BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
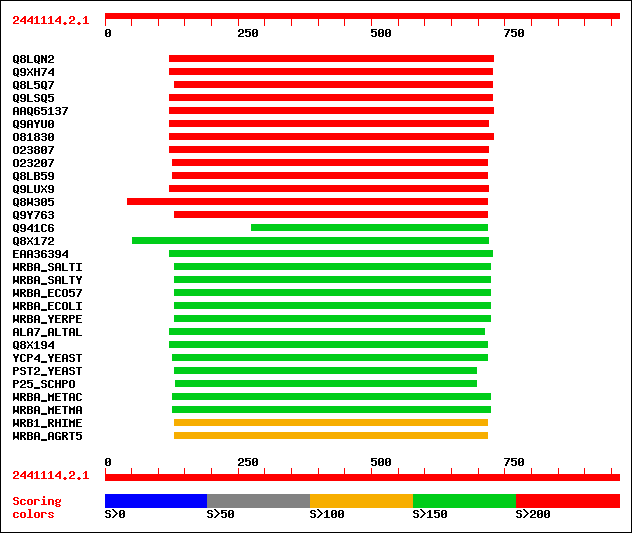
Query= 2441114.2.1
(964 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8LQN2|Q8LQN2 Putative 1,4-benzoquinone reductase. 363 2e-99
sptr|Q9XH74|Q9XH74 Hypothetical 21.7 kDa protein. 315 6e-85
sptr|Q8L5Q7|Q8L5Q7 Putative quinone oxidoreductase (EC 1.6.5.5). 315 8e-85
sptr|Q9LSQ5|Q9LSQ5 1,4-benzoquinone reductase-like, Trp represso... 306 3e-82
sptrnew|AAQ65137|AAQ65137 At4g27270. 305 6e-82
sptr|Q9AYU0|Q9AYU0 Quinone-oxidoreductase QR2. 304 1e-81
sptr|O81830|O81830 Hypothetical 22.3 kDa protein. 282 4e-75
sptr|O23807|O23807 LEDI-3 protein. 268 1e-70
sptr|O23207|O23207 Minor allergen. 247 2e-64
sptr|Q8LB59|Q8LB59 Minor allergen. 247 2e-64
sptr|Q9LUX9|Q9LUX9 1,4-benzoquinone reductase-like (Putative lig... 247 2e-64
sptr|Q8W305|Q8W305 Putative reductase. 213 4e-54
sptr|Q9Y763|Q9Y763 1,4-benzoquinone reductase. 204 1e-51
sptr|Q941C6|Q941C6 C7A10_610/C7A10_610 (At4g36690/C7A10_610). 188 8e-47
sptr|Q8X172|Q8X172 NADH:quinone oxidoreductase. 181 2e-44
sptrnew|EAA36394|EAA36394 Hypothetical protein. 180 2e-44
sw|Q8Z7N9|WRBA_SALTI Flavoprotein wrbA (Trp repressor binding pr... 179 5e-44
sw|Q8ZQ40|WRBA_SALTY Flavoprotein wrbA (Trp repressor binding pr... 179 7e-44
sw|Q8X4B4|WRBA_ECO57 Flavoprotein wrbA (Trp repressor binding pr... 175 1e-42
sw|P30849|WRBA_ECOLI Flavoprotein wrbA (Trp repressor binding pr... 174 2e-42
sw|Q8ZF61|WRBA_YERPE Flavoprotein wrbA (Trp repressor binding pr... 173 3e-42
sw|P42058|ALA7_ALTAL Minor allergen Alt a 7 (Alt a VII). 173 4e-42
sptr|Q8X194|Q8X194 Y20 protein. 172 8e-42
sw|P25349|YCP4_YEAST Hypothetical 26.4 kDa protein in CDC10-CIT2... 171 2e-41
sw|Q12335|PST2_YEAST Protoplast secreted protein 2 precursor. 164 1e-39
sw|P30821|P25_SCHPO P25 protein (Brefeldin A resistance protein). 163 4e-39
sw|P58796|WRBA_METAC Flavoprotein wrbA. 158 1e-37
sw|Q8PUV4|WRBA_METMA Flavoprotein wrbA. 151 1e-35
sw|Q92PU3|WRB1_RHIME Flavoprotein wrbA1. 146 4e-34
sw|P58795|WRBA_AGRT5 Flavoprotein wrbA. 144 1e-33
>sptr|Q8LQN2|Q8LQN2 Putative 1,4-benzoquinone reductase.
Length = 203
Score = 363 bits (931), Expect = 2e-99
Identities = 175/203 (86%), Positives = 183/203 (90%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MAVK+YVVYYSMYGHV KLA+EIKKGA S+EGVEAKIWQVPE L EEVLGKMGAPPKPDV
Sbjct: 1 MAVKVYVVYYSMYGHVAKLAEEIKKGASSIEGVEAKIWQVPETLHEEVLGKMGAPPKPDV 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
P ITPQEL EADGILFGFPTRFGMMAAQMKAFFDATGGLW EQSLAGKPAG+FFS
Sbjct: 61 PTITPQELTEADGILFGFPTRFGMMAAQMKAFFDATGGLWSEQSLAGKPAGIFFSTGTQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEV 660
PLTAITQLTHHGMVFVPVGYTFGAK+F M +VQGGSPYGAGTFAADGSRWP+E+
Sbjct: 121 GGQETTPLTAITQLTHHGMVFVPVGYTFGAKMFNMGEVQGGSPYGAGTFAADGSRWPTEM 180
Query: 661 ELEHAFHQGKYFAGIAKKLKGSA 729
ELEHAFHQGKYFAGIAKKLKGSA
Sbjct: 181 ELEHAFHQGKYFAGIAKKLKGSA 203
>sptr|Q9XH74|Q9XH74 Hypothetical 21.7 kDa protein.
