSRLTIIHRDL 3069
Query: 797 KASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVY 618
KA NILLDA+M PKISDFGMAR+FGG+Q Q NT RVVGT+GYMSPEYAMEGIFSVKSD+Y
Sbjct: 3070 KAGNILLDAEMSPKISDFGMARIFGGNQQQANTTRVVGTYGYMSPEYAMEGIFSVKSDIY 3129
Query: 617 GFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRC 438
FG+L+LEII+G R S H N+ Y+W W + NA +L+D + SC + +VLRC
Sbjct: 3130 SFGILLLEIISGFRISSPHLIMGFPNLIAYSWSLWKDGNARDLVDSSVVESCPLHEVLRC 3189
Query: 437 IHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPP 324
IHIALLC+QDH D+RP + +V+ ML N+++ LP P+ P
Sbjct: 3190 IHIALLCIQDHPDDRPLMSSVVFMLENNTAPLPQPKQP 3227
Score = 522 bits (1344), Expect = e-146
Identities = 330/808 (40%), Positives = 450/808 (55%), Gaps = 21/808 (2%)
Frame = -2
Query: 2690 DTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISP--RTVVWVANR 2517
D L Q L ++ S VF GFF+P S +LGIWYH+IS RT VWVANR
Sbjct: 1564 DQLTQANRLISPGDVLISKGRVFALGFFSPTASNQSF-FLGIWYHNISESERTYVWVANR 1622
Query: 2516 VAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGS 2337
P T+ S + TL ++ ++ + N T LW++N T+ G G Y+A+L D+G+
Sbjct: 1623 DNPITTPSFA-TLAISNSSNLVLSDSGNHT-----LWTTNVTATGGD-GAYAALL-DSGN 1674
Query: 2336 LEVRSEDGV-LWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALG 2160
L +R +G +W SF HPTDT+L GMR + + M +W DPS G +++
Sbjct: 1675 LVLRLPNGTTIWQSFDHPTDTLLMGMRFLVSYKAQ---VAMRCIAWKGPDDPSTGDFSIS 1731
Query: 2159 LDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPVLGN------- 2001
DP ++ Q ++W +G Y + FIG ++ S F+ + +
Sbjct: 1732 GDPSSNLQIFLW-NGTRPY--------IRFIGFGPSSMWSSVFSFSTSLIYETSVSTDDE 1782
Query: 2000 -YYTYTATNTSLQRFVVLPNGTDICYMV-RKSSQDWELVWYQPSNE--CEYYATCGPNAK 1833
Y YT ++ S + + L + ++ S+ W +V +PS C+ YA+CGP
Sbjct: 1783 FYIIYTTSDGSPYKRLQLDYTGTLKFLAWNDSASSWTVVVQRPSPTIVCDPYASCGPFGY 1842
Query: 1832 CTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWP 1653
C A+ +C CL GF P + + S+GC R L C D F+ M +K P
Sbjct: 1843 CDATA-AIPRCQCLDGFEPD-----GSNSSSRGCRRKQQLRCRGRD--DRFVTMAGMKVP 1894
Query: 1652 D-FSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTG-----CLAWGNELIDMHELQTGAY 1491
D F + + DE C C NCSC AY Y TG CL W EL D G
Sbjct: 1895 DKFLHVRNRSFDE--CAAECSRNCSCTAYAYANLTGADQARCLLWSGELADTGRANIGE- 1951
Query: 1490 TLNLKLPASEL-RGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHS 1314
L L+L S + + I KI + +L ++ CL W R I H
Sbjct: 1952 NLYLRLADSTVNKKKSDIPKIVLPVITSLLILMCICLA-WICKSRGI-----------HR 1999
Query: 1313 STQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPV 1134
S + Q+ + + S ++D + EL L+ I TAT+NFSD N LG+GGFG V
Sbjct: 2000 SKEIQKKHRLQHLKDSSELEND-----NLELPFICLEDIVTATNNFSDHNMLGKGGFGKV 2054
Query: 1133 YMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYE 954
Y G L GG+E+AVKRL + S QG+EEF+NEV+LIAKLQHRNLVRL+ CI +EK+L+YE
Sbjct: 2055 YKGVLEGGKEIAVKRLSKGSQQGVEEFRNEVVLIAKLQHRNLVRLISYCIHEDEKLLIYE 2114
Query: 953 YMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLD 774
Y+PNKSLD FLF+ +++ +LDW RF II+GIARGLLYLH+DSRL ++HRDLKASNILLD
Sbjct: 2115 YLPNKSLDTFLFDAKRKSVLDWTTRFMIIKGIARGLLYLHQDSRLTIIHRDLKASNILLD 2174
Query: 773 ADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILE 594
+M PKISDFGMAR+F G++ Q NT RVVGT+GYMSPEYA+EG FSVKSD Y FGVL+LE
Sbjct: 2175 TNMSPKISDFGMARIFEGNKQQENTTRVVGTYGYMSPEYALEGSFSVKSDTYSFGVLLLE 2234
Query: 593 IITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCV 414
+ AW W + NA +L+D IR SC + +VLRCI IAL CV
Sbjct: 2235 L---------------------AWSLWKDGNAMDLVDSSIRESCLLHEVLRCIQIALSCV 2273
Query: 413 QDHADERPDIPTVILMLSNDSSSLPNPR 330
QD RP + +++ ML N++++LP P+
Sbjct: 2274 QDDPTARPLMSSIVFMLENETAALPTPK 2301
Score = 483 bits (1244), Expect = e-135
Identities = 308/765 (40%), Positives = 416/765 (54%), Gaps = 29/765 (3%)
Frame = -2
Query: 2690 DTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVA 2511
D L G+ + + L+S G+F GFF P S Y+G+W+H+I RTVVWVANR
Sbjct: 20 DQLTLGKPIFPSEMLISKG-GIFALGFFPPANFSNSL-YVGVWFHNIPQRTVVWVANRDN 77
Query: 2510 PATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLE 2331
P T+ S S TL +T ++ + +LW++ + G SAVL DTG+
Sbjct: 78 PITTPS-SATLAITNSSGMV-----LSDSQGDILWTAKISVI-----GASAVLLDTGNFV 126
Query: 2330 VRSEDGV-LWDSFWHPTDTILSGMRITLQAPGRGPKERML--FTSWASETDPSPGRYALG 2160
+R +G +W SF HPTDTIL+GM + K ++ T+W S DPS G ++
Sbjct: 127 LRLANGTDIWQSFDHPTDTILAGMMFLMSY-----KSEIIGRLTAWRSHDDPSTGDFSFS 181
Query: 2159 LDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFT--PAIDPVLGNYYTYT 1986
LDP + Q W +G Y R+G V G + P S F ID YY+YT
Sbjct: 182 LDPSSDLQGMTW-NGTKPYCRNGVRTSVTVSGAQY-PSNSSLFMYQTLIDSGNKLYYSYT 239
Query: 1985 ATNTSLQRFVVLPN-GTDICYMVRKSSQDWELVWYQPS-NECEYYATCGPNAKCTASQDG 1812
+++S+ + L + GT + SS W L++ +P+ CE Y +CGP C +
Sbjct: 240 VSDSSIYTRLTLDSTGTMMFLSWDNSSSSWMLIFQRPAAGSCEVYGSCGPFGYCDFTGAV 299
Query: 1811 KAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPDFSYWVS 1632
A C CL GF P + GC R L C + G F+ + ++K PD +
Sbjct: 300 PA-CRCLDGFEPV-----DPSISQSGCRRKEELRC--GEGGHRFVSLPDMKVPDKFLQIR 351
Query: 1631 TVGDEPGCRTVCLNNCSCGAYVYTATTG---------CLAWGNELIDMHELQTGAYTLNL 1479
+ C C +NCSC AY Y + CL W EL+D + + L L
Sbjct: 352 NRSFDQ-CAAECSSNCSCKAYAYANLSSGGTMADPSRCLVWTGELVDSEKKASLGENLYL 410
Query: 1478 KLPASELRGHHPIWKIATIISAIVLFVLAACLLLWW--KHGRNIKDAVHGSWRSRHSSTQ 1305
+L + + + KI ++ V +L C++L W KH R +
Sbjct: 411 RLAEPPVGKKNRLLKI--VVPITVCMLLLTCIVLTWICKH--------------RGKQNK 454
Query: 1304 SQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYM- 1128
Q ML+ + + G++ + S I AT NF +SN LG GGFG VY
Sbjct: 455 EIQKRLMLEYPGT----SNELGGENVKFPFISFGDIVAATDNFCESNLLGRGGFGKVYKR 510
Query: 1127 ----------GTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPR 978
G L GG EVAVKRL SGQG+EEF+NEV+LIAKLQHRNLVRLLGCCI
Sbjct: 511 FPIYIDDNMKGILEGGTEVAVKRLNEGSGQGIEEFRNEVVLIAKLQHRNLVRLLGCCIHE 570
Query: 977 EEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDL 798
+EK+L+YEY+PNKSLDAFLF+ ++ +LDW RF II+GIA+GLLYLH+DSRL ++HRDL
Sbjct: 571 DEKLLIYEYLPNKSLDAFLFDATRKYVLDWPTRFKIIKGIAKGLLYLHQDSRLTIIHRDL 630
Query: 797 KASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVY 618
