BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
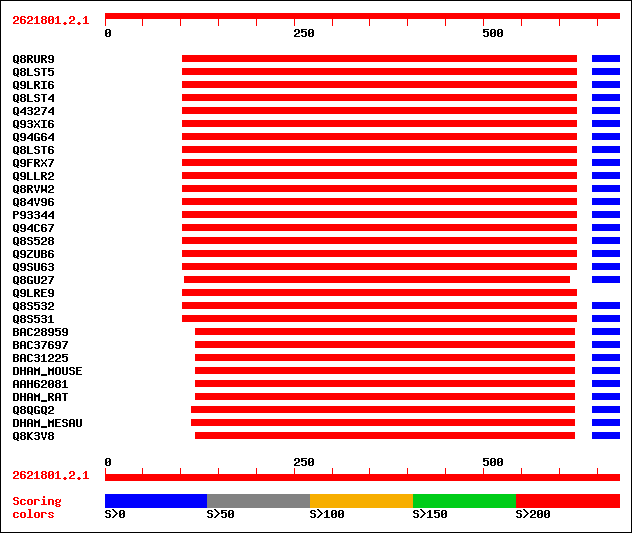
Query= 2621801.2.1
(679 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8RUR9|Q8RUR9 Mitochondrial aldehyde dehydrogenase RF2B. 350 9e-98
sptr|Q8LST5|Q8LST5 Mitochondrial aldehyde dehydrogenase. 336 1e-93
sptr|Q9LRI6|Q9LRI6 Mitochondrial aldehyde dehydrogenase ALDH2a. 309 1e-85
sptr|Q8LST4|Q8LST4 Mitochondrial aldehyde dehydrogenase. 296 1e-81
sptr|Q43274|Q43274 T CYTOPLASM MALE sterility RESTORER factor 2 ... 291 3e-80
sptr|Q93XI6|Q93XI6 Mitochondrial aldehyde dehydrogenase ALDH2. 288 2e-79
sptr|Q94G64|Q94G64 T-cytoplasm male sterility restorer factor 2. 288 3e-79
sptr|Q8LST6|Q8LST6 Mitochondrial aldehyde dehydrogenase. 287 6e-79
sptr|Q9FRX7|Q9FRX7 Aldehyde dehydrogenase ALDH2b. 283 1e-77
sptr|Q9LLR2|Q9LLR2 Aldehyde dehydrogenase. 283 1e-77
sptr|Q8RVW2|Q8RVW2 Aldehyde dehydrogenase (Fragment). 283 3e-77
sptr|Q84V96|Q84V96 Aldehyde dehydrogenase 1 precursor (EC 1.2.1.3). 281 6e-77
sptr|P93344|P93344 Aldehyde dehydrogenase (NAD+) (EC 1.2.1.3). 276 4e-75
sptr|Q94C67|Q94C67 Putative aldehyde dehydrogenase NAD+. 273 9e-75
sptr|Q8S528|Q8S528 Mitochondrial aldehyde dehydrogenase. 273 9e-75
sptr|Q9ZUB6|Q9ZUB6 F5O8.35 protein. 273 9e-75
sptr|Q9SU63|Q9SU63 Aldehyde dehydrogenase (NAD+)-like protein (E... 269 3e-73
sptr|Q8GU27|Q8GU27 Aldehyde dehydrogenase. 231 2e-61
sptr|Q9LRE9|Q9LRE9 Cytosolic aldehyde dehydrogenase. 228 4e-59
sptr|Q8S532|Q8S532 Cytosolic aldehyde dehydrogenase RF2C. 227 3e-58
sptr|Q8S531|Q8S531 Cytosolic aldehyde dehydrogenase RF2C. 222 6e-57
sptrnew|BAC28959|BAC28959 12 days embryo embryonic body between ... 210 2e-54
sptrnew|BAC37697|BAC37697 Adult male spinal cord cDNA, RIKEN ful... 210 2e-54
sptrnew|BAC31225|BAC31225 3 days neonate thymus cDNA, RIKEN full... 210 2e-54
sw|P47738|DHAM_MOUSE Aldehyde dehydrogenase, mitochondrial precu... 210 2e-54
sptrnew|AAH62081|AAH62081 Aldehyde dehydrogenase 2. 210 3e-54
sw|P11884|DHAM_RAT Aldehyde dehydrogenase, mitochondrial precurs... 210 3e-54
sptr|Q8QGQ2|Q8QGQ2 Aldehyde dehydrogenase 2. 210 3e-54
sw|P81178|DHAM_MESAU Aldehyde dehydrogenase, mitochondrial (EC 1... 210 3e-54
sptr|Q8K3V8|Q8K3V8 Mitochondrial aldehyde dehydrogenase (Fragment). 210 3e-54
>sptr|Q8RUR9|Q8RUR9 Mitochondrial aldehyde dehydrogenase RF2B.
