BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
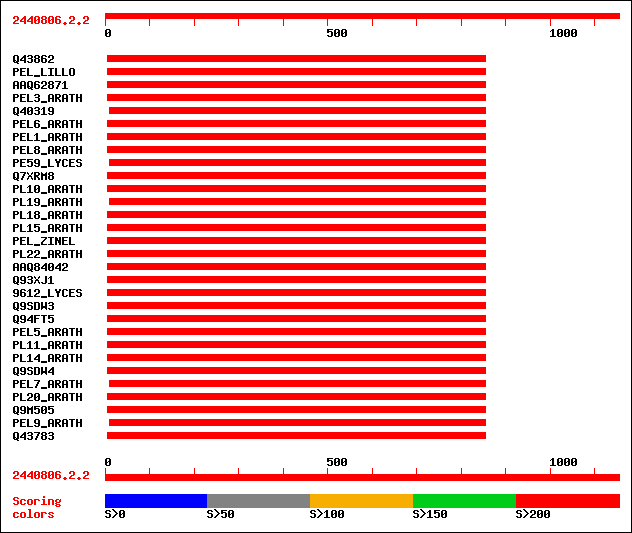
Query= 2440806.2.2
(1155 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q43862|Q43862 Pectate lyase homolog (EC 4.2.2.2). 535 e-151
sw|P40973|PEL_LILLO Pectate lyase precursor (EC 4.2.2.2). 350 2e-95
sptrnew|AAQ62871|AAQ62871 At1g14420. 319 6e-86
sw|Q9M9S2|PEL3_ARATH Probable pectate lyase 3 precursor (EC 4.2.... 319 6e-86
sptr|Q40319|Q40319 Pectate lyase homolog. 316 4e-85
sw|O64510|PEL6_ARATH Probable pectate lyase 6 precursor (EC 4.2.... 316 5e-85
sw|Q940Q1|PEL1_ARATH Probable pectate lyase 1 precursor (EC 4.2.... 308 8e-83
sw|Q9M8Z8|PEL8_ARATH Probable pectate lyase 8 precursor (EC 4.2.... 308 8e-83
sw|P15722|PE59_LYCES Probable pectate lyase P59 precursor (EC 4.... 306 4e-82
sptr|Q7XRM8|Q7XRM8 OSJNBa0095E20.8 protein. 305 7e-82
sw|Q9LJ42|PL10_ARATH Probable pectate lyase 10 precursor (EC 4.2... 305 7e-82
sw|Q9LFP5|PL19_ARATH Putative pectate lyase 19 precursor (EC 4.2... 304 1e-81
sw|Q9C5M8|PL18_ARATH Probable pectate lyase 18 precursor (EC 4.2... 304 2e-81
sw|Q944R1|PL15_ARATH Probable pectate lyase 15 precursor (EC 4.2... 303 3e-81
sw|O24554|PEL_ZINEL Pectate lyase precursor (EC 4.2.2.2) (ZePel). 303 3e-81
sw|Q93Z25|PL22_ARATH Probable pectate lyase 22 precursor (EC 4.2... 302 6e-81
sptrnew|AAQ84042|AAQ84042 Pectate lyase. 302 6e-81
sptr|Q93XJ1|Q93XJ1 Pectate lyase. 301 1e-80
sw|P24396|9612_LYCES Style development-specific protein 9612 pre... 300 2e-80
sptr|Q9SDW3|Q9SDW3 Pectate lyase 2. 300 4e-80
sptr|Q94FT5|Q94FT5 Pectate lyase (Fragment). 300 4e-80
sw|Q9FXD8|PEL5_ARATH Probable pectate lyase 5 precursor (EC 4.2.... 299 5e-80
sw|Q9LTZ0|PL11_ARATH Putative pectate lyase 11 precursor (EC 4.2... 299 6e-80
sw|Q9SVQ6|PL14_ARATH Putative pectate lyase 14 precursor (EC 4.2... 299 6e-80
sptr|Q9SDW4|Q9SDW4 Pectate lyase 1. 297 2e-79
sw|Q9SRH4|PEL7_ARATH Probable pectate lyase 7 precursor (EC 4.2.... 295 9e-79
sw|Q93WF1|PL20_ARATH Probable pectate lyase 20 precursor (EC 4.2... 295 9e-79
sptr|Q9M505|Q9M505 Pectate lyase. 291 2e-77
sw|Q9LRM5|PEL9_ARATH Putative pectate lyase 9 precursor (EC 4.2.... 289 6e-77
sptr|Q43783|Q43783 Pectate lyase (EC 4.2.2.2) (Fragment). 280 2e-74
>sptr|Q43862|Q43862 Pectate lyase homolog (EC 4.2.2.2).
Length = 438
Score = 535 bits (1378), Expect = e-151
Identities = 257/284 (90%), Positives = 257/284 (90%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQITXXXXXXXXXXXXXXXDS 184
DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQIT DS
Sbjct: 155 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQITLQNVQNVILHNLHIHDS 214
Query: 185 KAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITV 364
KAHSGGMIRDSKRHYGLRTR NVWIDHVSMSSCSDGLIDVVNGSTAITV
Sbjct: 215 KAHSGGMIRDSKRHYGLRTRSDGDGVSVLSSSNVWIDHVSMSSCSDGLIDVVNGSTAITV 274
Query: 365 SNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTH 544
SNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTH
Sbjct: 275 SNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTH 334
Query: 545 WIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAF 724
WIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAF
Sbjct: 335 WIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAF 394
Query: 725 FNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC
Sbjct: 395 FNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 438
>sw|P40973|PEL_LILLO Pectate lyase precursor (EC 4.2.2.2).