KASNILLD +M PKISDFG+AR+F G+Q Q NT RVVGT+GYMSPEY + G FSVKSD Y
Sbjct: 631 KASNILLDTEMNPKISDFGIARIFHGNQQQANTTRVVGTYGYMSPEYVLGGAFSVKSDTY 690
Query: 617 GFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELID 483
FGVL+LEI++G + S + ++ YAWR W + NA EL+D
Sbjct: 691 SFGVLLLEIVSGLKISSSKLTPNFFSLTAYAWRLWKDGNATELLD 735
Score = 330 bits (847), Expect = 5e-89
Identities = 252/769 (32%), Positives = 343/769 (44%), Gaps = 18/769 (2%)
Frame = -2
Query: 2699 QKTDTLRQGESLSGAATLVSS--PEGVFEAGFFAPDPKQPSRQYLGIWYHSISP-RTVVW 2529
Q D L + S A SS + F P P QP + + SP RT VW
Sbjct: 839 QSDDRLTPAKHSSSPAATSSSRMAASLLSVSFPLPPPTQPQACSTLAYGTTTSPERTYVW 898
Query: 2528 VANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQ 2349
VA R P T+ + L +T T L + D + GT ++NT + G GG +AVLQ
Sbjct: 899 VATRDNPITTHTARLAVTNTSGLVLSD---SKGT-------TANTVTIGG--GGATAVLQ 946
Query: 2348 DTGSLEVRSEDGVLWDSFWHPTDTILSGMRITLQAPGRGPK--ERMLFTSWASETDPSPG 2175
+TG+ +R GR K E + +W DPS
Sbjct: 947 NTGNFVLRY---------------------------GRTYKNHEAVRVVAWRGRRDPSTC 979
Query: 2174 RYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDPVLGNYY 1995
++L DP G + G WRSG WNG G L R ++ +D Y
Sbjct: 980 EFSLSGDPDQWGLHIVIWHGASPSWRSGVWNGATATG-----LTRYIWSQIVDNGEEIYA 1034
Query: 1994 TYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQD 1815
Y A + L + + G S W + +P + C +Y CGP C +
Sbjct: 1035 IYNAADGILTHWKLDYTGNVSFRAWNNVSSTWTSPFERPGHGCLHYGACGPFGYCDITGS 1094
Query: 1814 GKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPDFSYWV 1635
+ +C CL GF P N+ S+GC R L C
Sbjct: 1095 FQ-ECKCLDGFEPADGFSLNS---SRGCRRKEELRC------------------------ 1126
Query: 1634 STVGDEPGCRTVCLNNCSCGAYVYT-----ATTG----CLAWGNELIDMHELQTGAYTLN 1482
G C C NCSC AY Y TTG CL W EL+D + L
Sbjct: 1127 ---GPVEECADECDRNCSCTAYAYANLRTILTTGDPSRCLVWMGELLDSEKASAVGENLY 1183
Query: 1481 LKLPASELRGHHPIWKIAT-IISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRH--SS 1311
L+L S + I KI I+ +++ +C++L R I+ R++
Sbjct: 1184 LRLAGSPAVNNKNIVKIVLPAIACLLILTACSCVVLCKCESRGIR-------RNKEVLKK 1236
Query: 1310 TQSQQNSAMLDI-SQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPV 1134
T+ SA D Q++ F D S + + +AT+ F ++N LG+GGFG
Sbjct: 1237 TELGYLSAFHDSWDQNLEFPD------------ISYEDLTSATNGFHETNMLGKGGFG-- 1282
Query: 1133 YMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYE 954
+H+NLVRLLGCCI +EK+L+YE
Sbjct: 1283 -------------------------------------KHKNLVRLLGCCIHGDEKLLIYE 1305
Query: 953 YMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLD 774
Y+PNKSLD FLF+ + ++DW+ RF+II+G+ARGLLYLH+DSR+ ++HRDLK SNILLD
Sbjct: 1306 YLPNKSLDKFLFDHAMKSVIDWQTRFNIIKGVARGLLYLHQDSRMMIIHRDLKTSNILLD 1365
Query: 773 ADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILE 594
A+M PKISDFGMAR+FG + Q +T RVVGT+GYM+PEYAMEGIFSVKSD Y FGVL+LE
Sbjct: 1366 AEMNPKISDFGMARIFGNSEQQASTRRVVGTYGYMAPEYAMEGIFSVKSDTYSFGVLLLE 1425
Query: 593 IITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQV 447
I AW W + A +D ++ SC + +V
Sbjct: 1426 I---------------------AWNLWKDGMAEAFVDKMVLESCLLNEV 1453
>sptr|Q9SYA0|Q9SYA0 T25B24.15 protein.
Length = 829
Score = 585 bits (1509), Expect = e-166
Identities = 339/794 (42%), Positives = 454/794 (57%), Gaps = 12/794 (1%)
Frame = -2
Query: 2666 LSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPS 2487
LS TL S+ E V+E GFF+P+ Q QY+GIW+ PR VVWVANR P T ++
Sbjct: 33 LSMGQTLSSANE-VYELGFFSPNNTQD--QYVGIWFKDTIPRVVVWVANREKPVTDSTAY 89
Query: 2486 LTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEV--RSEDG 2313
L ++ +G L +L+G +GT +WSS T + G A L D+G+L+V +
Sbjct: 90 LAISSSGSLLLLNGK--HGT-----VWSSGVTFSSS---GCRAELSDSGNLKVIDNVSER 139
Query: 2312 VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQA 2133
LW SF H DT+L +T E+ + TSW S TDPSPG + + P Q
Sbjct: 140 ALWQSFDHLGDTLLHTSSLTYNL---ATAEKRVLTSWKSYTDPSPGDFLGQITPQVPSQG 196
Query: 2132 YIWKDGNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGNYYTYTATNTSLQRFV 1956
++ + G+ YWRSG W F GIP+ Y FT D Y TY + L R
Sbjct: 197 FVMR-GSTPYWRSGPWAKTRFTGIPFMDESYTGPFTLHQDVNGSGYLTYFQRDYKLSRIT 255
Query: 1955 VLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHP 1776
+ G+ M R + WEL + P C++Y CGP C S C C +GF P
Sbjct: 256 LTSEGS--IKMFRDNGMGWELYYEAPKKLCDFYGACGPFGLCVMSPS--PMCKCFRGFVP 311
Query: 1775 KLQEQWNAGNWSQGCIRSPPLGCETNQSG---DGFLPMGNIKWPDFSYWVSTVGDEPGCR 1605
K E+W GNW+ GC+R L C N +G D F + NIK PDF + S+V E C
Sbjct: 312 KSVEEWKRGNWTGGCVRHTELDCLGNSTGEDADDFHQIANIKPPDFYEFASSVNAEE-CH 370
Query: 1604 TVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKI-A 1428
C++NCSC A+ Y GCL W +L+D + L+++L SEL G+ I A
Sbjct: 371 QRCVHNCSCLAFAYIKGIGCLVWNQDLMDAVQFSATGELLSIRLARSELDGNKRKKTIVA 430
Query: 1427 TIISAIVLFVLAACLLLWWKH-----GRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSI 1263
+I+S + +L W+ G + + +S ++ A +
Sbjct: 431 SIVSLTLFMILGFTAFGVWRCRVEHIGNILMTLLSNDLLLLFNSFACKRKKAHISKDA-- 488
Query: 1262 RFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLC 1083
+ +D++ L + + I+ AT+NFS SNKLG+GGFG VY G L G+E+AVKRL
Sbjct: 489 -WKNDLKPQDVPGLDFFDMHTIQNATNNFSLSNKLGQGGFGSVYKGKLQDGKEIAVKRLS 547
Query: 1082 RNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQ 903
+SGQG EEF NE++LI+KLQHRNLVR+LGCCI EEK+L+YE+M NKSLD FLF+ K+
Sbjct: 548 SSSGQGKEEFMNEIVLISKLQHRNLVRVLGCCIEEEEKLLIYEFMVNKSLDTFLFDSRKR 607
Query: 902 RLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFG 723
+DW KRFDII+GIARGLLYLH DSRLRV+HRDLK SNILLD M PKISDFG+ARM+
Sbjct: 608 LEIDWPKRFDIIQGIARGLLYLHHDSRLRVIHRDLKVSNILLDEKMNPKISDFGLARMYQ 667
Query: 722 GDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSL 543
G + Q NT RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+G++ F +
Sbjct: 668 GTEYQDNTRRVVGTLGYMSPEYAWTGMFSEKSDIYSFGVLMLEIISGEKISRFSYGVEGK 727
Query: 542 NIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILML 363
+ YAW W+E +L+D + SC +V RCI I LLCVQ +RP+ ++ ML
Sbjct: 728 TLIAYAWESWSEYRGIDLLDQDLADSCHPLEVGRCIQIGLLCVQHQPADRPNTLELLAML 787
Query: 362 SNDSSSLPNPRPPT 321
+ +S LP+P+ PT
Sbjct: 788 TT-TSDLPSPKQPT 800
>sptr|O64781|O64781 T1F9.12.