Length = 550
Score = 350 bits (897), Expect(2) = 9e-98
Identities = 173/174 (99%), Positives = 174/174 (100%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVG+RGFYIQPTVFA
Sbjct: 377 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGDRGFYIQPTVFA 436
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR
Sbjct: 437 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 496
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL
Sbjct: 497 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 550
Score = 29.6 bits (65), Expect(2) = 9e-98
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 358 FVHERVYDEFVE 369
>sptr|Q8LST5|Q8LST5 Mitochondrial aldehyde dehydrogenase.
Length = 551
Score = 336 bits (862), Expect(2) = 1e-93
Identities = 165/174 (94%), Positives = 171/174 (98%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR+GVEQGPQIDG+QFNKILRYVQSGVDSGATLV GGDRVG RGFYIQPTVFA
Sbjct: 378 KRVVGDPFRNGVEQGPQIDGDQFNKILRYVQSGVDSGATLVTGGDRVGSRGFYIQPTVFA 437
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
DAKD+MKIAREEIFGPVQTILKFSG+EEVIRRANAT YGLAAGVFTRSLDAANTLSRALR
Sbjct: 438 DAKDDMKIAREEIFGPVQTILKFSGMEEVIRRANATHYGLAAGVFTRSLDAANTLSRALR 497
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVWVNCYDVFDATIPFGGYKMSGVGREKG+YALRNYLQTKAVVTPIK+PAWL
Sbjct: 498 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGVYALRNYLQTKAVVTPIKDPAWL 551
Score = 29.6 bits (65), Expect(2) = 1e-93
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 359 FVHERVYDEFVE 370
>sptr|Q9LRI6|Q9LRI6 Mitochondrial aldehyde dehydrogenase ALDH2a.
Length = 553
Score = 309 bits (792), Expect(2) = 1e-85
Identities = 150/174 (86%), Positives = 163/174 (93%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
+RVVGDPFR GVEQGPQIDGEQF KIL+YV+SGVDSGATLVAGGDR G RGFYIQPTVFA
Sbjct: 380 QRVVGDPFRTGVEQGPQIDGEQFKKILQYVKSGVDSGATLVAGGDRAGSRGFYIQPTVFA 439
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +DEMKIA+EEIFGPVQ+ILKFS VEEV+RRANATPYGLAAGVFT+ LDAANTL+RALR
Sbjct: 440 DVEDEMKIAQEEIFGPVQSILKFSTVEEVVRRANATPYGLAAGVFTQRLDAANTLARALR 499
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVWVN YDVFDA +PFGGYKMSGVGREKG+Y+LRNYLQTKAVVTPIK+ AWL
Sbjct: 500 VGTVWVNTYDVFDAAVPFGGYKMSGVGREKGVYSLRNYLQTKAVVTPIKDAAWL 553
Score = 29.6 bits (65), Expect(2) = 1e-85
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 361 FVHERVYDEFVE 372
>sptr|Q8LST4|Q8LST4 Mitochondrial aldehyde dehydrogenase.
Length = 547
Score = 296 bits (757), Expect(2) = 1e-81
Identities = 139/174 (79%), Positives = 159/174 (91%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR GVEQGPQID EQFNKILRY++SGVDSGA LV GGDR+GE+G+YIQPT+F+
Sbjct: 374 KRVVGDPFRKGVEQGPQIDDEQFNKILRYIRSGVDSGANLVTGGDRLGEKGYYIQPTIFS 433
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D MKIA+EEIFGPVQ+ILKF + EVI+RANA+PYGLAAGVFT SLD ANTL+RALR
Sbjct: 434 DVQDGMKIAQEEIFGPVQSILKFKDLNEVIKRANASPYGLAAGVFTNSLDTANTLTRALR 493
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVW+NC+D+FDA IPFGGYKMSG+GREKGI +L+NYLQ KAVVTPIKN AWL
Sbjct: 494 AGTVWINCFDIFDAAIPFGGYKMSGIGREKGIDSLKNYLQVKAVVTPIKNAAWL 547
Score = 29.6 bits (65), Expect(2) = 1e-81
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 355 FVHERVYDEFVE 366
>sptr|Q43274|Q43274 T CYTOPLASM MALE sterility RESTORER factor 2 (EC
1.2.1.3) (Aldehyde dehydrogenase (NAD+)).