Length = 204
Score = 315 bits (807), Expect = 6e-85
Identities = 147/202 (72%), Positives = 173/202 (85%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MA K+Y+VYYSMYGHV KLA+EI KGA SVEGVEAK+WQV E L ++VLGKMGAPPK +V
Sbjct: 1 MATKVYIVYYSMYGHVEKLAEEILKGAASVEGVEAKLWQVAETLQDDVLGKMGAPPKSEV 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
P+I+P +L+EADG+LFGFPTRFGMMAAQ KAFFD+TGGLWR Q+LAGKPAG+F+S
Sbjct: 61 PIISPNDLSEADGLLFGFPTRFGMMAAQFKAFFDSTGGLWRTQALAGKPAGIFYSTGSQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEV 660
LTAITQL HHGMVFVP+GY+FGA +F M+Q++GGSPYGAGT+A DG+R PSE+
Sbjct: 121 GGQETTALTAITQLVHHGMVFVPIGYSFGAGMFEMEQIKGGSPYGAGTYAGDGTRQPSEL 180
Query: 661 ELEHAFHQGKYFAGIAKKLKGS 726
EL+ AFHQGKYFAGIAKKLKGS
Sbjct: 181 ELQQAFHQGKYFAGIAKKLKGS 202
>sptr|Q8L5Q7|Q8L5Q7 Putative quinone oxidoreductase (EC 1.6.5.5).
Length = 204
Score = 315 bits (806), Expect = 8e-85
Identities = 147/199 (73%), Positives = 170/199 (85%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
K+Y+VYYS YGHV KLA+EI+KGA SVEGVEAK+WQVPE L E+VLGKMGAPPK DVP+I
Sbjct: 5 KVYIVYYSTYGHVHKLAEEIQKGAASVEGVEAKLWQVPETLPEDVLGKMGAPPKSDVPII 64
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TP EL EADG+LFGFPTRFGMMAAQ KAF DATGGLWR Q+LAGKPAG+F+S
Sbjct: 65 TPNELPEADGLLFGFPTRFGMMAAQFKAFMDATGGLWRTQALAGKPAGIFYSTGSQGGGQ 124
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEVELE 669
PLT+ITQL HHG++FVP+GYTFGA +F +++V+GGSPYGAGT+A DGSR P+E+EL
Sbjct: 125 ETTPLTSITQLVHHGLIFVPIGYTFGAGMFEIEKVKGGSPYGAGTYAGDGSRQPTELELA 184
Query: 670 HAFHQGKYFAGIAKKLKGS 726
AFHQGKYFAGIAKKLKGS
Sbjct: 185 QAFHQGKYFAGIAKKLKGS 203
>sptr|Q9LSQ5|Q9LSQ5 1,4-benzoquinone reductase-like, Trp repressor
binding protein-like (1,4-benzoquinone reductase-like
protein).
Length = 204
Score = 306 bits (784), Expect = 3e-82
Identities = 145/202 (71%), Positives = 165/202 (81%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MA K+Y+VYYSMYGHV KLA+EI+KGA SVEGVEAK+WQVPE L EE L KM APPK +
Sbjct: 1 MATKVYIVYYSMYGHVEKLAEEIRKGAASVEGVEAKLWQVPETLHEEALSKMSAPPKSES 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
P+ITP ELAEADG +FGFPTRFGMMAAQ KAF DATGGLWR Q+LAGKPAG+F+S
Sbjct: 61 PIITPNELAEADGFVFGFPTRFGMMAAQFKAFLDATGGLWRAQALAGKPAGIFYSTGSQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEV 660
LTAITQL HHGM+FVP+GYTFGA +F M+ V+GGSPYGAGTFA DGSR P+E+
Sbjct: 121 GGQETTALTAITQLVHHGMLFVPIGYTFGAGMFEMENVKGGSPYGAGTFAGDGSRQPTEL 180
Query: 661 ELEHAFHQGKYFAGIAKKLKGS 726
EL+ AFHQG+Y A I KKLKGS
Sbjct: 181 ELQQAFHQGQYIASITKKLKGS 202
>sptrnew|AAQ65137|AAQ65137 At4g27270.
Length = 205
Score = 305 bits (781), Expect = 6e-82
Identities = 145/203 (71%), Positives = 163/203 (80%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MA K+Y+VYYSMYGHV KLA EI+KGA SV+GVEA +WQVPE L E+VL KM APPK D
Sbjct: 1 MATKVYIVYYSMYGHVEKLAQEIRKGAASVDGVEAILWQVPETLQEDVLSKMSAPPKSDA 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
P+ITP ELAEADG +FGFPTRFGMMAAQ KAF DATGGLWR Q LAGKPAG+F+S
Sbjct: 61 PIITPNELAEADGFIFGFPTRFGMMAAQFKAFLDATGGLWRTQQLAGKPAGIFYSTGSQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEV 660
LTAITQL HHGM+FVP+GYTFGA +F M+ V+GGSPYGAGTFA DGSR P+E+
Sbjct: 121 GGQETTALTAITQLVHHGMIFVPIGYTFGAGMFEMENVKGGSPYGAGTFAGDGSRQPTEL 180
Query: 661 ELEHAFHQGKYFAGIAKKLKGSA 729
EL AFHQGKY A I+KKLKG A
Sbjct: 181 ELGQAFHQGKYIAAISKKLKGPA 203
>sptr|Q9AYU0|Q9AYU0 Quinone-oxidoreductase QR2.