Length = 831
Score = 585 bits (1509), Expect = e-166
Identities = 340/786 (43%), Positives = 460/786 (58%), Gaps = 11/786 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP+GV+E GFF+P+ + +QY+GIW+ +I+P+ VVWVANR P T + +LT++ G
Sbjct: 56 LSSPDGVYELGFFSPNNSR--KQYVGIWFKNIAPQVVVWVANRDKPVTKTAANLTISSNG 113
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSEDG--VLWDSFW 2292
L +LDGT ++WS T A A L DTG+L V + LW SF
Sbjct: 114 SLILLDGTQ-------DVIWS---TGEAFTSNKCHAELLDTGNLVVIDDVSGKTLWKSFE 163
Query: 2291 HPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGN 2112
+ +T+L + P RG K R+L TSW S +DPSPG + L P Q I + G+
Sbjct: 164 NLGNTMLPQSSVMYDIP-RG-KNRVL-TSWRSNSDPSPGEFTLEFTPQVPPQGLI-RRGS 219
Query: 2111 VTYWRSGQWNGVNFIGIPWRPL-YRSGFTPAIDPVLGNY-YTYTATNTSLQRFVVLPNGT 1938
YWRSG W F GIP Y S FT D G ++Y+ +V L +
Sbjct: 220 SPYWRSGPWAKTRFSGIPGIDASYVSPFTVLQDVAKGTASFSYSMLRNYKLSYVTLTSEG 279
Query: 1937 DICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQW 1758
+ ++ + W+L + P++ C+ Y CGP C S++ KC CLKGF PK ++W
Sbjct: 280 KM-KILWNDGKSWKLHFEAPTSSCDLYRACGPFGLCVRSRN--PKCICLKGFVPKSDDEW 336
Query: 1757 NAGNWSQGCIRSPPLGCETNQSG-------DGFLPMGNIKWPDFSYWVSTVGDEPGCRTV 1599
GNW+ GC+R L C TN S D F M +K PD Y ++ + C
Sbjct: 337 KKGNWTSGCVRRTQLSCHTNSSTKTQGKETDSFYHMTRVKTPDL-YQLAGFLNAEQCYQD 395
Query: 1598 CLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIATII 1419
CL NCSC A+ Y + GCL W EL+D + + +L+L+L +SEL G + I
Sbjct: 396 CLGNCSCTAFAYISGIGCLVWNRELVDTVQFLSDGESLSLRLASSELAGSNRTKIILGTT 455
Query: 1418 SAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDVED 1239
++ +FV+ A + SWR R + Q++ N + SQ + D+E
Sbjct: 456 VSLSIFVILVF-------------AAYKSWRYR--TKQNEPNPMFIHSSQDA-WAKDMEP 499
Query: 1238 GKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLE 1059
+ ++ + IRTAT+NFS SNKLG+GGFGPVY G L G+E+AVKRL +SGQG +
Sbjct: 500 QDVSGVNLFDMHTIRTATNNFSSSNKLGQGGFGPVYKGKLVDGKEIAVKRLSSSSGQGTD 559
Query: 1058 EFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKR 879
EF NE+ LI+KLQH+NLVRLLGCCI EEK+L+YEY+ NKSLD FLF+ + +DW+KR
Sbjct: 560 EFMNEIRLISKLQHKNLVRLLGCCIKGEEKLLIYEYLVNKSLDVFLFDSTLKFEIDWQKR 619
Query: 878 FDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQFNT 699
F+II+G+ARGLLYLHRDSRLRV+HRDLK SNILLD M PKISDFG+ARM G Q Q NT
Sbjct: 620 FNIIQGVARGLLYLHRDSRLRVIHRDLKVSNILLDEKMIPKISDFGLARMSQGTQYQDNT 679
Query: 698 NRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWR 519
RVVGT GYM+PEYA G+FS KSD+Y FGVL+LEII G++ F E+ + YAW
Sbjct: 680 RRVVGTLGYMAPEYAWTGVFSEKSDIYSFGVLLLEIIIGEKISRF--SEEGKTLLAYAWE 737
Query: 518 QWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDSSSLP 339
W E +L+D + S +V RC+ I LLCVQ +RP+ ++ ML+ S LP
Sbjct: 738 SWCETKGVDLLDQALADSSHPAEVGRCVQIGLLCVQHQPADRPNTLELMSMLTT-ISELP 796
Query: 338 NPRPPT 321
+P+ PT
Sbjct: 797 SPKQPT 802
>sptr|Q9SXB4|Q9SXB4 T28P6.6 protein.
Length = 820
Score = 584 bits (1506), Expect = e-165
Identities = 340/827 (41%), Positives = 479/827 (57%), Gaps = 12/827 (1%)
Frame = -2
Query: 2666 LSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPS 2487
L+ + T+VSS F GFF+P + +Y GIWY+S+S +TV+WVAN+ P +S
Sbjct: 36 LNDSETIVSSFR-TFRFGFFSP--VNSTSRYAGIWYNSVSVQTVIWVANKDKPINDSSGV 92
Query: 2486 LTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVR--SEDG 2313
++++ G+L V DG +LWS+N +++A + +L D+G+L ++ S D
Sbjct: 93 ISVSQDGNLVVTDGQRR-------VLWSTNVSTQASANSTVAELL-DSGNLVLKEASSDA 144
Query: 2312 VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQA 2133
LW+SF +PTD+ L M + A G + TSW S +DPSPG Y L +
Sbjct: 145 YLWESFKYPTDSWLPNMLVGTNARIGGGN--VTITSWKSPSDPSPGSYTAALVLAAYPEL 202
Query: 2132 YIWKDGN--VTYWRSGQWNGVNFIGIPWRPLYRSGFTPAI---DPVLGNYYTYTATNTSL 1968
+I + N T WRSG WNG F G+P +Y F D G+ A +++L
Sbjct: 203 FIMNNNNNNSTVWRSGPWNGQMFNGLP--DVYAGVFLYRFIVNDDTNGSVTMSYANDSTL 260
Query: 1967 QRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLK 1788
+ F + G+ I ++ ++W + P+ EC+ Y CG A C ++ C+C++
Sbjct: 261 RYFYMDYRGSVIRRDWSETRRNWTVGLQVPATECDNYRRCGEFATCNPRKN--PLCSCIR 318
Query: 1787 GFHPKLQEQWNAGNWSQGCIRSPPLGCE---TNQSGDGFLPMGNIKWPDFSYWVSTVGDE 1617
GF P+ +WN GNWS GC R PL CE N S DGFL + +K PDF+ + E
Sbjct: 319 GFRPRNLIEWNNGNWSGGCTRRVPLQCERQNNNGSADGFLRLRRMKLPDFAR--RSEASE 376
Query: 1616 PGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELR--GHHP 1443
P C CL CSC A + GC+ W L+D EL L ++L SE++ P
Sbjct: 377 PECLRTCLQTCSCIAAAHGLGYGCMIWNGSLVDSQELSASGLDLYIRLAHSEIKTKDKRP 436
Query: 1442 IWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSI 1263
I I TI++ + FV+AAC+LL R I + R + ++ A+ ++
Sbjct: 437 IL-IGTILAGGI-FVVAACVLL----ARRIVMKKRAKKKGRDAEQIFERVEALAGGNK-- 488
Query: 1262 RFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLC 1083
GK EL ++ + AT+NFS NKLG+GGFGPVY G L G+E+AVKRL
Sbjct: 489 --------GKLKELPLFEFQVLAAATNNFSLRNKLGQGGFGPVYKGKLQEGQEIAVKRLS 540
Query: 1082 RNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQ 903
R SGQGLEE NEV++I+KLQHRNLV+LLGCCI EE++LVYE+MP KSLD +LF+ +
Sbjct: 541 RASGQGLEELVNEVVVISKLQHRNLVKLLGCCIAGEERMLVYEFMPKKSLDYYLFDSRRA 600
Query: 902 RLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFG 723
+LLDWK RF+II GI RGLLYLHRDSRLR++HRDLKASNILLD ++ PKISDFG+AR+F
Sbjct: 601 KLLDWKTRFNIINGICRGLLYLHRDSRLRIIHRDLKASNILLDENLIPKISDFGLARIFP 660
Query: 722 GDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSL 543
G++++ NT RVVGT+GYM+PEYAM G+FS KSDV+ GV++LEII+G+R +
Sbjct: 661 GNEDEANTRRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIISGRR-------NSNS 713
Query: 542 NIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILML 363
+ Y W WNE L+DP I +++ +CIHI LLCVQ+ A++RP + TV ML
Sbjct: 714 TLLAYVWSIWNEGEINSLVDPEIFDLLFEKEIHKCIHIGLLCVQEAANDRPSVSTVCSML 773
Query: 362 SNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
S++ + +P P+ P + R I VT+T + GR
Sbjct: 774 SSEIADIPEPKQPAFISRNNVPEAESSENSDLKDSINNVTITDVTGR 820
>sptr|Q9SY95|Q9SY95 T25B24.10 protein.
Length = 802
Score = 582 bits (1501), Expect = e-165
Identities = 332/781 (42%), Positives = 450/781 (57%), Gaps = 6/781 (0%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G+FE GFF+P+ + Y+GIW+ I PRTVVWVANR T A+ L ++ G
Sbjct: 33 LSSPNGIFELGFFSPNNSR--NLYVGIWFKGIIPRTVVWVANRENSVTDATADLAISSNG 90
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE-DGV-LWDSFW 2292
L + DG + +WS+ T + G SA L D+G+L V + G+ LW SF
Sbjct: 91 SLLLFDGKHST-------VWSTGETFASN---GSSAELSDSGNLLVIDKVSGITLWQSFE 140
Query: 2291 HPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGN 2112
H DT+L + + PG G E+ + +SW S TDP PG + + Q +I + G+
Sbjct: 141 HLGDTMLPYSSL-MYNPGTG--EKRVLSSWKSYTDPLPGEFVGYITTQVPPQGFIMR-GS 196
Query: 2111 VTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGNYYTYTATNTSLQRFVVLPNGTD 1935
YWRSG W F G+P Y F+ D Y+++ N V+ G+
Sbjct: 197 KPYWRSGPWAKTRFTGVPLTDESYTHPFSVQQDANGSVYFSHLQRNFKRSLLVLTSEGS- 255
Query: 1934 ICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWN 1755
+ + DW L P+N C++Y CGP C S KC C KGF P+ E+W
Sbjct: 256 -LKVTHHNGTDWVLNIDVPANTCDFYGVCGPFGLCVMSIP--PKCKCFKGFVPQFSEEWK 312
Query: 1754 AGNWSQGCIRSPPLGCETNQSG---DGFLPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNC 1584
GNW+ GC+R L C+ N +G + F P+ NIK PDF +VS+ G C CL+NC
Sbjct: 313 RGNWTGGCVRRTELLCQGNSTGRHVNVFHPVANIKPPDFYEFVSS-GSAEECYQSCLHNC 371
Query: 1583 SCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIATIISAIVL 1404
SC A+ Y GCL W EL+D+ + G L+++L +SE+ G+ I I +I L
Sbjct: 372 SCLAFAYINGIGCLIWNQELMDVMQFSVGGELLSIRLASSEMGGNQRKKTIIASIVSISL 431
Query: 1403 FVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHE 1224
FV A A G WR R + N+ + +S + +D++
Sbjct: 432 FVTLA-------------SAAFGFWRYR-----LKHNAIVSKVSLQGAWRNDLKSEDVSG 473
Query: 1223 LKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNE 1044
L + + I AT+NFS NKLG+GGFGPVY G L G+E+AVKRL +SGQG EEF NE
Sbjct: 474 LYFFEMKTIEIATNNFSLVNKLGQGGFGPVYKGKLQDGKEIAVKRLSSSSGQGKEEFMNE 533
Query: 1043 VILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIE 864
++LI+KLQH NLVR+LGCCI EE++LVYE+M NKSLD F+F+ K+ +DW KRF II+
Sbjct: 534 ILLISKLQHINLVRILGCCIEGEERLLVYEFMVNKSLDTFIFDSRKRVEIDWPKRFSIIQ 593
Query: 863 GIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVG 684
GIARGLLYLHRDSRLR++HRD+K SNILLD M PKISDFG+ARM+ G + Q NT R+VG
Sbjct: 594 GIARGLLYLHRDSRLRIIHRDVKVSNILLDDKMNPKISDFGLARMYEGTKYQDNTRRIVG 653
Query: 683 TFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNED 504
T GYMSPEYA G+FS KSD Y FGVL+LE+I+G++ F ++ N+ YAW W E+
Sbjct: 654 TLGYMSPEYAWTGVFSEKSDTYSFGVLLLEVISGEKISRFSYDKERKNLLAYAWESWCEN 713
Query: 503 NAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPP 324
+D SC +V RC+ I LLCVQ +RP+ ++ ML+ +S LP P+ P
Sbjct: 714 GGVGFLDKDATDSCHPSEVGRCVQIGLLCVQHQPADRPNTLELLSMLTT-TSDLPLPKEP 772
Query: 323 T 321
T
Sbjct: 773 T 773
>sptr|Q8H7E6|Q8H7E6 Hypothetical protein.