Length = 549
Score = 291 bits (745), Expect(2) = 3e-80
Identities = 138/174 (79%), Positives = 157/174 (90%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR GVEQGPQID EQFNKILRY++ GVD GATLV GGDR+G++GFYIQPT+F+
Sbjct: 376 KRVVGDPFRKGVEQGPQIDDEQFNKILRYIRYGVDGGATLVTGGDRLGDKGFYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D MKIA+EEIFGPVQ+ILKF + EVI+RANA+ YGLAAGVFT SLD ANTL+RALR
Sbjct: 436 DVQDGMKIAQEEIFGPVQSILKFKDLNEVIKRANASQYGLAAGVFTNSLDTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVWVNC+DVFDA IPFGGYKMSG+GREKG+ +L+NYLQ KAVVTPIKN AWL
Sbjct: 496 AGTVWVNCFDVFDAAIPFGGYKMSGIGREKGVDSLKNYLQVKAVVTPIKNAAWL 549
Score = 29.6 bits (65), Expect(2) = 3e-80
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 357 FVHERVYDEFVE 368
>sptr|Q93XI6|Q93XI6 Mitochondrial aldehyde dehydrogenase ALDH2.
Length = 549
Score = 288 bits (738), Expect(2) = 2e-79
Identities = 133/174 (76%), Positives = 159/174 (91%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR GVEQGPQID EQF KILRY++SGVDSGATLV GGD++G++G+YIQPT+F+
Sbjct: 376 KRVVGDPFRKGVEQGPQIDDEQFKKILRYIKSGVDSGATLVTGGDKLGDKGYYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D+MKIA+EEIFGPVQ+I KF+ + EVI+RANA+ YGLAAGVFT +LD ANTL+RALR
Sbjct: 436 DVQDDMKIAQEEIFGPVQSIFKFNDLNEVIKRANASQYGLAAGVFTNNLDTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGT+WVNC+D+FDA IPFGGYKMSG+GREKGI +L+NYLQ KAVVT +KNPAWL
Sbjct: 496 AGTIWVNCFDIFDAAIPFGGYKMSGIGREKGIDSLKNYLQVKAVVTALKNPAWL 549
Score = 29.6 bits (65), Expect(2) = 2e-79
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 357 FVHERVYDEFVE 368
>sptr|Q94G64|Q94G64 T-cytoplasm male sterility restorer factor 2.
Length = 549
Score = 288 bits (737), Expect(2) = 3e-79
Identities = 137/174 (78%), Positives = 156/174 (89%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR GVEQGPQID EQF KILRY++ GVD GATLV GGDR+G++GFYIQPT+F+
Sbjct: 376 KRVVGDPFRKGVEQGPQIDDEQFKKILRYIRYGVDGGATLVTGGDRLGDKGFYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D MKIA+EEIFGPVQ+ILKF + EVI+RANA+ YGLAAGVFT SLD ANTL+RALR
Sbjct: 436 DVQDGMKIAQEEIFGPVQSILKFKDLNEVIKRANASQYGLAAGVFTNSLDTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVWVNC+DVFDA IPFGGYKMSG+GREKG+ +L+NYLQ KAVVTPIKN AWL
Sbjct: 496 AGTVWVNCFDVFDAAIPFGGYKMSGMGREKGVDSLKNYLQVKAVVTPIKNAAWL 549
Score = 29.6 bits (65), Expect(2) = 3e-79
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 357 FVHERVYDEFVE 368
>sptr|Q8LST6|Q8LST6 Mitochondrial aldehyde dehydrogenase.
Length = 549
Score = 287 bits (734), Expect(2) = 6e-79
Identities = 133/174 (76%), Positives = 158/174 (90%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR GVEQGPQID EQF KILRY++SGVDSGATLV GGD++G++G+YIQPT+F+
Sbjct: 376 KRVVGDPFRKGVEQGPQIDDEQFKKILRYIKSGVDSGATLVTGGDKLGDKGYYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D MKIA+EEIFGPVQ+I KF+ + EVI+RANA+ YGLAAGVFT +LD ANTL+RALR
Sbjct: 436 DVQDGMKIAQEEIFGPVQSIFKFNDLNEVIKRANASQYGLAAGVFTNNLDTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGT+WVNC+D+FDA IPFGGYKMSG+GREKGI +L+NYLQ KAVVT +KNPAWL
Sbjct: 496 AGTIWVNCFDIFDAAIPFGGYKMSGIGREKGIDSLKNYLQVKAVVTALKNPAWL 549
Score = 29.6 bits (65), Expect(2) = 6e-79
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 357 FVHERVYDEFVE 368
>sptr|Q9FRX7|Q9FRX7 Aldehyde dehydrogenase ALDH2b.