Length = 205
Score = 304 bits (779), Expect = 1e-81
Identities = 145/201 (72%), Positives = 167/201 (83%), Gaps = 1/201 (0%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MA K+Y+VYYS YGHV +LA EIKKGA SV VE K+WQVPEILS+EVLGKM APPK DV
Sbjct: 1 MATKVYIVYYSTYGHVERLAQEIKKGAESVGNVEVKLWQVPEILSDEVLGKMWAPPKSDV 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
PVITP EL EADGI+FGFPTRFGMMAAQ KAFFD+TGGLW+ Q+LAGKPAG+FFS
Sbjct: 61 PVITPDELVEADGIIFGFPTRFGMMAAQFKAFFDSTGGLWKTQALAGKPAGIFFSTGTQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSE 657
LTAITQLTHHGM++VP+GYTFGA +F M++++GGSPYGAGTFA ADGSR PS+
Sbjct: 121 GGQETTALTAITQLTHHGMIYVPIGYTFGADMFNMEKIKGGSPYGAGTFAGADGSRQPSD 180
Query: 658 VELEHAFHQGKYFAGIAKKLK 720
+EL+ AFHQG Y AGI KK+K
Sbjct: 181 IELKQAFHQGMYIAGITKKIK 201
>sptr|O81830|O81830 Hypothetical 22.3 kDa protein.
Length = 211
Score = 282 bits (722), Expect = 4e-75
Identities = 139/209 (66%), Positives = 160/209 (76%), Gaps = 6/209 (2%)
Frame = +1
Query: 121 MAVKIYVVY----YSMYGHVG--KLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGA 282
MA K+Y+VY + + G + KLA EI+KGA SV+GVEA +WQVPE L E+VL KM A
Sbjct: 1 MATKVYIVYVKIIFCLSGAIDNLKLAQEIRKGAASVDGVEAILWQVPETLQEDVLSKMSA 60
Query: 283 PPKPDVPVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFF 462
PPK D P+ITP ELAEADG +FGFPTRFGMMAAQ KAF DATGGLWR Q LAGKPAG+F+
Sbjct: 61 PPKSDAPIITPNELAEADGFIFGFPTRFGMMAAQFKAFLDATGGLWRTQQLAGKPAGIFY 120
Query: 463 SXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGS 642
S LTAITQL HHGM+FVP+GYTFGA +F M+ V+GGSPYGAGTFA DGS
Sbjct: 121 STGSQGGGQETTALTAITQLVHHGMIFVPIGYTFGAGMFEMENVKGGSPYGAGTFAGDGS 180
Query: 643 RWPSEVELEHAFHQGKYFAGIAKKLKGSA 729
R P+E+EL AFHQGKY A I+KKLKG A
Sbjct: 181 RQPTELELGQAFHQGKYIAAISKKLKGPA 209
>sptr|O23807|O23807 LEDI-3 protein.
Length = 201
Score = 268 bits (684), Expect = 1e-70
Identities = 134/201 (66%), Positives = 155/201 (77%), Gaps = 1/201 (0%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MAVK+Y+VYYS YGHV KLA+E+KKGA SV+GVE K+WQV E L E+VLGKM APPK D
Sbjct: 1 MAVKLYIVYYSTYGHVLKLAEEMKKGAESVDGVEVKLWQVAETLPEDVLGKMYAPPKGDA 60
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
P+ITP ELAEADG LFGFPTRFGMM AQ KAFFD+TGGLW+ L GKPAG F S
Sbjct: 61 PIITPDELAEADGYLFGFPTRFGMMPAQFKAFFDSTGGLWQSGGLVGKPAGFFISSGSQG 120
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAA-DGSRWPSE 657
LTAITQL HHGM++VP G TFGA L ++++GG+ YGAGTFA DG+R P+E
Sbjct: 121 GGQETTALTAITQLVHHGMIYVPTGGTFGAGLH-KNEIRGGTLYGAGTFAGPDGARQPTE 179
Query: 658 VELEHAFHQGKYFAGIAKKLK 720
+ELE AFHQGKY AGI K+LK
Sbjct: 180 LELELAFHQGKYTAGIVKRLK 200
>sptr|O23207|O23207 Minor allergen.
Length = 273
Score = 247 bits (631), Expect = 2e-64
Identities = 124/198 (62%), Positives = 146/198 (73%), Gaps = 1/198 (0%)
Frame = +1
Query: 127 VKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKP-DVP 303
+KI+VV+YSMYGHV LA +KKG SVEGVEA +++VPE LS+EV+ +M AP K ++P
Sbjct: 73 LKIFVVFYSMYGHVESLAKRMKKGVDSVEGVEATLYRVPETLSQEVVEQMKAPVKDLEIP 132
Query: 304 VITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXX 483
IT EL ADG LFGFPTR+G MAAQMKAFFD+TG LW+EQSLAGKPAG F S
Sbjct: 133 EITAAELTAADGFLFGFPTRYGCMAAQMKAFFDSTGSLWKEQSLAGKPAGFFVSTGTQGG 192
Query: 484 XXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEVE 663
TAITQL HHGM+FVP+GYTFGA +F MD ++GGSPYGAG FA DGSR +E E
Sbjct: 193 GQETTAWTAITQLVHHGMLFVPIGYTFGAGMFKMDSIRGGSPYGAGVFAGDGSREATETE 252
Query: 664 LEHAFHQGKYFAGIAKKL 717
L A HQG Y A I K+L
Sbjct: 253 LALAEHQGNYMAAIVKRL 270
>sptr|Q8LB59|Q8LB59 Minor allergen.
Length = 273
Score = 247 bits (631), Expect = 2e-64
Identities = 124/198 (62%), Positives = 146/198 (73%), Gaps = 1/198 (0%)
Frame = +1
Query: 127 VKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKP-DVP 303
+KI+VV+YSMYGHV LA +KKG SVEGVEA +++VPE LS+EV+ +M AP K ++P
Sbjct: 73 LKIFVVFYSMYGHVESLAKRMKKGVDSVEGVEATLYRVPETLSQEVVEQMKAPVKDLEIP 132
Query: 304 VITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXX 483
IT EL ADG LFGFPTR+G MAAQMKAFFD+TG LW+EQSLAGKPAG F S
Sbjct: 133 EITAAELTAADGFLFGFPTRYGCMAAQMKAFFDSTGSLWKEQSLAGKPAGFFVSTGTQGG 192
Query: 484 XXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEVE 663
TAITQL HHGM+FVP+GYTFGA +F MD ++GGSPYGAG FA DGSR +E E
Sbjct: 193 GQETTAWTAITQLVHHGMLFVPIGYTFGAGMFKMDSIRGGSPYGAGVFAGDGSREATETE 252
Query: 664 LEHAFHQGKYFAGIAKKL 717
L A HQG Y A I K+L
Sbjct: 253 LALAEHQGNYMAAIVKRL 270
>sptr|Q9LUX9|Q9LUX9 1,4-benzoquinone reductase-like (Putative light
harvesting pigment protein).