Length = 808
Score = 582 bits (1500), Expect = e-164
Identities = 341/823 (41%), Positives = 466/823 (56%), Gaps = 15/823 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ QY+GIW+ I+PR VVWVANR P T+ +LT++ G
Sbjct: 42 LSSPGGFYELGFFSPNNSH--NQYVGIWFKKITPRVVVWVANREKPITNPVANLTISRNG 99
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LD + ++WS T R A L DTG+L + + + +LW SF
Sbjct: 100 SLILLDSSKN-------VVWS---TRRPSISNKCHAKLLDTGNLVIVDDVSENLLWQSFE 149
Query: 2291 HPTDTIL--SGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKD 2118
+P DT+L S + L A G E+ + +SW S TDPSPG + + L P Q +
Sbjct: 150 NPGDTMLPYSSLMYNL-ATG----EKRVLSSWKSHTDPSPGDFVVRLTPQVPAQIVTMR- 203
Query: 2117 GNVTYWRSGQWNGVNFIGIPWRP-LYRSGFTPAIDPVLGN-YYTYTATNTSLQRFVVLPN 1944
G+ Y RSG W F G+P Y S F+ + D G ++Y ++ L R ++
Sbjct: 204 GSSVYKRSGPWAKTGFTGVPLMDESYTSPFSLSQDVGNGTGLFSYLQRSSELTRVIITSE 263
Query: 1943 GTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQE 1764
G R + W L + P+N C+ Y CGP C S KC C+KGF PK +E
Sbjct: 264 G--YLKTFRYNGTGWVLDFITPANLCDLYGACGPFGLCVTSNP--TKCKCMKGFVPKYKE 319
Query: 1763 QWNAGNWSQGCIRSPPLGCETNQSG-------DGFLPMGNIKWPDFSYWVSTVGDEPGCR 1605
+W GN + GC+R L C+ N S D F + N+K PD + S V D C
Sbjct: 320 EWKRGNMTSGCMRRTELSCQANLSTKTQGKGVDVFYRLANVKPPDLYEYASFV-DADQCH 378
Query: 1604 TVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKIAT 1425
CL+NCSC A+ Y GCL W +ELID G L+++L +SEL G+ I
Sbjct: 379 QGCLSNCSCSAFAYITGIGCLLWNHELIDTVRYSVGGEFLSIRLASSELAGNRRTKIIVG 438
Query: 1424 IISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDV 1245
IS + +LA +W++ D SW++ +
Sbjct: 439 SISLSIFVILAFGSYKYWRYRAKQND----SWKN------------------------GL 470
Query: 1244 EDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQG 1065
E + L + ++ IRTAT+NF+ SNKLG+GGFGPVY GTL +++AVKRL +SGQG
Sbjct: 471 EPQEISGLTFFEMNTIRTATNNFNVSNKLGQGGFGPVYKGTLSDKKDIAVKRLSSSSGQG 530
Query: 1064 LEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWK 885
EEF NE+ LI+KLQHRNLVRLLGCCI EEK+L+YE++ NKSLD FLF+ + +DW
Sbjct: 531 TEEFMNEIKLISKLQHRNLVRLLGCCIDGEEKLLIYEFLVNKSLDTFLFDLTLKLQIDWP 590
Query: 884 KRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQF 705
KRF+II+G++RGLLYLHRDS +RV+HRDLK SNILLD M PKISDFG+ARMF G Q++
Sbjct: 591 KRFNIIQGVSRGLLYLHRDSCMRVIHRDLKVSNILLDEKMNPKISDFGLARMFQGTQHKT 650
Query: 704 NTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYA 525
RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+GK+ SF C E+ + G+A
Sbjct: 651 TLVRVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIISGKKISSFCCGEEGKTLLGHA 710
Query: 524 WRQWNEDNAAELIDPVIRASCS--VRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDS 351
W W E +L+D I +SCS +V RC+ I LLC+Q A +RP+I V+ M+++ +
Sbjct: 711 WECWLETGGVDLLDEDISSSCSPVEVEVARCVQIGLLCIQQQAIDRPNIAQVVTMMTS-A 769
Query: 350 SSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
+ LP P+ P L+ + VT T+++GR
Sbjct: 770 TDLPRPKQPLFALQ----IQDQESVVSVSKSVNHVTQTEIYGR 808
>sptr|Q8L454|Q8L454 Putative serine/threonine kinase.
Length = 807
Score = 580 bits (1494), Expect = e-164
Identities = 343/825 (41%), Positives = 463/825 (56%), Gaps = 20/825 (2%)
Frame = -2
Query: 2720 SLCCVAAQ-KTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISP 2544
S CC+++ TD+L + +S T+VS+ E F GFF+P + +Y+GIWY ++
Sbjct: 24 STCCLSSTITTDSLLPNKQISDGQTIVSANE-TFTLGFFSPGTS--TYRYVGIWYSNVPN 80
Query: 2543 RTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGY 2364
RTVVWVANR P S L +G+L +LDG SS T +
Sbjct: 81 RTVVWVANRNNPVLDTSGILMFDTSGNLVILDGRG-----------SSFTVAYGSGAKDT 129
Query: 2363 SAVLQDTGSLEVRSEDG---VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASE 2193
A + D+G+L +RS + W SF +PTDT L GM + G + L TSW S
Sbjct: 130 EATILDSGNLVLRSVSNRSRLRWQSFDYPTDTWLQGMNL-----GFVGAQNQLLTSWRSS 184
Query: 2192 TDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPAIDP 2013
DP+ G Y+ G+DP G +IW+ GNV YW+SG WNG ++ FT +
Sbjct: 185 DDPAIGDYSFGMDPNEKGDFFIWERGNV-YWKSGLWNGQSY-----------NFTESES- 231
Query: 2012 VLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQ------------DWELVWYQPSNE 1869
+ Y T+L + +P + Y++ S Q W ++ P
Sbjct: 232 -MSFLYVSNDARTTLS-YSSIPASGMVRYVLDHSGQLKLLERMDFVLHQWLVLGSWPEGS 289
Query: 1868 CEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSG 1689
C+ Y+ CG C +QD + +C C KGF+P W++G+ +GCIR + C G
Sbjct: 290 CKAYSPCGAFGICAGNQDWQNRCKCPKGFNPGDGVGWSSGDTRRGCIRQTNMHCV----G 345
Query: 1688 DGFLPMGNIKWPDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHE 1509
D F M ++ P + +S++ + C + CL NCSC AY C W ++++ E
Sbjct: 346 DKFFQMPDMGLPGNATTISSITGQKQCESTCLTNCSCTAYAVLQDK-CSLWYGNIMNLRE 404
Query: 1508 LQTG--AYTLNLKLPASELRGHH-PIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVH 1338
++G T L+L ASEL P+ IA +S++ + A+ + LW
Sbjct: 405 GESGDAVGTFYLRLAASELESRGTPVVLIAATVSSVAFLIFASLIFLWM----------- 453
Query: 1337 GSWRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKL 1158
WR + + +D +I+ + E G SH + I AT FS NKL
Sbjct: 454 --WRQKSKAKG-------VDTDSAIKLWESEETG-SH-FTSFCFSEIADATCKFSLENKL 502
Query: 1157 GEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPR 978
GEGGFGPVY G LP G+E+AVKRL +SGQGL EFKNE++LIAKLQHRNLVRLLGCCI
Sbjct: 503 GEGGFGPVYKGNLPEGQEIAVKRLAAHSGQGLLEFKNEIMLIAKLQHRNLVRLLGCCIQG 562
Query: 977 EEKILVYEYMPNKSLDAFLFNPEK-QRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRD 801
EEKIL+YEYMPNKSLD FLF + Q L+ IIEGIA+GLLYLH+ SR R++HRD
Sbjct: 563 EEKILIYEYMPNKSLDFFLFAGQVIQCGLE-----GIIEGIAQGLLYLHKHSRFRIIHRD 617
Query: 800 LKASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDV 621
LKASNILLD DM PKISDFGMAR+FG + + NTNRVVGT+GYM+PEYAMEGIFSVKSDV
Sbjct: 618 LKASNILLDIDMNPKISDFGMARIFGSKETEANTNRVVGTYGYMAPEYAMEGIFSVKSDV 677
Query: 620 YGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLR 441
+ FGVL+LEI++G R FH +SLN+ YAW W E +EL DP I +C +VLR
Sbjct: 678 FSFGVLLLEIVSGIRNAGFHQRGNSLNLLCYAWELWKEGRWSELADPSIYNACPEHKVLR 737
Query: 440 CIHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLRG 306
CIH+ L+CVQ+ RP + +I L N+S++LP P+ P + G
Sbjct: 738 CIHVGLMCVQESPINRPTMTEIISALDNESTTLPEPKQPAFVSAG 782
>sptr|Q9SXB5|Q9SXB5 T28P6.5 protein.