Length = 549
Score = 283 bits (724), Expect(2) = 1e-77
Identities = 132/174 (75%), Positives = 157/174 (90%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPF++GVEQGPQID EQFNKILRY++ GVDSGA LV GGDR+G++G+YIQPT+F+
Sbjct: 376 KRVVGDPFKNGVEQGPQIDDEQFNKILRYIKYGVDSGANLVTGGDRLGDKGYYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D M+IA+EEIFGPVQ+ILKF+ + EVI+RANA+ YGLAAGVFT +L+ ANTL+RALR
Sbjct: 436 DVQDNMRIAQEEIFGPVQSILKFNDLNEVIKRANASQYGLAAGVFTNNLNTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVWVNC+DVFDA IPFGGYK SG+GREKGI +L+NYLQ KAVVTPIKN AWL
Sbjct: 496 VGTVWVNCFDVFDAAIPFGGYKQSGIGREKGIDSLKNYLQVKAVVTPIKNAAWL 549
Score = 29.3 bits (64), Expect(2) = 1e-77
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHER+YDEFVE
Sbjct: 357 FVHERIYDEFVE 368
>sptr|Q9LLR2|Q9LLR2 Aldehyde dehydrogenase.
Length = 549
Score = 283 bits (724), Expect(2) = 1e-77
Identities = 132/174 (75%), Positives = 157/174 (90%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPF++GVEQGPQID EQFNKILRY++ GVDSGA LV GGDR+G++G+YIQPT+F+
Sbjct: 376 KRVVGDPFKNGVEQGPQIDDEQFNKILRYIKYGVDSGANLVTGGDRLGDKGYYIQPTIFS 435
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D +D M+IA+EEIFGPVQ+ILKF+ + EVI+RANA+ YGLAAGVFT +L+ ANTL+RALR
Sbjct: 436 DVQDNMRIAQEEIFGPVQSILKFNDLNEVIKRANASQYGLAAGVFTNNLNTANTLTRALR 495
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVWVNC+DVFDA IPFGGYK SG+GREKGI +L+NYLQ KAVVTPIKN AWL
Sbjct: 496 VGTVWVNCFDVFDAAIPFGGYKQSGIGREKGIDSLKNYLQVKAVVTPIKNAAWL 549
Score = 29.3 bits (64), Expect(2) = 1e-77
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHER+YDEFVE
Sbjct: 357 FVHERIYDEFVE 368
>sptr|Q8RVW2|Q8RVW2 Aldehyde dehydrogenase (Fragment).
Length = 230
Score = 283 bits (723), Expect(2) = 3e-77
Identities = 132/174 (75%), Positives = 156/174 (89%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
+RVVGDPF+ GVEQGPQID EQFNKILRY++ GVD GA+LV GG+R+G +G+YIQPT+F+
Sbjct: 57 RRVVGDPFKKGVEQGPQIDDEQFNKILRYIKLGVDGGASLVTGGERIGSKGYYIQPTIFS 116
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D KD+M IA+EEIFGPVQ+ILKF+ + EVIRRANA+ YGLAAGVFT ++D ANTL+RALR
Sbjct: 117 DVKDDMVIAKEEIFGPVQSILKFNDLNEVIRRANASRYGLAAGVFTSNIDKANTLTRALR 176
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVWVNC+DVFDA IPFGGYKMSG GREKGI +L+ YLQTKAVVTP+KNPAWL
Sbjct: 177 VGTVWVNCFDVFDAAIPFGGYKMSGQGREKGIDSLKGYLQTKAVVTPLKNPAWL 230
Score = 28.1 bits (61), Expect(2) = 3e-77
Identities = 10/12 (83%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHE++YDEFVE
Sbjct: 38 FVHEKIYDEFVE 49
>sptr|Q84V96|Q84V96 Aldehyde dehydrogenase 1 precursor (EC 1.2.1.3).
Length = 542
Score = 281 bits (720), Expect(2) = 6e-77
Identities = 130/174 (74%), Positives = 155/174 (89%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
+RVVGDPF+ GVEQGPQID EQF KILRY++SG++S ATL GGDR+G +GFY+QPTVF+
Sbjct: 369 RRVVGDPFKKGVEQGPQIDTEQFEKILRYIKSGIESNATLECGGDRLGSKGFYVQPTVFS 428
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
+ +D+M IA++EIFGPVQTI KF ++EVIRRAN+T YGLAAGVFT++L ANTL RALR
Sbjct: 429 NVQDDMLIAQDEIFGPVQTIFKFKEIDEVIRRANSTRYGLAAGVFTQNLATANTLMRALR 488
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVW+NC+DVFDA IPFGGYKMSG+GREKGIY+L NYLQ KAVVTP+KNPAWL
Sbjct: 489 AGTVWINCFDVFDAAIPFGGYKMSGIGREKGIYSLHNYLQVKAVVTPLKNPAWL 542
Score = 28.5 bits (62), Expect(2) = 6e-77
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEF+E
Sbjct: 350 FVHERVYDEFLE 361
>sptr|P93344|P93344 Aldehyde dehydrogenase (NAD+) (EC 1.2.1.3).