Length = 207
Score = 247 bits (630), Expect = 2e-64
Identities = 120/202 (59%), Positives = 155/202 (76%), Gaps = 2/202 (0%)
Frame = +1
Query: 121 MAV-KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPD 297
MAV KIY+VYYS++GHV +A E+ +G SV VEA +WQVPE L E++L K+ A P+PD
Sbjct: 1 MAVTKIYIVYYSLHGHVETMAREVLRGVNSVPDVEATLWQVPETLPEKILEKVKAVPRPD 60
Query: 298 -VPVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXX 474
VP I P++LAEADG +FGFP+RFG+MA+Q+ FFD T LW Q+LAGKPAG+F+S
Sbjct: 61 DVPDIRPEQLAEADGFMFGFPSRFGVMASQVMTFFDNTNDLWTTQALAGKPAGIFWSTGF 120
Query: 475 XXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFAADGSRWPS 654
LTA+T+L HHGM+FVPVGYTFG ++ M +V+GGSPYG+GT+AADGSR P+
Sbjct: 121 HGGGQELTALTAVTKLAHHGMIFVPVGYTFGKSMYEMGEVKGGSPYGSGTYAADGSREPT 180
Query: 655 EVELEHAFHQGKYFAGIAKKLK 720
E+E++ A + GKYFAGIAKKLK
Sbjct: 181 ELEIQQANYHGKYFAGIAKKLK 202
>sptr|Q8W305|Q8W305 Putative reductase.
Length = 252
Score = 213 bits (541), Expect = 4e-54
Identities = 113/232 (48%), Positives = 143/232 (61%), Gaps = 7/232 (3%)
Frame = +1
Query: 43 SSAPARPQEFPHLPLWSGHRSSETSPMAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVE 222
++A A P H + + +P V+IYVV+YSMYGHV LA + +G SV G
Sbjct: 16 AAAAAAPASHSHQRERTSIPAPAAAPRPVRIYVVFYSMYGHVRLLARAVARGVGSVPGAR 75
Query: 223 AKIWQVPEILSEEVLGKM-------GAPPKPDVPVITPQELAEADGILFGFPTRFGMMAA 381
A +++VPE L VL +M G + +PV+ P L +ADG LFGFP RFG M A
Sbjct: 76 AILFRVPETLPPAVLARMEADDGGGGGDGEDVIPVVDPDGLPDADGFLFGFPARFGAMPA 135
Query: 382 QMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYT 561
QM+AFFD+T L R Q LAGKPAG+F S TAITQL HHGM+FVP+GYT
Sbjct: 136 QMQAFFDSTVPLCRHQRLAGKPAGLFVSTGTQAGGQETTAWTAITQLAHHGMLFVPIGYT 195
Query: 562 FGAKLFGMDQVQGGSPYGAGTFAADGSRWPSEVELEHAFHQGKYFAGIAKKL 717
FG + M +++GGSPYGAG F+ DGSR PSE+EL A H GKY A + KK+
Sbjct: 196 FGEGMLEMGELRGGSPYGAGVFSGDGSRPPSELELALAEHHGKYMATLVKKM 247
>sptr|Q9Y763|Q9Y763 1,4-benzoquinone reductase.
Length = 201
Score = 204 bits (520), Expect = 1e-51
Identities = 108/198 (54%), Positives = 136/198 (68%), Gaps = 2/198 (1%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
K+ ++ YSMYGH+ KLA E +K + G A I+Q+PE L EEVL KM APPKP+ PVI
Sbjct: 3 KVAIIIYSMYGHIAKLA-EAEKAGIEEAGGSATIYQIPETLPEEVLAKMHAPPKPEYPVI 61
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TP++L E D +FG PTR+G Q KAF+DATGGLW + +LAGK A +F S
Sbjct: 62 TPEKLPEFDAFVFGIPTRYGNFPGQWKAFWDATGGLWAQGALAGKYASVFVSTGTPGGGQ 121
Query: 490 XXXPLTAITQLTHHGMVFVPVGY-TFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVE 663
L +I+ LTHHG+VFVP+GY T A+L + +V+GGSP+GAGTFA ADGSR PS +E
Sbjct: 122 ESTVLNSISTLTHHGIVFVPLGYSTTFAQLANLSEVRGGSPWGAGTFAGADGSRSPSALE 181
Query: 664 LEHAFHQGKYFAGIAKKL 717
LE A QGKYF I KK+
Sbjct: 182 LELATAQGKYFWNIIKKV 199
>sptr|Q941C6|Q941C6 C7A10_610/C7A10_610 (At4g36690/C7A10_610).
Length = 152
Score = 188 bits (478), Expect = 8e-47
Identities = 94/149 (63%), Positives = 106/149 (71%), Gaps = 1/149 (0%)
Frame = +1
Query: 274 MGAPPKP-DVPVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPA 450
M AP K ++P IT EL ADG LFGFPTR+G MAAQMKAFFD+TG LW+EQSLAGKPA
Sbjct: 1 MKAPVKDLEIPEITAAELTAADGFLFGFPTRYGCMAAQMKAFFDSTGSLWKEQSLAGKPA 60
Query: 451 GMFFSXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA 630
G F S TAITQL HHGM+FVP+GYTFGA +F MD ++GGSPYGAG FA
Sbjct: 61 GFFVSTGTQGGGQETTAWTAITQLVHHGMLFVPIGYTFGAGMFKMDSIRGGSPYGAGVFA 120
Query: 631 ADGSRWPSEVELEHAFHQGKYFAGIAKKL 717
DGSR +E EL A HQG Y A I K+L
Sbjct: 121 GDGSREATETELALAEHQGNYMAAIVKRL 149
>sptr|Q8X172|Q8X172 NADH:quinone oxidoreductase.