Length = 820
Score = 579 bits (1493), Expect = e-164
Identities = 331/830 (39%), Positives = 474/830 (57%), Gaps = 14/830 (1%)
Frame = -2
Query: 2669 SLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASP 2490
+L+ + T+VSS F GFF+P + +Y GIWY+SI +TV+WVAN+ P +S
Sbjct: 35 TLNDSETIVSSFR-TFRFGFFSP--VNSTNRYAGIWYNSIPVQTVIWVANKDTPINDSSG 91
Query: 2489 SLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVR--SED 2316
++++ G+L V DG +LWS+N ++RA + +L+ +G+L ++ + D
Sbjct: 92 VISISEDGNLVVTDGQRR-------VLWSTNVSTRASANSTVAELLE-SGNLVLKDANTD 143
Query: 2315 GVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQ 2136
LW+SF +PTD+ L M + A G + TSW + +DPSPG Y L +
Sbjct: 144 AYLWESFKYPTDSWLPNMLVGTNARTGGGN--ITITSWTNPSDPSPGSYTAALVLAPYPE 201
Query: 2135 AYIWK--DGNVTYWRSGQWNGVNFIGIP--WRPLYRSGFTPAIDPVLGNYYTYTATNTSL 1968
+I+ D N T WRSG WNG+ F G+P + L+ F D G+ A +++L
Sbjct: 202 LFIFNNNDNNATVWRSGPWNGLMFNGLPDVYPGLFLYRFK-VNDDTNGSATMSYANDSTL 260
Query: 1967 QRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLK 1788
+ + G I ++ ++W L P+ EC+ Y+ CG C ++ C+C+K
Sbjct: 261 RHLYLDYRGFAIRRDWSEARRNWTLGSQVPATECDIYSRCGQYTTCNPRKN--PHCSCIK 318
Query: 1787 GFHPKLQEQWNAGNWSQGCIRSPPLGCETNQ---SGDGFLPMGNIKWPDFSYWVSTVGDE 1617
GF P+ +WN GNWS GCIR PL CE S D FL + +K PDF+ + E
Sbjct: 319 GFRPRNLIEWNNGNWSGGCIRKLPLQCERQNNKGSADRFLKLQRMKMPDFAR--RSEASE 376
Query: 1616 PGCRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELR--GHHP 1443
P C CL +CSC A+ + GC+ W L+D L L+++L SE + P
Sbjct: 377 PECFMTCLQSCSCIAFAHGLGYGCMIWNRSLVDSQVLSASGMDLSIRLAHSEFKTQDRRP 436
Query: 1442 IWKIATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSI 1263
I ++ I FV+A C+LL +R + + D Q
Sbjct: 437 ILIGTSLAGGI--FVVATCVLL-----------------ARRIVMKKRAKKKGTDAEQIF 477
Query: 1262 RFDDDVEDG---KSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVK 1092
+ + + G K EL ++ + TAT NFS SNKLG+GGFGPVY G L G+E+AVK
Sbjct: 478 KRVEALAGGSREKLKELPLFEFQVLATATDNFSLSNKLGQGGFGPVYKGMLLEGQEIAVK 537
Query: 1091 RLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNP 912
RL + SGQGLEE EV++I+KLQHRNLV+L GCCI EE++LVYE+MP KSLD ++F+P
Sbjct: 538 RLSQASGQGLEELVTEVVVISKLQHRNLVKLFGCCIAGEERMLVYEFMPKKSLDFYIFDP 597
Query: 911 EKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMAR 732
+ +LLDW RF+II GI RGLLYLHRDSRLR++HRDLKASNILLD ++ PKISDFG+AR
Sbjct: 598 REAKLLDWNTRFEIINGICRGLLYLHRDSRLRIIHRDLKASNILLDENLIPKISDFGLAR 657
Query: 731 MFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHE 552
+F G++++ NT RVVGT+GYM+PEYAM G+FS KSDV+ GV++LEII+G+R
Sbjct: 658 IFPGNEDEANTRRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIISGRR-------N 710
Query: 551 DSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVI 372
+ + W WNE ++DP I +++ +C+HIALLCVQD A++RP + TV
Sbjct: 711 SHSTLLAHVWSIWNEGEINGMVDPEIFDQLFEKEIRKCVHIALLCVQDAANDRPSVSTVC 770
Query: 371 LMLSNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXXXXIGTVTMTQLHGR 222
+MLS++ + +P P+ P M R I VT+T + GR
Sbjct: 771 MMLSSEVADIPEPKQPAFMPRNVGLEAEFSESIALKASINNVTITDVSGR 820
>sptr|O64782|O64782 T1F9.13 (At1g61380/T1F9_13).
Length = 805
Score = 575 bits (1481), Expect = e-162
Identities = 341/788 (43%), Positives = 445/788 (56%), Gaps = 14/788 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ Q QY+GIW+ I PR VVWVANR P TS++ +LT++ G
Sbjct: 35 LSSPGGFYELGFFSPNNTQ--NQYVGIWFKKIVPRVVVWVANRDTPVTSSAANLTISSNG 92
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LDG ++WS T +A A L DTG+ V + LW SF
Sbjct: 93 SLILLDGKQ-------DVIWS---TGKAFTSNKCHAELLDTGNFVVIDDVSGNKLWQSFE 142
Query: 2291 HPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGN 2112
H +T+L + +L K+R+L T+W S +DPSPG ++L + P Q I + G+
Sbjct: 143 HLGNTMLP--QSSLMYDTSNGKKRVL-TTWKSNSDPSPGEFSLEITPQIPTQGLI-RRGS 198
Query: 2111 VTYWRSGQWNGVNFIGIPWRPL-YRSGFTPAIDPVLGN-YYTY-TATNTSLQRFVVLPNG 1941
V YWR G W F GI Y S F+ D G ++Y T N +L + P G
Sbjct: 199 VPYWRCGPWAKTRFSGISGIDASYVSPFSVVQDTAAGTGSFSYSTLRNYNLSYVTLTPEG 258
Query: 1940 TDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQ 1761
++ +W+L P N C+ Y CGP C S KC CLKGF PK E+
Sbjct: 259 K--MKILWDDGNNWKLHLSLPENPCDLYGRCGPYGLCVRSDP--PKCECLKGFVPKSDEE 314
Query: 1760 WNAGNWSQGCIRSPPLGCETNQS-------GDGFLPMGNIKWPDFSYWVSTVGDEPGCRT 1602
W GNW+ GC+R L C+ S D F M ++K PD + S + E C
Sbjct: 315 WGKGNWTSGCVRRTKLSCQAKSSMKTQGKDTDIFYRMTDVKTPDLHQFASFLNAEQ-CYQ 373
Query: 1601 VCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRG--HHPIWKIA 1428
CL NCSC A+ Y + GCL W EL D + + L ++L +SEL G I
Sbjct: 374 GCLGNCSCTAFAYISGIGCLVWNGELADTVQFLSSGEFLFIRLASSELAGSSRRKIIVGT 433
Query: 1427 TIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDD 1248
T+ +I L ++ A ++LW R R + +N + F
Sbjct: 434 TVSLSIFLILVFAAIMLW---------------RYRAKQNDAWKNGFERQDVSGVNF--- 475
Query: 1247 VEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQ 1068
+ + IRTAT+NFS SNKLG+GGFGPVY G L G+E+ VKRL +SGQ
Sbjct: 476 -----------FEMHTIRTATNNFSPSNKLGQGGFGPVYKGKLVDGKEIGVKRLASSSGQ 524
Query: 1067 GLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDW 888
G EEF NE+ LI+KLQHRNLVRLLG CI EEK+L+YE+M NKSLD F+F+P + LDW
Sbjct: 525 GTEEFMNEITLISKLQHRNLVRLLGYCIDGEEKLLIYEFMVNKSLDIFIFDPCLKFELDW 584
Query: 887 KKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQ 708
KRF+II+GIARGLLYLHRDSRLRV+HRDLK SNILLD M PKISDFG+ARMF G Q Q
Sbjct: 585 PKRFNIIQGIARGLLYLHRDSRLRVIHRDLKVSNILLDDRMNPKISDFGLARMFQGTQYQ 644
Query: 707 FNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGY 528
NT RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+GKR F ++S + Y
Sbjct: 645 DNTRRVVGTLGYMSPEYAWAGLFSEKSDIYSFGVLMLEIISGKRISRFIYGDESKGLLAY 704
Query: 527 AWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDSS 348
W W E + L+D + +C +V RC+ I LLCVQ A +RP+ V+ ML++ ++
Sbjct: 705 TWDSWCETGGSNLLDRDLTDTCQAFEVARCVQIGLLCVQHEAVDRPNTLQVLSMLTS-AT 763
Query: 347 SLPNPRPP 324
LP P+ P
Sbjct: 764 DLPVPKQP 771
>sptr|Q8LEY0|Q8LEY0 Receptor kinase, putative.