Length = 542
Score = 276 bits (707), Expect(2) = 4e-75
Identities = 127/174 (72%), Positives = 155/174 (89%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KR VGDPF+ G EQGPQID +QF+KI+ Y++SG+DSGATL GG+R+GERG+YI+PTVF+
Sbjct: 369 KRTVGDPFKSGTEQGPQIDSKQFDKIMNYIRSGIDSGATLETGGERLGERGYYIKPTVFS 428
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
+ KD+M IA++EIFGPVQ+ILKF V++VIRRAN + YGLAAGVFT+++D ANTL+RALR
Sbjct: 429 NVKDDMLIAQDEIFGPVQSILKFKDVDDVIRRANNSRYGLAAGVFTQNIDTANTLTRALR 488
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVWVNC+D FDATIPFGGYKMSG GREKG Y+L+NYLQ KAVVTP+KNPAWL
Sbjct: 489 VGTVWVNCFDTFDATIPFGGYKMSGHGREKGEYSLKNYLQVKAVVTPLKNPAWL 542
Score = 27.3 bits (59), Expect(2) = 4e-75
Identities = 10/12 (83%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHE+VYDEF+E
Sbjct: 350 FVHEKVYDEFLE 361
>sptr|Q94C67|Q94C67 Putative aldehyde dehydrogenase NAD+.
Length = 534
Score = 273 bits (698), Expect(2) = 9e-75
Identities = 126/174 (72%), Positives = 152/174 (87%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KR VGDPF+ G+EQGPQ+D EQFNKIL+Y++ GV++GATL AGGDR+G +G+YIQPTVF+
Sbjct: 361 KRNVGDPFKSGIEQGPQVDSEQFNKILKYIKHGVEAGATLQAGGDRLGSKGYYIQPTVFS 420
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D KD+M IA +EIFGPVQTILKF ++EVI RAN + YGLAAGVFT++LD A+ L RALR
Sbjct: 421 DVKDDMLIATDEIFGPVQTILKFKDLDEVIARANNSRYGLAAGVFTQNLDTAHRLMRALR 480
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVW+NC+DV DA+IPFGGYKMSG+GREKGIY+L NYLQ KAVVT +KNPAWL
Sbjct: 481 VGTVWINCFDVLDASIPFGGYKMSGIGREKGIYSLNNYLQVKAVVTSLKNPAWL 534
Score = 29.6 bits (65), Expect(2) = 9e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 342 FVHERVYDEFVE 353
>sptr|Q8S528|Q8S528 Mitochondrial aldehyde dehydrogenase.
Length = 534
Score = 273 bits (698), Expect(2) = 9e-75
Identities = 126/174 (72%), Positives = 152/174 (87%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KR VGDPF+ G+EQGPQ+D EQFNKIL+Y++ GV++GATL AGGDR+G +G+YIQPTVF+
Sbjct: 361 KRNVGDPFKSGIEQGPQVDSEQFNKILKYIKHGVEAGATLQAGGDRLGSKGYYIQPTVFS 420
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D KD+M IA +EIFGPVQTILKF ++EVI RAN + YGLAAGVFT++LD A+ L RALR
Sbjct: 421 DVKDDMLIATDEIFGPVQTILKFKDLDEVIARANNSRYGLAAGVFTQNLDTAHRLMRALR 480
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVW+NC+DV DA+IPFGGYKMSG+GREKGIY+L NYLQ KAVVT +KNPAWL
Sbjct: 481 VGTVWINCFDVLDASIPFGGYKMSGIGREKGIYSLNNYLQVKAVVTSLKNPAWL 534
Score = 29.6 bits (65), Expect(2) = 9e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 342 FVHERVYDEFVE 353
>sptr|Q9ZUB6|Q9ZUB6 F5O8.35 protein.