Length = 257
Score = 181 bits (458), Expect = 2e-44
Identities = 100/230 (43%), Positives = 141/230 (61%), Gaps = 7/230 (3%)
Frame = +1
Query: 52 PARPQEFPHLPLWSGHR-----SSETSPMAVKIYVVYYSMYGHVGKLADEIKKGALSVEG 216
PA P P +G + +++T+ + ++ +V Y+MYGHV KLA+ IK G + G
Sbjct: 28 PAESTNTPAPPTSTGTKPTTTQTTDTTMSSPRLAIVIYTMYGHVAKLAEAIKSG-IEGAG 86
Query: 217 VEAKIWQVPEILSEEVLGKMGAPPKPDVPVITPQELAEADGILFGFPTRFGMMAAQMKAF 396
A I+QV E LS E+L + APPKPD PV+ P +L DG LFG PTR+G Q KAF
Sbjct: 87 GNASIFQVAETLSPEILNLVKAPPKPDYPVMDPLDLKNYDGFLFGIPTRYGNFPVQWKAF 146
Query: 397 FDATGGLWREQSLAGKPAGMFFSXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYTFG-AK 573
+D+TG LW +L GK AG+F S + A++ L HHG+++VP+GY + A+
Sbjct: 147 WDSTGPLWASTALCGKYAGLFVSTGSPGGGQESTLMAAMSTLVHHGVIYVPLGYKYTFAQ 206
Query: 574 LFGMDQVQGGSPYGAGTFA-ADGSRWPSEVELEHAFHQGKYFAGIAKKLK 720
L + +V+GGSP+GAGTFA +DGSR P+ +ELE A QGK F ++K
Sbjct: 207 LANLTEVRGGSPWGAGTFANSDGSRQPTPLELEIANLQGKSFYEYVARVK 256
>sptrnew|EAA36394|EAA36394 Hypothetical protein.
Length = 244
Score = 180 bits (457), Expect = 2e-44
Identities = 104/209 (49%), Positives = 131/209 (62%), Gaps = 7/209 (3%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKP-D 297
MA KI +VYYSMYGH+ +LA+ K G + G A ++QVPE LS+EVL KM APPKP D
Sbjct: 1 MAPKIAIVYYSMYGHIRQLAEAAKAG-IEKAGGTADLYQVPETLSDEVLAKMYAPPKPTD 59
Query: 298 VPVIT-PQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXX 474
+PVI P L E DG LFG PTR+G AQ +AF+D TGGLW L GK AG+F S
Sbjct: 60 IPVIEDPAILKEYDGFLFGIPTRYGNFPAQWRAFWDKTGGLWATGGLYGKAAGLFISTAG 119
Query: 475 XXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFG----MDQVQGGSPYGAGTFA-ADG 639
+ A++ L HHG+++VP+GY AK+FG + V GGSP+G+GT + DG
Sbjct: 120 LGGGQESTAIAAMSTLAHHGIIYVPLGY---AKVFGELSDLSAVHGGSPWGSGTLSGGDG 176
Query: 640 SRWPSEVELEHAFHQGKYFAGIAKKLKGS 726
SR PSE EL+ A QG+ F KL S
Sbjct: 177 SRQPSESELKVAGIQGEEFYNTLSKLTAS 205
>sw|Q8Z7N9|WRBA_SALTI Flavoprotein wrbA (Trp repressor binding
protein).
Length = 197
Score = 179 bits (454), Expect = 5e-44
Identities = 98/199 (49%), Positives = 125/199 (62%), Gaps = 1/199 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
KI V+YYSMYGH+ +A + +GA V+G E I +VPE + E+ K G + + PV
Sbjct: 2 KILVLYYSMYGHIETMAHAVAEGAKKVDGAEVIIKRVPETMPPEIFAKAGGKTQ-NAPVA 60
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TPQELA+ D I+FG PTRFG M+ QM+ F D TGGLW +L GK G+F S
Sbjct: 61 TPQELADYDAIIFGTPTRFGNMSGQMRTFLDQTGGLWASGALYGKLGGVFSSTGTGGGQE 120
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVEL 666
T T L HHGMV VP+GY+ +LF + QV+GG+PYGA T A DGSR PS+ EL
Sbjct: 121 QTITST-WTTLAHHGMVIVPIGYS-AQELFDVSQVRGGTPYGATTIAGGDGSRQPSQEEL 178
Query: 667 EHAFHQGKYFAGIAKKLKG 723
A +QG+Y AG+A KL G
Sbjct: 179 SIARYQGEYVAGLAVKLNG 197
>sw|Q8ZQ40|WRBA_SALTY Flavoprotein wrbA (Trp repressor binding
protein).
Length = 197
Score = 179 bits (453), Expect = 7e-44
Identities = 98/199 (49%), Positives = 123/199 (61%), Gaps = 1/199 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
KI V+YYSMYGH+ +A + +GA V+G E I +VPE + E+ K G + + PV
Sbjct: 2 KILVLYYSMYGHIETMAHAVAEGAKKVDGAEVIIKRVPETMPPEIFAKAGGKTQ-NAPVA 60
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TPQELA+ D I+FG PTRFG M+ QM+ F D TGGLW SL GK G FS
Sbjct: 61 TPQELADYDAIIFGTPTRFGNMSGQMRTFLDQTGGLWASGSLYGK-LGSVFSSTGTGGGQ 119
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVEL 666
+ T L HHGMV VP+GY +LF + QV+GG+PYGA T A DGSR PS+ EL
Sbjct: 120 EQTITSTWTTLAHHGMVIVPIGYA-AQELFDVSQVRGGTPYGATTIAGGDGSRQPSQEEL 178
Query: 667 EHAFHQGKYFAGIAKKLKG 723
A +QG+Y AG+A KL G
Sbjct: 179 SIARYQGEYVAGLAVKLNG 197
>sw|Q8X4B4|WRBA_ECO57 Flavoprotein wrbA (Trp repressor binding
protein).