Length = 805
Score = 573 bits (1478), Expect = e-162
Identities = 340/788 (43%), Positives = 445/788 (56%), Gaps = 14/788 (1%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVTG 2466
+SSP G +E GFF+P+ + QY+GIW+ I PR VVWVANR P TS++ +LT++ G
Sbjct: 35 LSSPGGFYELGFFSPNNTR--NQYVGIWFKKIVPRVVVWVANRDTPVTSSAANLTISSNG 92
Query: 2465 DLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSE--DGVLWDSFW 2292
L +LDG ++WS T +A A L DTG+ V + LW SF
Sbjct: 93 SLILLDGKE-------DVIWS---TGKAFSSNKCHAQLLDTGNFVVIDDVSGNKLWQSFE 142
Query: 2291 HPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSGQAYIWKDGN 2112
H +T+L + +L K+R+L T+W S +DPSPG ++L + P Q I + G+
Sbjct: 143 HLGNTMLP--QSSLMYDTSNGKKRVL-TTWKSNSDPSPGEFSLEITPQIPTQGLI-RRGS 198
Query: 2111 VTYWRSGQWNGVNFIGIPWRPL-YRSGFTPAIDPVLGN-YYTY-TATNTSLQRFVVLPNG 1941
V YWR G W F GI Y S F+ D G ++Y T N +L + P G
Sbjct: 199 VPYWRCGPWAKTRFSGISGIDASYVSPFSVVQDTAAGTGSFSYSTLRNYNLSYVTLTPEG 258
Query: 1940 TDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFHPKLQEQ 1761
++ DW+L P N C+ Y CGP C S KC CLKGF PK E+
Sbjct: 259 Q--MKILWDDGNDWKLHLSLPENPCDLYGRCGPYGLCVRSDP--PKCECLKGFVPKSDEE 314
Query: 1760 WNAGNWSQGCIRSPPLGCETNQS-------GDGFLPMGNIKWPDFSYWVSTVGDEPGCRT 1602
W GNW+ GC+R L C+ S D F M ++K PD + S + E C
Sbjct: 315 WGKGNWTSGCVRRTKLSCQAKSSMKTQGKDTDIFYRMTDVKTPDLHQFASFLNAEQ-CYQ 373
Query: 1601 VCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRG--HHPIWKIA 1428
CL NCSC A+ Y + GCL W EL D + + L ++L +SEL G I
Sbjct: 374 GCLGNCSCTAFAYISGIGCLVWNGELADTVQFLSSGEILFIRLASSELAGSSRRKIIVGT 433
Query: 1427 TIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDD 1248
T+ +I L ++ A ++LW R R + +N + F
Sbjct: 434 TVSLSIFLILVFAAIMLW---------------RYRAKQNDAWKNGFERQDVSGVNF--- 475
Query: 1247 VEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQ 1068
+ + IRTAT+NFS SNKLG+GGFGPVY G L G+E+ VKRL +SGQ
Sbjct: 476 -----------FEMHTIRTATNNFSPSNKLGQGGFGPVYKGKLVDGKEIGVKRLASSSGQ 524
Query: 1067 GLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDW 888
G EEF NE+ LI+KLQHRNLVRLLG CI EEK+L+YE+M NKSLD F+F+P + LDW
Sbjct: 525 GTEEFMNEITLISKLQHRNLVRLLGYCIDGEEKLLIYEFMVNKSLDIFIFDPCLKFELDW 584
Query: 887 KKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQ 708
KRF+II+GIARGLLYLHRDSRLRV+HR+LK SNILLD M PKISDFG+ARMF G Q Q
Sbjct: 585 PKRFNIIQGIARGLLYLHRDSRLRVIHRNLKVSNILLDDRMNPKISDFGLARMFQGTQYQ 644
Query: 707 FNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGY 528
NT RVVGT GYMSPEYA G+FS KSD+Y FGVL+LEII+GKR F ++S + Y
Sbjct: 645 DNTRRVVGTLGYMSPEYAWAGLFSEKSDIYSFGVLMLEIISGKRISRFIYGDESKGLLAY 704
Query: 527 AWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILMLSNDSS 348
W W E + L+D + +C +V RC+ I LLCVQ A +RP+ V+ ML++ ++
Sbjct: 705 TWDSWCETGGSNLLDRDLTDTCQAFEVARCVQIGLLCVQHEAVDRPNTLQVLSMLTS-AT 763
Query: 347 SLPNPRPP 324
LP P+ P
Sbjct: 764 DLPVPKQP 771
>sptr|Q9ZR08|Q9ZR08 Putative receptor kinase.
Length = 852
Score = 571 bits (1472), Expect = e-161
Identities = 338/839 (40%), Positives = 473/839 (56%), Gaps = 37/839 (4%)
Frame = -2
Query: 2714 CCVAAQKTDTLRQGESL---SGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISP 2544
C VA Q + TL +G +L S TLVS+ + FE GFF P+ R+YLGIW++++ P
Sbjct: 19 CFVAVQDSKTLFKGSTLINDSHGETLVSAGQR-FELGFFTPNGSSDERRYLGIWFYNLHP 77
Query: 2543 RTVVWVANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGY 2364
TVVWVANR +P S T++ G+L V+D + W + +
Sbjct: 78 LTVVWVANRESPVLDRSCIFTISKDGNLEVIDSKGR-------VYWDTGVKP-SSVSAER 129
Query: 2363 SAVLQDTGSLEVRS---EDGVLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASE 2193
L D G+L + S E V+W SF +PTDT L GMR+ E M +SW S
Sbjct: 130 MVKLMDNGNLVLISDGNEANVVWQSFQNPTDTFLPGMRMD---------ENMTLSSWRSF 180
Query: 2192 TDPSPGRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPL----YRSGFTP 2025
DPS G + +D Q IWK ++ YW+SG FIG P + S FT
Sbjct: 181 NDPSHGNFTFQMDQEEDKQFIIWKR-SMRYWKSGISG--KFIGSDEMPYAISYFLSNFTE 237
Query: 2024 AI---DPVLGNYYTYTATNTSLQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYA 1854
+ + + +T TNT RF + +G Y + W +W +P +EC Y
Sbjct: 238 TVTVHNASVPPLFTSLYTNT---RFTMSSSG-QAQYFRLDGERFWAQIWAEPRDECSVYN 293
Query: 1853 TCGPNAKCTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPL-GCETNQSGDGFL 1677
CG C + + C CL GF P E+W G++S GC R + G + GD FL
Sbjct: 294 ACGNFGSCNSKNE--EMCKCLPGFRPNFLEKWVKGDFSGGCSRESRICGKDGVVVGDMFL 351
Query: 1676 PMGNIKW--PDFSYWVSTVGDEPGCRTVCLNNCSCGAYVYT------ATTGCLAWGNELI 1521
+ ++ PD + +E CR CLNNC C AY Y + T C W +L
Sbjct: 352 NLSVVEVGSPDSQF---DAHNEKECRAECLNNCQCQAYSYEEVDILQSNTKCWIWLEDLN 408
Query: 1520 DMHELQTGAYTLNLKLPASELRGHHPIWK--------------IATIISAIVLFVLAACL 1383
++ E G+ + +++ ++ H + + T SA +L VL++
Sbjct: 409 NLKEGYLGSRNVFIRVAVPDIGSHVERGRGRYGEAKTPVVLIIVVTFTSAAILVVLSSTA 468
Query: 1382 LLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLD 1203
+ R + + R H + + ++ +S RF D D + ++ + L+
Sbjct: 469 SYVFLQRRKVNKELGSIPRGVHLCDSERH---IKELIESGRFKQD--DSQGIDVPSFELE 523
Query: 1202 RIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKL 1023
I ATSNFS++NKLG+GGFGPVY G PG +E+AVKRL R SGQGLEEFKNEV+LIAKL
Sbjct: 524 TILYATSNFSNANKLGQGGFGPVYKGMFPGDQEIAVKRLSRCSGQGLEEFKNEVVLIAKL 583
Query: 1022 QHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLL 843
QHRNLVRLLG C+ EEK+L+YEYMP+KSLD F+F+ + + LDWK R +II GIARGLL
Sbjct: 584 QHRNLVRLLGYCVAGEEKLLLYEYMPHKSLDFFIFDRKLCQRLDWKMRCNIILGIARGLL 643
Query: 842 YLHRDSRLRVVHRDLKASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSP 663
YLH+DSRLR++HRDLK SNILLD +M PKISDFG+AR+FGG + NTNRVVGT+GYMSP
Sbjct: 644 YLHQDSRLRIIHRDLKTSNILLDEEMNPKISDFGLARIFGGSETSANTNRVVGTYGYMSP 703
Query: 662 EYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELID 483
EYA+EG+FS KSDV+ FGV+++E I+GKR FH E SL++ G+AW W + EL+D
Sbjct: 704 EYALEGLFSFKSDVFSFGVVVIETISGKRNTGFHEPEKSLSLLGHAWDLWKAERGIELLD 763
Query: 482 PVIRASCSVRQVLRCIHIALLCVQDHADERPDIPTVILML-SNDSSSLPNPRPPTLMLR 309
++ SC L+C+++ LLCVQ+ ++RP + V+ ML S+++++LP P+ P +LR
Sbjct: 764 QALQESCETEGFLKCLNVGLLCVQEDPNDRPTMSNVVFMLGSSEAATLPTPKQPAFVLR 822
>sptr|O81833|O81833 Putative receptor protein kinase.