Length = 519
Score = 273 bits (698), Expect(2) = 9e-75
Identities = 126/174 (72%), Positives = 152/174 (87%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KR VGDPF+ G+EQGPQ+D EQFNKIL+Y++ GV++GATL AGGDR+G +G+YIQPTVF+
Sbjct: 346 KRNVGDPFKSGIEQGPQVDSEQFNKILKYIKHGVEAGATLQAGGDRLGSKGYYIQPTVFS 405
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D KD+M IA +EIFGPVQTILKF ++EVI RAN + YGLAAGVFT++LD A+ L RALR
Sbjct: 406 DVKDDMLIATDEIFGPVQTILKFKDLDEVIARANNSRYGLAAGVFTQNLDTAHRLMRALR 465
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
GTVW+NC+DV DA+IPFGGYKMSG+GREKGIY+L NYLQ KAVVT +KNPAWL
Sbjct: 466 VGTVWINCFDVLDASIPFGGYKMSGIGREKGIYSLNNYLQVKAVVTSLKNPAWL 519
Score = 29.6 bits (65), Expect(2) = 9e-75
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHERVYDEFVE
Sbjct: 327 FVHERVYDEFVE 338
>sptr|Q9SU63|Q9SU63 Aldehyde dehydrogenase (NAD+)-like protein (EC
1.2.1.3) (Putative aldehyde dehydrogenase (NAD+)
protein) (AT3g48000/T17F15_130) (Aldehyde dehydrogenase
ALDH2a) (NAD+).
Length = 538
Score = 269 bits (688), Expect(2) = 3e-73
Identities = 123/174 (70%), Positives = 154/174 (88%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
KRVVGDPFR G+EQGPQID +QF K+++Y++SG++S ATL GGD++G++G++IQPTVF+
Sbjct: 365 KRVVGDPFRKGIEQGPQIDLKQFEKVMKYIKSGIESNATLECGGDQIGDKGYFIQPTVFS 424
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
+ KD+M IA++EIFGPVQ+ILKFS V+EVI+RAN T YGLAAGVFT++LD AN +SRAL+
Sbjct: 425 NVKDDMLIAQDEIFGPVQSILKFSDVDEVIKRANETKYGLAAGVFTKNLDTANRVSRALK 484
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AGTVWVNC+DVFDA IPFGGYKMSG GREKGIY+L NYLQ KAVVT + PAW+
Sbjct: 485 AGTVWVNCFDVFDAAIPFGGYKMSGNGREKGIYSLNNYLQIKAVVTALNKPAWI 538
Score = 28.5 bits (62), Expect(2) = 3e-73
Identities = 11/12 (91%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FVHE+VYDEFVE
Sbjct: 346 FVHEKVYDEFVE 357
>sptr|Q8GU27|Q8GU27 Aldehyde dehydrogenase.
Length = 523
Score = 231 bits (589), Expect(2) = 2e-61
Identities = 108/170 (63%), Positives = 132/170 (77%)
Frame = -3
Query: 614 VGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFADAK 435
VGDPF +QGPQ+D QF+KIL YV+ G GA L GG RVG +G+Y+ PTVF++
Sbjct: 353 VGDPFSGQYDQGPQVDNAQFSKILSYVEHGKAEGAKLNVGGCRVGNKGYYVAPTVFSNVT 412
Query: 434 DEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRAGT 255
D MKIAREEIFGPVQ+ILK+S +EVIRRANA+ YGLA+GVF++ LD NTL R L AGT
Sbjct: 413 DNMKIAREEIFGPVQSILKYSTFDEVIRRANASDYGLASGVFSKDLDTVNTLVRGLHAGT 472
Query: 254 VWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAW 105
VWVNCY++FD+ +PFGG+K SG+GREKG YAL NY + KAV P+ NPAW
Sbjct: 473 VWVNCYNLFDSAVPFGGFKTSGIGREKGEYALSNYTKVKAVYMPLVNPAW 522
Score = 26.9 bits (58), Expect(2) = 2e-61
Identities = 9/12 (75%), Positives = 12/12 (100%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
+VHE++YDEFVE
Sbjct: 331 YVHEKIYDEFVE 342
>sptr|Q9LRE9|Q9LRE9 Cytosolic aldehyde dehydrogenase.
Length = 502
Score = 228 bits (582), Expect = 4e-59
Identities = 105/174 (60%), Positives = 132/174 (75%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
K VVGDPF V QGPQID EQ+ KIL+Y+ G GATLV GG GE G+YI+PT+F
Sbjct: 329 KSVVGDPFNPRVHQGPQIDKEQYEKILKYIDIGKREGATLVTGGKPCGENGYYIEPTIFT 388
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D K+EM IA+EEIFGPV ++KF VEE I++AN+T YGLAAG+ T+++D ANT+SR++R
Sbjct: 389 DVKEEMSIAQEEIFGPVMALMKFKTVEEAIQKANSTRYGLAAGIVTKNIDVANTVSRSIR 448
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AG +W+NCY FD +PFGGYKMSG G++ G+ AL YL TKAVVTP+ N WL
Sbjct: 449 AGAIWINCYLGFDPDVPFGGYKMSGFGKDMGMDALEKYLHTKAVVTPLYNTPWL 502
>sptr|Q8S532|Q8S532 Cytosolic aldehyde dehydrogenase RF2C.