Length = 197
Score = 175 bits (443), Expect = 1e-42
Identities = 96/199 (48%), Positives = 123/199 (61%), Gaps = 1/199 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
K+ V+YYSMYGH+ +A + +GA V+G E + +VPE +S ++ K G + PV
Sbjct: 2 KVLVLYYSMYGHIETMARAVAEGASKVDGAEVVVKRVPETMSPQLFEKAGGKTQT-APVA 60
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TPQELA D I+FG PTRFG M+ QM+ F D TGGLW +L GK A +F S
Sbjct: 61 TPQELANYDAIIFGTPTRFGNMSGQMRTFLDQTGGLWASGALYGKLASVFSSTGTGGGQE 120
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVEL 666
T T L HHGMV VP+GY +LF + QV+GG+PYGA T A DGSR PS+ EL
Sbjct: 121 QTITST-WTTLAHHGMVIVPIGYA-AQELFDVSQVRGGTPYGATTIAGGDGSRQPSQEEL 178
Query: 667 EHAFHQGKYFAGIAKKLKG 723
A +QG+Y AG+A KL G
Sbjct: 179 SIARYQGEYVAGLAVKLNG 197
>sw|P30849|WRBA_ECOLI Flavoprotein wrbA (Trp repressor binding
protein).
Length = 197
Score = 174 bits (440), Expect = 2e-42
Identities = 95/199 (47%), Positives = 123/199 (61%), Gaps = 1/199 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
K+ V+YYSMYGH+ +A + +GA V+G E + +VPE + ++ K G + PV
Sbjct: 2 KVLVLYYSMYGHIETMARAVAEGASKVDGAEVVVKRVPETMPPQLFEKAGGKTQT-APVA 60
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TPQELA+ D I+FG PTRFG M+ QM+ F D TGGLW +L GK A +F S
Sbjct: 61 TPQELADYDAIIFGTPTRFGNMSGQMRTFLDQTGGLWASGALYGKLASVFSSTGTGGGQE 120
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVEL 666
T T L HHGMV VP+GY +LF + QV+GG+PYGA T A DGSR PS+ EL
Sbjct: 121 QTITST-WTTLAHHGMVIVPIGYA-AQELFDVSQVRGGTPYGATTIAGGDGSRQPSQEEL 178
Query: 667 EHAFHQGKYFAGIAKKLKG 723
A +QG+Y AG+A KL G
Sbjct: 179 SIARYQGEYVAGLAVKLNG 197
>sw|Q8ZF61|WRBA_YERPE Flavoprotein wrbA (Trp repressor binding
protein).
Length = 199
Score = 173 bits (439), Expect = 3e-42
Identities = 99/200 (49%), Positives = 118/200 (59%), Gaps = 2/200 (1%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
KI V+YYSMYGH+ LA I +GA V GV+ I +VPE + E K G PV
Sbjct: 3 KILVLYYSMYGHIETLAGAIAEGARKVSGVDVTIKRVPETMPAEAFAKAGGKTNQQAPVA 62
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
TP ELA+ DGI+FG PTRFG M+ QM+ F D TGGLW +L GK A +F S
Sbjct: 63 TPHELADYDGIIFGTPTRFGNMSGQMRTFLDQTGGLWASGALYGKVASVFASTGTGGGQE 122
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAK-LFGMDQVQGGSPYGAGTFA-ADGSRWPSEVE 663
T T L HHG + VP+GY GAK LF + Q +GG+PYGA T A DGSR PS E
Sbjct: 123 HTITST-WTTLAHHGFIIVPIGY--GAKELFDVSQTRGGTPYGATTIAGGDGSRQPSAEE 179
Query: 664 LEHAFHQGKYFAGIAKKLKG 723
L A QG++ A I KLKG
Sbjct: 180 LAIARFQGEHVAKITAKLKG 199
>sw|P42058|ALA7_ALTAL Minor allergen Alt a 7 (Alt a VII).
Length = 204
Score = 173 bits (438), Expect = 4e-42
Identities = 100/202 (49%), Positives = 131/202 (64%), Gaps = 5/202 (2%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPD- 297
MA KI +VYYSMYGH+ K+AD KG + G +AK++QV E L +EVL KM APPK
Sbjct: 1 MAPKIAIVYYSMYGHIKKMADAELKG-IQEAGGDAKLFQVAETLPQEVLDKMYAPPKDSS 59
Query: 298 VPVIT-PQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXX 474
VPV+ P L E DGILFG PTR+G AQ K F+D TG W++ + GK AG+F S
Sbjct: 60 VPVLEDPAVLEEFDGILFGIPTRYGNFPAQFKTFWDKTGKQWQQGAFWGKYAGVFVSTGT 119
Query: 475 XXXXXXXXPLTAITQLTHHGMVFVPVGY-TFGAKLFGMDQVQGGSPYGAGTFAA-DGSRW 648
+T+++ L HG ++VP+GY T + L +D+V GGSP+GAGTF+A DGSR
Sbjct: 120 LGGGQETTAITSMSTLVDHGFIYVPLGYKTAFSMLANLDEVHGGSPWGAGTFSAGDGSRQ 179
Query: 649 PSEVELEHAFHQGK-YFAGIAK 711
PSE+EL A QGK ++ +AK
Sbjct: 180 PSELELNIAQAQGKAFYEAVAK 201
>sptr|Q8X194|Q8X194 Y20 protein.