Length = 815
Score = 571 bits (1472), Expect = e-161
Identities = 330/800 (41%), Positives = 464/800 (58%), Gaps = 26/800 (3%)
Frame = -2
Query: 2645 VSSPEGVFEAGFFAPDPK-QPSRQYLGIWYHSISPRTVVWVANRVAPATSASPSLTLTVT 2469
+SSP+ VF+ GFF+ D + QP ++LG+WY + P VVWVANR P S L L+
Sbjct: 40 LSSPDQVFQLGFFSLDQEEQPQHRFLGLWY--MEPFAVVWVANRNNPLYGTSGFLNLSSL 97
Query: 2468 GDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSEDG---VLWDS 2298
GDL++ DG + LWSS+++S + + +L+ + S + S DG VLW S
Sbjct: 98 GDLQLFDG-------EHKALWSSSSSSTKASKTANNPLLKISCSGNLISSDGEEAVLWQS 150
Query: 2297 FWHPTDTILSGMRITLQAPGRGPKERMLFT--SWASETDPSPGRYALGLDPGNSGQAYIW 2124
F +P +TIL+GM++ G+ K +M ++ SW + DPSPG + L LD Q +
Sbjct: 151 FDYPMNTILAGMKL-----GKNFKTQMEWSLSSWKTLKDPSPGDFTLSLDTRGLPQLILR 205
Query: 2123 KDGNVTY-WRSGQWNGVNFIGIPW----RPLYRSGFTPAIDPVLGNYYTYTATNTSLQRF 1959
K+G+ +Y +R G WNG++F G P L+ FT + V Y++T + + R
Sbjct: 206 KNGDSSYSYRLGSWNGLSFTGAPAMGRENSLFDYKFTSSAQEV---NYSWTPRHRIVSR- 261
Query: 1958 VVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGFH 1779
+VL N + ++ W L P +EC+YY+ CG A C + C+CL+GF
Sbjct: 262 LVLNNTGKLHRFIQSKQNQWILANTAPEDECDYYSICGAYAVCGINSKNTPSCSCLQGFK 321
Query: 1778 PKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPDFSY-WVSTVGDEP--GC 1608
PK +WN + GC+ P CE D F+ +K PD S+ W + C
Sbjct: 322 PKSGRKWNISRGAYGCVHEIPTNCEKK---DAFVKFPGLKLPDTSWSWYDAKNEMTLEDC 378
Query: 1607 RTVCLNNCSCGAYVYT----ATTGCLAWGNELIDMHELQTGAYTLNLKLPAS--ELRGHH 1446
+ C +NCSC AY T GCL W +L+DM E + + +++ + E +G
Sbjct: 379 KIKCSSNCSCTAYANTDIREGGKGCLLWFGDLVDMREYSSFGQDVYIRMGFAKIEFKGRE 438
Query: 1445 PIWKI--ATIISAIVLFVLAACL---LLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAML 1281
+ + + + A+VL V+ AC ++ G N + +
Sbjct: 439 VVGMVVGSVVAIAVVLVVVFACFRKKIMKRYRGENFRKGIE------------------- 479
Query: 1280 DISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEV 1101
++D++ L ++ I AT +FS N LG GGFGPVY G L G+E+
Sbjct: 480 --------EEDLD------LPIFDRKTISIATDDFSYVNFLGRGGFGPVYKGKLEDGQEI 525
Query: 1100 AVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFL 921
AVKRL NSGQG+EEFKNEV LIAKLQHRNLVRLLGCCI EE +L+YEYMPNKSLD F+
Sbjct: 526 AVKRLSANSGQGVEEFKNEVKLIAKLQHRNLVRLLGCCIQGEECMLIYEYMPNKSLDFFI 585
Query: 920 FNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDFG 741
F+ + LDWKKR +II G+ARG+LYLH+DSRLR++HRDLKA N+LLD DM PKISDFG
Sbjct: 586 FDERRSTELDWKKRMNIINGVARGILYLHQDSRLRIIHRDLKAGNVLLDNDMNPKISDFG 645
Query: 740 MARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSFH 561
+A+ FGGDQ++ +TNRVVGT+GYM PEYA++G FSVKSDV+ FGVL+LEIITGK F
Sbjct: 646 LAKSFGGDQSESSTNRVVGTYGYMPPEYAIDGHFSVKSDVFSFGVLVLEIITGKTNRGFR 705
Query: 560 CHEDSLNIAGYAWRQWNEDNAAELIDPV-IRASCSVRQVLRCIHIALLCVQDHADERPDI 384
+ LN+ G+ W+ W ED E+ + + + + +VLRCIH+ALLCVQ ++RP +
Sbjct: 706 HADHDLNLLGHVWKMWVEDREIEVPEEEWLEETSVIPEVLRCIHVALLCVQQKPEDRPTM 765
Query: 383 PTVILMLSNDSSSLPNPRPP 324
+V+LM +D SSLP+P P
Sbjct: 766 ASVVLMFGSD-SSLPHPTQP 784
>sptr|O64771|O64771 T1F9.2.
Length = 817
Score = 571 bits (1472), Expect = e-161
Identities = 335/812 (41%), Positives = 453/812 (55%), Gaps = 25/812 (3%)
Frame = -2
Query: 2672 ESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVAPATSAS 2493
ES +SS GV+E GFF+ + Q QY+GIW+ I PR VVWVANR P T ++
Sbjct: 29 ESPLSIGKTLSSSNGVYELGFFSFNNSQ--NQYVGIWFKGIIPRVVVWVANREKPVTDSA 86
Query: 2492 PSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQDTGSLEVRSEDG 2313
+LT++ G L + + + ++WS T + G A L D G+L V +
Sbjct: 87 ANLTISSNGSLLLFNENHS-------VVWSIGETFASN---GSRAELTDNGNLVVIDNNS 136
Query: 2312 --VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSPGRYALGLDPGNSG 2139
LW+SF H DT+L + E+ + TSW S TDPSPG + + + P
Sbjct: 137 GRTLWESFEHFGDTMLPFSNLMYNL---ATGEKRVLTSWKSHTDPSPGDFTVQITPQVPS 193
Query: 2138 QAYIWKDGNVTYWRSGQWNGVNFIGIP-WRPLYRSGFTPAIDPVLGNYYTYTATNTSLQR 1962
QA + G+ TYWRSG W F GIP Y S F+ D +TY N L
Sbjct: 194 QACTMR-GSKTYWRSGPWAKTRFTGIPVMDDTYTSPFSLQQDTNGSGSFTYFERNFKLSY 252
Query: 1961 FVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAKCTASQDGKAKCTCLKGF 1782
++ G+ + + + DWEL + P N C+ Y CGP C S KC C KGF
Sbjct: 253 IMITSEGS--LKIFQHNGMDWELNFEAPENSCDIYGFCGPFGICVMSVP--PKCKCFKGF 308
Query: 1781 HPKLQEQWNAGNWSQGCIRSPPLGCETNQSG---DGFLPMGNIKWPDFSYWVSTVGDEPG 1611
PK E+W GNW+ GC+R L C+ N +G +GF + NIK PDF + S V D G
Sbjct: 309 VPKSIEEWKRGNWTDGCVRHTELHCQGNTNGKTVNGFYHVANIKPPDFYEFASFV-DAEG 367
Query: 1610 CRTVCLNNCSCGAYVYTATTGCLAWGNELIDMHELQTGAYTLNLKLPASELRGHHPIWKI 1431
C +CL+NCSC A+ Y GCL W +L+D + G L+++L +SEL G+ K
Sbjct: 368 CYQICLHNCSCLAFAYINGIGCLMWNQDLMDAVQFSAGGEILSIRLASSELGGN----KR 423
Query: 1430 ATIISAIVLFVLAACLLLWWKHGRNIKDAVHGSWRSRHSSTQSQQNSAMLDISQSIRFDD 1251
II A +L HG + + +S ++ + I+ +++
Sbjct: 424 NKIIVASILM-----------HGNTL------------TIIESLVSAKISKIASKEAWNN 460
Query: 1250 DVEDGKSHELKVYSLDRIRTATSNFSDSNKLGEGGFGPVYMGTLPGGEEVAVKRLCRNSG 1071
D+E LK + ++ I+TAT NFS SNKLG+GGFG VY G L G+E+AVKRL +SG
Sbjct: 461 DLEPQDVSGLKFFEMNTIQTATDNFSLSNKLGQGGFGSVYKGKLQDGKEIAVKRLSSSSG 520
Query: 1070 QGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSLDAFLF--------- 918
QG EEF NE++LI+KLQH+NLVR+LGCCI EE++LVYE++ NKSLD FLF
Sbjct: 521 QGKEEFMNEIVLISKLQHKNLVRILGCCIEGEERLLVYEFLLNKSLDTFLFVLIVSIRYY 580
Query: 917 --NPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKISDF 744
+ K+ +DW KRF+IIEGIARGL YLHRDS LRV+HRDLK SNILLD M PKISDF
Sbjct: 581 CLDSRKRLEIDWPKRFNIIEGIARGLHYLHRDSCLRVIHRDLKVSNILLDEKMNPKISDF 640
Query: 743 GMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKRAVSF 564
G+ARM+ G + Q NT RV GT GYM+PEYA G+FS KSD+Y FGV++LEIITG++ F
Sbjct: 641 GLARMYQGTEYQDNTRRVAGTLGYMAPEYAWTGMFSEKSDIYSFGVILLEIITGEKISRF 700
Query: 563 HCHEDSLNIAGY--------AWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQD 408
+ Y AW W E +L+D + SC +V RC+ I LLCVQ
Sbjct: 701 SYGRQGKTLLAYVNLKPKQQAWESWCESGGIDLLDKDVADSCHPLEVERCVQIGLLCVQH 760
Query: 407 HADERPDIPTVILMLSNDSSSLPNPRPPTLML 312
+RP+ ++ ML+ +S L +P+ PT ++
Sbjct: 761 QPADRPNTMELLSMLTT-TSDLTSPKQPTFVV 791
>sptr|Q8LQN4|Q8LQN4 Putative receptor kinase.