Length = 502
Score = 227 bits (578), Expect(2) = 3e-58
Identities = 102/174 (58%), Positives = 132/174 (75%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFA 444
K VVGDPF V QGPQ+D +Q+ K+LRY+ G GATLV GG G++G+YI+PT+F
Sbjct: 329 KSVVGDPFNPSVSQGPQVDKDQYEKVLRYIDIGKREGATLVTGGKPCGDKGYYIEPTIFT 388
Query: 443 DAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALR 264
D KD+M IA++EIFGPV ++KF VEEVI++AN T YGLAAG+ T+++D ANT+SR++R
Sbjct: 389 DVKDDMTIAQDEIFGPVMALMKFKTVEEVIQKANNTRYGLAAGIVTKNIDVANTVSRSIR 448
Query: 263 AGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
AG +W+NCY FD PFGGYKMSG G++ G+ AL YLQTK VVTP+ N WL
Sbjct: 449 AGAIWINCYFAFDPDAPFGGYKMSGFGKDMGMDALDKYLQTKTVVTPLYNTPWL 502
Score = 20.8 bits (42), Expect(2) = 3e-58
Identities = 7/12 (58%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
+V E +YDEFV+
Sbjct: 310 YVQEGIYDEFVK 321
>sptr|Q8S531|Q8S531 Cytosolic aldehyde dehydrogenase RF2C.
Length = 503
Score = 222 bits (566), Expect(2) = 6e-57
Identities = 102/175 (58%), Positives = 132/175 (75%), Gaps = 1/175 (0%)
Frame = -3
Query: 623 KRVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGE-RGFYIQPTVF 447
K VVGDPF V QGPQ+D +Q+ K+LRY+ G GATLV GG G+ +G+YI+PT+F
Sbjct: 329 KSVVGDPFNPSVSQGPQVDKDQYEKVLRYIDIGKREGATLVTGGKPCGDNKGYYIEPTIF 388
Query: 446 ADAKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRAL 267
D KD+M IA++EIFGPV ++KF VEEVI++AN T YGLAAG+ T+++D ANT+SR++
Sbjct: 389 TDVKDDMTIAQDEIFGPVMALMKFKTVEEVIQKANNTRYGLAAGIVTKNIDVANTVSRSI 448
Query: 266 RAGTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPIKNPAWL 102
RAG +W+NCY FD PFGGYKMSG G++ G+ AL YLQTK VVTP+ N WL
Sbjct: 449 RAGAIWINCYFAFDPDAPFGGYKMSGFGKDMGMDALDKYLQTKTVVTPLYNTPWL 503
Score = 20.8 bits (42), Expect(2) = 6e-57
Identities = 7/12 (58%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
+V E +YDEFV+
Sbjct: 310 YVQEGIYDEFVK 321
>sptrnew|BAC28959|BAC28959 12 days embryo embryonic body between
diaphragm region and neck cDNA, RIKEN full-length
enriched library, clone:9430093D07 product:aldehyde
dehydrogenase 2, mitochondrial, full insert sequence.
Length = 519
Score = 210 bits (535), Expect(2) = 2e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANDSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 24.6 bits (52), Expect(2) = 2e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQENVYDEFVE 339
>sptrnew|BAC37697|BAC37697 Adult male spinal cord cDNA, RIKEN
full-length enriched library, clone:A330064L07
product:aldehyde dehydrogenase 2, mitochondrial, full
insert sequence.
Length = 519
Score = 210 bits (535), Expect(2) = 2e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANDSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 24.6 bits (52), Expect(2) = 2e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQENVYDEFVE 339
>sptrnew|BAC31225|BAC31225 3 days neonate thymus cDNA, RIKEN
full-length enriched library, clone:A630082J17
product:aldehyde dehydrogenase 2, mitochondrial, full
insert sequence.
Length = 519
Score = 210 bits (535), Expect(2) = 2e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANDSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 24.6 bits (52), Expect(2) = 2e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQENVYDEFVE 339
>sw|P47738|DHAM_MOUSE Aldehyde dehydrogenase, mitochondrial
precursor (EC 1.2.1.3) (ALDH class 2) (AHD-M1) (ALDHI)
(ALDH-E2).
Length = 519
Score = 210 bits (535), Expect(2) = 2e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANDSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 24.6 bits (52), Expect(2) = 2e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQENVYDEFVE 339
>sptrnew|AAH62081|AAH62081 Aldehyde dehydrogenase 2.