Length = 203
Score = 172 bits (435), Expect = 8e-42
Identities = 94/201 (46%), Positives = 123/201 (61%), Gaps = 2/201 (0%)
Frame = +1
Query: 121 MAVKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDV 300
MA KI +V+YS+YGH+ KLA+ KKG + G A I+Q+ E L +EVL KM AP K
Sbjct: 1 MAPKIAIVFYSLYGHIQKLAEAEKKG-IEAAGGTADIYQIAETLPQEVLDKMHAPGKSSY 59
Query: 301 PVITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXX 480
PV P L D LFG PTR+G Q KAF+D TGG+W GK AG+F S
Sbjct: 60 PVAEPATLLNYDAFLFGIPTRYGNFPGQWKAFWDKTGGIWSTGGFWGKYAGLFVSTGTPG 119
Query: 481 XXXXXXPLTAITQLTHHGMVFVPVGY-TFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPS 654
+ A++ L HHG+++VP+GY T L +++V+GGSP+GAGT+A ADGSR PS
Sbjct: 120 GGQESTNIAAMSTLAHHGIIYVPLGYKTTFPILANLNEVRGGSPWGAGTYAGADGSRQPS 179
Query: 655 EVELEHAFHQGKYFAGIAKKL 717
+EL+ A QGK F G K+
Sbjct: 180 ALELQLAEEQGKAFYGAVSKV 200
>sw|P25349|YCP4_YEAST Hypothetical 26.4 kDa protein in CDC10-CIT2
intergenic region.
Length = 247
Score = 171 bits (432), Expect = 2e-41
Identities = 97/200 (48%), Positives = 124/200 (62%), Gaps = 3/200 (1%)
Frame = +1
Query: 127 VKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKP-DVP 303
VKI ++ YS YGH+ LA +KKG + G +A I++V E L +EVL KM AP KP D+P
Sbjct: 2 VKIAIITYSTYGHIDVLAQAVKKG-VEAAGGKADIYRVEETLPDEVLTKMNAPQKPEDIP 60
Query: 304 VITPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXX 483
V T + L E D LFG PTRFG + AQ AF+D TGGLW + SL GK AG+F S
Sbjct: 61 VATEKTLLEYDAFLFGVPTRFGNLPAQWSAFWDKTGGLWAKGSLNGKAAGIFVSTSSYGG 120
Query: 484 XXXXXPLTAITQLTHHGMVFVPVGYTFG-AKLFGMDQVQGGSPYGAGTFAA-DGSRWPSE 657
++ L HHG++F+P+GY A+L +++V GGSP+GAGT A DGSR S
Sbjct: 121 GQESTVKACLSYLAHHGIIFLPLGYKNSFAELASIEEVHGGSPWGAGTLAGPDGSRTASP 180
Query: 658 VELEHAFHQGKYFAGIAKKL 717
+EL A QGK F AKKL
Sbjct: 181 LELRIAEIQGKTFYETAKKL 200
>sw|Q12335|PST2_YEAST Protoplast secreted protein 2 precursor.
Length = 198
Score = 164 bits (416), Expect = 1e-39
Identities = 90/191 (47%), Positives = 118/191 (61%), Gaps = 2/191 (1%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
++ ++ Y++YGHV A+ KKG + G A I+QV E LS EV+ +G PKPD P+
Sbjct: 3 RVAIIIYTLYGHVAATAEAEKKG-IEAAGGSADIYQVEETLSPEVVKALGGAPKPDYPIA 61
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
T L E D LFG PTRFG AQ KAF+D TGGLW + +L GK AG F S
Sbjct: 62 TQDTLTEYDAFLFGIPTRFGNFPAQWKAFWDRTGGLWAKGALHGKVAGCFVSTGTGGGNE 121
Query: 490 XXXPLTAITQLTHHGMVFVPVGY-TFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVE 663
+ +++ L HHG++FVP+GY A+L MD+V GGSP+GAGT A +DGSR PS +E
Sbjct: 122 ATI-MNSLSTLAHHGIIFVPLGYKNVFAELTNMDEVHGGSPWGAGTIAGSDGSRSPSALE 180
Query: 664 LEHAFHQGKYF 696
L+ QGK F
Sbjct: 181 LQVHEIQGKTF 191
>sw|P30821|P25_SCHPO P25 protein (Brefeldin A resistance protein).
Length = 202
Score = 163 bits (412), Expect = 4e-39
Identities = 91/191 (47%), Positives = 121/191 (63%), Gaps = 3/191 (1%)
Frame = +1
Query: 133 IYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVIT 312
+ +V YS YGHV KLA E +K + G +A I+Q PE LS E+L KM A PKP+ PV+T
Sbjct: 7 VAIVIYSTYGHVVKLA-EAEKAGIEKAGGKAVIYQFPETLSPEILEKMHAAPKPNYPVVT 65
Query: 313 PQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXXX 492
L + D LFG+PTR+G AQ + F+D+TGGLW + +L GK G FFS
Sbjct: 66 LDVLTQYDAFLFGYPTRYGTPPAQFRTFWDSTGGLWVQGALHGKYFGQFFSTGTLGGGQE 125
Query: 493 XXPLTAITQLTHHGMVFVPVGY--TFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVE 663
LTA+T HHGM+FVP+GY TF + + ++ + GGS +GAG++A ADGSR S+ E
Sbjct: 126 STALTAMTSFVHHGMIFVPLGYKNTF-SLMANVESIHGGSSWGAGSYAGADGSRNVSDDE 184
Query: 664 LEHAFHQGKYF 696
LE A QG+ F
Sbjct: 185 LEIARIQGETF 195
>sw|P58796|WRBA_METAC Flavoprotein wrbA.