Length = 835
Score = 568 bits (1464), Expect = e-160
Identities = 345/854 (40%), Positives = 471/854 (55%), Gaps = 31/854 (3%)
Frame = -2
Query: 2690 DTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVWVANRVA 2511
DT+ G L+ TLVS + F GFF P + Y+G+WY+ +S RTVVWVANR
Sbjct: 28 DTVVPGRPLAANETLVSGGDANFVLGFFTPPGANST--YVGVWYNKVSVRTVVWVANRED 85
Query: 2510 PA---TSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSR-AGPRGGYSAVLQDT 2343
P + +P TL+V+ GT A ++ ++WS ++ A P +A + D+
Sbjct: 86 PLPGDVADNPDATLSVSPT-----GTLAIVAGNSTVVWSVTPAAKLASP----TARIMDS 136
Query: 2342 GSLEVR--SEDGVLWDSFWHPTDTILSGMRITLQ-APGRGPKERMLFTSWASETDPSPGR 2172
G+L + + GV W F +PTDT+L MR+ + GR T+W S +DPSPG
Sbjct: 137 GNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRN----RTLTAWKSPSDPSPGP 192
Query: 2171 YALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFTPA-IDPVLGNYY 1995
+ +D Q +IW +G WRSG W+GV F G+P Y SGFT + I+ Y
Sbjct: 193 VVMAMDTSGDPQVFIW-NGAEKVWRSGPWDGVQFTGVPDTVTY-SGFTFSFINNAKEVTY 250
Query: 1994 TYTATNTSLQRFVVLPNGTDICYMVRKSSQ-----DWELVWYQPSNECEYYATCGPNAKC 1830
++ N S+ + L N T ++++S+ W L WY P ++C+ + CG N C
Sbjct: 251 SFQVHNVSIISRLGL-NSTGSYGLLQRSTWVEAAGTWNLYWYAPKDQCDEVSPCGANGVC 309
Query: 1829 TASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWPD 1650
+ C+CL+GF PK E W + GC+RS PL C+ DGF+ + + K PD
Sbjct: 310 DTNN--LPVCSCLRGFTPKSPEAWALRDGRAGCVRSTPLDCQNGT--DGFVAVEHAKVPD 365
Query: 1649 FSYWVSTVG-DEPGCRTVCLNNCSCGAYV----------YTATTGCLAWGNELIDMHELQ 1503
V +G CR CL NCSC AY + A TGC+ W L D+
Sbjct: 366 TERSVVDLGLSLEQCRKACLMNCSCTAYASANVSGGGRGHGAGTGCVMWTTGLTDLRVYP 425
Query: 1502 TGAYTLNLKLPASEL------RGHHPIWKIATIISAIVLFVLAACLLLWWKHGRNIKDAV 1341
L ++L A++L I I IS++ + A L+W + + +
Sbjct: 426 EFGQDLFVRLAAADLGLTSKSNKARVIIAIVVSISSVTFLSVLAGFLVWTRKKKRARKTG 485
Query: 1340 HGSWRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNK 1161
W ST + S DDD+E L ++ L I AT FS +NK
Sbjct: 486 SSKWSGGSRSTGRRYEG-------SSHHDDDLE------LPIFDLGTIAAATDGFSINNK 532
Query: 1160 LGEGGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIP 981
LGEGGFGPVY G L G+E+AVK L + S QGL+EFKNEV+LIAKLQHRNLVRLLG I
Sbjct: 533 LGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQGLDEFKNEVMLIAKLQHRNLVRLLGFSIS 592
Query: 980 REEKILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRD 801
+E+ILVYEYM NKSLD FLF R+ IIEGI RGLLYLH+DSR R++HRD
Sbjct: 593 GQERILVYEYMANKSLDYFLF-----------ARYRIIEGITRGLLYLHQDSRYRIIHRD 641
Query: 800 LKASNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDV 621
LKASN+LLD +M PKISDFGMARMFG ++ + NT +VVGT+GYMSPEYAM+G+FSVKSDV
Sbjct: 642 LKASNVLLDKEMTPKISDFGMARMFGSEETEINTRKVVGTYGYMSPEYAMDGVFSVKSDV 701
Query: 620 YGFGVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLR 441
+ FGVL+LEII+G+R + + + LN+ G+AW WNE + EL D + S +VL+
Sbjct: 702 FSFGVLLLEIISGRRNRGVYSYSNHLNLLGHAWSLWNEGKSLELADETMNGSFDSDEVLK 761
Query: 440 CIHIALLCVQDHADERPDIPTVILML-SNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXXX 264
CI + LLCVQ++ D+RP + V+LML + D+++LP P+ P R
Sbjct: 762 CIRVGLLCVQENPDDRPLMSQVLLMLATTDATTLPTPKQPGFAARRILMETDTSSSKPDC 821
Query: 263 XXIGTVTMTQLHGR 222
+ T+T L GR
Sbjct: 822 SIFDSATVTILEGR 835
>sptr|Q7XHV3|Q7XHV3 Putative receptor protein kinase.
Length = 860
Score = 566 bits (1458), Expect = e-160
Identities = 337/855 (39%), Positives = 467/855 (54%), Gaps = 26/855 (3%)
Frame = -2
Query: 2708 VAAQKTDTLRQGESLSGAATLVSSPEGVFEAGFFAPDPKQPSRQYLGIWYHSISPRTVVW 2529
+AA +D + SL+ LVS+ GVFE GFF P + ++LGIWY I P TVVW
Sbjct: 23 LAASASDNILANSSLADGQKLVSAG-GVFELGFFTPPGSTTAARFLGIWYRDIDPPTVVW 81
Query: 2528 VANRVAPATSASPSLTLTVTGDLRVLDGTAANGTADAPLLWSSNTTSRAGPRGGYSAVLQ 2349
VANR AP + + SL + V G G G ++WSS S +A L
Sbjct: 82 VANRDAPVSGTAGSLAVVVNGGGGGGGGRLVLGDGSGRVVWSS-APSNVTASDPVAARLL 140
Query: 2348 DTGSLEVRSEDG---VLWDSFWHPTDTILSGMRITLQAPGRGPKERMLFTSWASETDPSP 2178
D+G+ + G V+W SF +P+DT+L GM+ +R L T+W S DPSP
Sbjct: 141 DSGNFVLAGGGGSGDVIWQSFDYPSDTLLPGMKFGWDLTTG--LDRYL-TTWRSAGDPSP 197
Query: 2177 GRYALGLDPGNSGQAYIWKDGNVTYWRSGQWNGVNFIGIPWRPLYRSGFT-PAIDPVLGN 2001
G Y +DP + + +IW +G +R+G W+G+ F G P + F +
Sbjct: 198 GDYTFKIDPRGAPEGFIWYNGTSPVYRNGPWDGLQFSGEPEMEPNNTSFRFEFVANRTDV 257
Query: 2000 YYTYTATNTS----LQRFVVLPNGTDICYMVRKSSQDWELVWYQPSNECEYYATCGPNAK 1833
YYT+ L RFV+ + Y+ + W L W P ++C+ YA CG
Sbjct: 258 YYTFVVDGGGGGGVLSRFVLNQSSAQR-YVWLPQAGGWSLYWSLPRDQCDQYAHCGAYGV 316
Query: 1832 CTASQDGKAKCTCLKGFHPKLQEQWNAGNWSQGCIRSPPLGCETNQSGDGFLPMGNIKWP 1653
C + C C GF P W + S GC R L C +GDGFLP+ +K P
Sbjct: 317 CDVG--AASMCGCPAGFAPASPRNWELRDSSAGCARRTRLNC----TGDGFLPLRGVKLP 370
Query: 1652 DFSY-WVSTVGDEPGCRTVCLNNCSCGAY----VYTATTGCLAWGNELIDMHELQTGAYT 1488
D + V CR CL NCSC AY V +GC+ W + L+D+ + G
Sbjct: 371 DTTNATVDAAIAVDQCRARCLANCSCVAYAASDVRGGGSGCIMWSSPLVDIRKFSYGGED 430
Query: 1487 LNLKLPASELRGH-------HPIWKIATIISAIVLFVLAACLLLWWKHGRN-IKDAVHGS 1332
L ++L AS+L + + + + +S +VL LAA +W K RN + + V
Sbjct: 431 LFMRLAASDLPTNGDDSSRKNTVLAVVLSLSGVVLLALAA-FFVWDKLFRNKVANPVRFQ 489
Query: 1331 WRSRHSSTQSQQNSAMLDISQSIRFDDDVEDGKSHELKVYSLDRIRTATSNFSDSNKLGE 1152
R +S S S L+ Q + +D+ + ++ + I +T NF++ KLGE
Sbjct: 490 SPQRFTSFDS---SIPLNQVQDRKMEDETRHSNELNVTLFDFNTIAFSTDNFANLAKLGE 546
Query: 1151 GGFGPVYMGTLPGGEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREE 972
GGFGPVY G L GG+ VAVKRL + S QGL+EFKNEV+LIA+LQH NLVRLLGCCI EE
Sbjct: 547 GGFGPVYKGELDGGQTVAVKRLSKFSTQGLDEFKNEVMLIARLQHVNLVRLLGCCIHGEE 606
Query: 971 KILVYEYMPNKSLDAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKA 792
++LVYEYM NKSLD F+F+ + L+W KRF+II GIARGLLYLH+DSR +++HRDLKA
Sbjct: 607 RMLVYEYMENKSLDNFIFDKARSAQLNWSKRFNIILGIARGLLYLHQDSRFKIIHRDLKA 666
Query: 791 SNILLDADMKPKISDFGMARMFGGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGF 612
NILLD DM PKISDFG+AR+FG D + +T +VVGT+GYMSPEYAM+G+FSVKSDV+ F
Sbjct: 667 GNILLDGDMNPKISDFGVARIFGDDTDS-HTRKVVGTYGYMSPEYAMDGVFSVKSDVFSF 725
Query: 611 GVLILEIITGKRAVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIR-----ASCSVRQV 447
GVL+LE+++G++ + + ++ +AWR W E NA L+D + S +V
Sbjct: 726 GVLVLELVSGRKNRGMYSSGEQTSLLSHAWRLWREGNALALLDEAVAGGGGGGGYSRSEV 785
Query: 446 LRCIHIALLCVQDHADERPDIPTVILMLSNDSSSLPNPRPPTLMLRGXXXXXXXXXXXXX 267
LRC+ + LLCVQ+ ++RP + V +ML N S+ +P PR P
Sbjct: 786 LRCVQVGLLCVQERPEDRPHMAAVFMMLGNLSAVVPQPRHPGFCSDRGGGGGSTDGEWSS 845
Query: 266 XXXIGTVTMTQLHGR 222
+ VT+T + GR
Sbjct: 846 TCTVNDVTVTIVEGR 860
Database: /db/trembl-ebi/tmp/swall
Posted date: Nov 7, 2003 8:22 PM
Number of letters in database: 415,658,970
Number of sequences in database: 1,302,759
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,104,019,532
Number of Sequences: 1302759
Number of extensions: 50591523
Number of successful extensions: 179165
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 139518
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 162964
length of database: 415,658,970
effective HSP length: 132
effective length of database: 243,694,782
effective search space used: 192275182998
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