Length = 519
Score = 210 bits (535), Expect(2) = 3e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANNSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 23.9 bits (50), Expect(2) = 3e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQEDVYDEFVE 339
>sw|P11884|DHAM_RAT Aldehyde dehydrogenase, mitochondrial precursor
(EC 1.2.1.3) (ALDH class 2) (ALDH1) (ALDH-E2).
Length = 519
Score = 210 bits (535), Expect(2) = 3e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 348 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 407
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 408 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANNSKYGLAAAVFTKDLDKANYLSQALQA 467
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 468 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 514
Score = 23.9 bits (50), Expect(2) = 3e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 328 FVQEDVYDEFVE 339
>sptr|Q8QGQ2|Q8QGQ2 Aldehyde dehydrogenase 2.
Length = 516
Score = 210 bits (535), Expect(2) = 3e-54
Identities = 103/171 (60%), Positives = 122/171 (71%), Gaps = 2/171 (1%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
R+VGDPF EQGPQ+D +QF K+L Y+ SG GA L+ GG ERG++IQPTVF D
Sbjct: 345 RIVGDPFDLNTEQGPQVDEDQFKKVLGYISSGKREGAKLMCGGAPAAERGYFIQPTVFGD 404
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD+MKIAREEIFGPV ILKF +EEVI RAN + YGLAA VFT+++D AN +S LRA
Sbjct: 405 VKDDMKIAREEIFGPVMQILKFKSLEEVIERANDSKYGLAAAVFTQNIDKANYISHGLRA 464
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVV--TPIKN 114
GTVW+NCY+VF PFGGYK SG+GRE G Y L Y + K V P KN
Sbjct: 465 GTVWINCYNVFGVQAPFGGYKASGIGRELGEYGLDIYTEVKTVTIKVPQKN 515
Score = 23.9 bits (50), Expect(2) = 3e-54
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E +YDEFVE
Sbjct: 325 FVQESIYDEFVE 336
>sw|P81178|DHAM_MESAU Aldehyde dehydrogenase, mitochondrial (EC
1.2.1.3) (ALDH class 2) (ALDH1) (ALDH-E2).
Length = 500
Score = 210 bits (535), Expect(2) = 3e-54
Identities = 105/171 (61%), Positives = 123/171 (71%), Gaps = 2/171 (1%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 329 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 388
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 389 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANNSKYGLAAAVFTKDLDKANYLSQALQA 448
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVV--TPIKN 114
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V P KN
Sbjct: 449 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTIKVPQKN 499
Score = 23.9 bits (50), Expect(2) = 3e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 309 FVQEDVYDEFVE 320
>sptr|Q8K3V8|Q8K3V8 Mitochondrial aldehyde dehydrogenase (Fragment).
Length = 488
Score = 210 bits (535), Expect(2) = 3e-54
Identities = 102/167 (61%), Positives = 121/167 (72%)
Frame = -3
Query: 620 RVVGDPFRDGVEQGPQIDGEQFNKILRYVQSGVDSGATLVAGGDRVGERGFYIQPTVFAD 441
RVVG+PF EQGPQ+D QF KIL Y++SG GA L+ GG +RG++IQPTVF D
Sbjct: 317 RVVGNPFDSRTEQGPQVDETQFKKILGYIKSGQQEGAKLLCGGGAAADRGYFIQPTVFGD 376
Query: 440 AKDEMKIAREEIFGPVQTILKFSGVEEVIRRANATPYGLAAGVFTRSLDAANTLSRALRA 261
KD M IA+EEIFGPV ILKF +EEV+ RAN + YGLAA VFT+ LD AN LS+AL+A
Sbjct: 377 VKDGMTIAKEEIFGPVMQILKFKTIEEVVGRANNSKYGLAAAVFTKDLDKANYLSQALQA 436
Query: 260 GTVWVNCYDVFDATIPFGGYKMSGVGREKGIYALRNYLQTKAVVTPI 120
GTVW+NCYDVF A PFGGYKMSG GRE G Y L+ Y + K V +
Sbjct: 437 GTVWINCYDVFGAQSPFGGYKMSGSGRELGEYGLQAYTEVKTVTVKV 483
Score = 23.9 bits (50), Expect(2) = 3e-54
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 679 FVHERVYDEFVE 644
FV E VYDEFVE
Sbjct: 297 FVQEDVYDEFVE 308
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 481,101,531
Number of Sequences: 1395590
Number of extensions: 11130050
Number of successful extensions: 52611
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 43871
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 50537
length of database: 442,889,342
effective HSP length: 119
effective length of database: 276,814,132
effective search space used: 29342297992
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