Length = 209
Score = 158 bits (399), Expect = 1e-37
Identities = 92/208 (44%), Positives = 120/208 (57%), Gaps = 9/208 (4%)
Frame = +1
Query: 127 VKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPK----P 294
VK+ +++YSMYGHV ++A+ + GA VEG E I+QVPE L EEVL KMGA
Sbjct: 2 VKVNIIFYSMYGHVYRMAEAVAAGAREVEGAEVGIYQVPETLPEEVLEKMGAIETKKLFA 61
Query: 295 DVPVIT----PQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFF 462
+PV+T + LA AD ++FG PTR+G M AQM+A D GGLW + GK +F
Sbjct: 62 HIPVLTREMNEEVLAGADALIFGTPTRYGNMTAQMRAVLDGLGGLWNRDAFVGKVGSVFT 121
Query: 463 SXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADG 639
S LT L H GM+ V + Y+ + MD++ GGSPYG T A DG
Sbjct: 122 SSGTQHGGQESTILTFHVTLLHLGMILVGLPYS-EKRQTRMDEITGGSPYGVSTIAGGDG 180
Query: 640 SRWPSEVELEHAFHQGKYFAGIAKKLKG 723
SR PSE EL A +QG++ IAKK+ G
Sbjct: 181 SRQPSENELAMARYQGRHVTLIAKKIAG 208
>sw|Q8PUV4|WRBA_METMA Flavoprotein wrbA.
Length = 209
Score = 151 bits (381), Expect = 1e-35
Identities = 88/208 (42%), Positives = 120/208 (57%), Gaps = 9/208 (4%)
Frame = +1
Query: 127 VKIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPD--- 297
VK+ ++++S++ H+ ++A+ + GA VEG E I+QVPE L E+VL KMGA
Sbjct: 2 VKVNIIFHSVHAHIYRMAEAVAAGAREVEGAEVGIYQVPETLPEDVLEKMGAIETKKLFA 61
Query: 298 -VPVIT----PQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFF 462
+PV+T LA AD ++FG PTR+GMM AQM+A D G LW E + GK +F
Sbjct: 62 HIPVVTRDMYEDVLAGADALIFGTPTRYGMMTAQMRAVLDGLGKLWSEDAFVGKVGSVFT 121
Query: 463 SXXXXXXXXXXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADG 639
S L+ L H GMV V + Y + M+++ GGSPYGA T A DG
Sbjct: 122 SSGTQHGGQESTILSFHVTLLHLGMVIVGLPYAEKRQTI-MNEITGGSPYGASTIAGGDG 180
Query: 640 SRWPSEVELEHAFHQGKYFAGIAKKLKG 723
SR PSE ELE A +QG++ IAKK+ G
Sbjct: 181 SRQPSENELEMARYQGRHVTQIAKKIAG 208
>sw|Q92PU3|WRB1_RHIME Flavoprotein wrbA1.
Length = 199
Score = 146 bits (369), Expect = 4e-34
Identities = 83/197 (42%), Positives = 113/197 (57%), Gaps = 1/197 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
++ V+YYS YGH+ +A + +GA S G E + +VPE++ E+V PV
Sbjct: 3 RVLVLYYSAYGHIETMAHAVAEGARSA-GAEVAVKRVPELVPEDVAKASHFKLDQPAPVA 61
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
T +ELA+ D I+FG TR+G +A+Q++ F D TGGLW + L GK F S
Sbjct: 62 TVEELADYDAIIFGAGTRYGTVASQLRNFIDQTGGLWAKGKLVGKVGSAFTSSATQHGGQ 121
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTF-AADGSRWPSEVEL 666
L I + HHGMV V + Y F ++ G+++V+GGSPYGA T DGSR PS VEL
Sbjct: 122 ESTILGLIPTMMHHGMVVVGLPYAFQGQM-GVEEVKGGSPYGASTITGGDGSRQPSAVEL 180
Query: 667 EHAFHQGKYFAGIAKKL 717
E A QG + A IA KL
Sbjct: 181 EAARFQGAHVARIAAKL 197
>sw|P58795|WRBA_AGRT5 Flavoprotein wrbA.
Length = 199
Score = 144 bits (364), Expect = 1e-33
Identities = 83/197 (42%), Positives = 110/197 (55%), Gaps = 1/197 (0%)
Frame = +1
Query: 130 KIYVVYYSMYGHVGKLADEIKKGALSVEGVEAKIWQVPEILSEEVLGKMGAPPKPDVPVI 309
K+ V+YYS YGH+ +A + +G S G EA + +VPE++ EEV PV
Sbjct: 3 KVLVLYYSSYGHIETMAYAVAEGVEST-GAEAVVKRVPELVPEEVAKSSHFKMDQPAPVA 61
Query: 310 TPQELAEADGILFGFPTRFGMMAAQMKAFFDATGGLWREQSLAGKPAGMFFSXXXXXXXX 489
T ELAE D I+ G TRFG +A+QM+ F+D TGGLW L GK F S
Sbjct: 62 TVDELAEYDAIIVGAGTRFGTVASQMRNFWDQTGGLWFSGKLVGKVGSAFTSSATQHGGQ 121
Query: 490 XXXPLTAITQLTHHGMVFVPVGYTFGAKLFGMDQVQGGSPYGAGTFA-ADGSRWPSEVEL 666
L I HHGM V + Y F ++ G+D+++GGSPYGA T DGSR PS +EL
Sbjct: 122 ESTILGFIPTFLHHGMAVVGLPYAFQGQM-GVDEIKGGSPYGASTITDGDGSRQPSAIEL 180
Query: 667 EHAFHQGKYFAGIAKKL 717
+ A +QG + A +A KL
Sbjct: 181 DAARYQGAHVAKLAAKL 197
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 786,681,514
Number of Sequences: 1272877
Number of extensions: 18502899
Number of successful extensions: 67516
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 60836
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 67104
length of database: 406,886,216
effective HSP length: 123
effective length of database: 250,322,345
effective search space used: 49313501965
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