Length = 434
Score = 350 bits (899), Expect = 2e-95
Identities = 170/285 (59%), Positives = 204/285 (71%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
DRPLWI+F + MVI L+QELI+N++KTIDGRGA V I AQ+T D
Sbjct: 150 DRPLWIIFGKSMVIRLKQELIINNDKTIDGRGANVQIAGGAQLTVQFVHNVIIHGIHIHD 209
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K GG+IRDS++H G+RTR N+WIDHVS++ CSDGLIDV+ GSTAIT
Sbjct: 210 IKPGEGGLIRDSEKHSGIRTRSDGDGISIIGSSNIWIDHVSLARCSDGLIDVILGSTAIT 269
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN H T+HD VML GAS+ QD +MQVTVAFNHFGRGLVQRMPRCRYGF HVVNNDYT
Sbjct: 270 ISNCHLTEHDDVMLLGASDTYTQDEIMQVTVAFNHFGRGLVQRMPRCRYGFVHVVNNDYT 329
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HWIMYA+GG+ +PTIISQGNR+IAP AKEVTKR+Y ++ +W WKSQGD+ ++GA
Sbjct: 330 HWIMYAVGGSQHPTIISQGNRYIAPHIEAAKEVTKRDYAEPAEWSKWTWKSQGDLFVSGA 389
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF ESGG E KY + D I AK G +V +LTRF+G L C C
Sbjct: 390 FFVESGGPFENKYSKKDLIKAKPGTFVQRLTRFSGALNCKENMEC 434
>sptrnew|AAQ62871|AAQ62871 At1g14420.
Length = 459
Score = 319 bits (817), Expect = 6e-86
Identities = 159/292 (54%), Positives = 200/292 (68%), Gaps = 8/292 (2%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWI+FAR M+I+L+QEL++ +KTIDGRGA+V+IM A +T
Sbjct: 169 DGPLWIIFARSMIIKLQQELMITSDKTIDGRGARVYIMEGAGLTLQFVNNVIIHNIYVKH 228
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
+GG+IRDS+ H GLRT+ N+WIDHVSM+ C+DG+ID ++GSTA+T
Sbjct: 229 IVPGNGGLIRDSEAHIGLRTKSDGDGISLFGATNIWIDHVSMTRCADGMIDAIDGSTAVT 288
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SNSHFTDH VMLFGA ++ D MQ+TVAFNHFG+ L QRMPRCRYG HVVNNDYT
Sbjct: 289 ISNSHFTDHQEVMLFGARDEHVIDKKMQITVAFNHFGKRLEQRMPRCRYGTIHVVNNDYT 348
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGGNMNPTIISQGNRFIAP + AK++TKREYTPY ++K W W+S+GD +NGA
Sbjct: 349 HWEMYAIGGNMNPTIISQGNRFIAPPNEEAKQITKREYTPYGEWKSWNWQSEGDYFLNGA 408
Query: 722 FFNESGGQNERKY-------DRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F +SG N ++F P K G V +LT AG L C +G+ C
Sbjct: 409 YFVQSGKANAWSSKPKTPLPNKFTIRP-KPGTMVRKLTMDAGVLGCKLGEAC 459
>sw|Q9M9S2|PEL3_ARATH Probable pectate lyase 3 precursor (EC 4.2.2.2)
(Pectate lyase A2).
Length = 459
Score = 319 bits (817), Expect = 6e-86
Identities = 159/292 (54%), Positives = 200/292 (68%), Gaps = 8/292 (2%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWI+FAR M+I+L+QEL++ +KTIDGRGA+V+IM A +T
Sbjct: 169 DGPLWIIFARSMIIKLQQELMITSDKTIDGRGARVYIMEGAGLTLQFVNNVIIHNIYVKH 228
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
+GG+IRDS+ H GLRT+ N+WIDHVSM+ C+DG+ID ++GSTA+T
Sbjct: 229 IVPGNGGLIRDSEAHIGLRTKSDGDGISLFGATNIWIDHVSMTRCADGMIDAIDGSTAVT 288
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SNSHFTDH VMLFGA ++ D MQ+TVAFNHFG+ L QRMPRCRYG HVVNNDYT
Sbjct: 289 ISNSHFTDHQEVMLFGARDEHVIDKKMQITVAFNHFGKRLEQRMPRCRYGTIHVVNNDYT 348
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGGNMNPTIISQGNRFIAP + AK++TKREYTPY ++K W W+S+GD +NGA
Sbjct: 349 HWEMYAIGGNMNPTIISQGNRFIAPPNEEAKQITKREYTPYGEWKSWNWQSEGDYFLNGA 408
Query: 722 FFNESGGQNERKY-------DRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F +SG N ++F P K G V +LT AG L C +G+ C
Sbjct: 409 YFVQSGKANAWSSKPKTPLPNKFTIRP-KPGTMVRKLTMDAGVLGCKLGEAC 459
>sptr|Q40319|Q40319 Pectate lyase homolog.
Length = 450
Score = 316 bits (810), Expect = 4e-85
Identities = 153/284 (53%), Positives = 193/284 (67%), Gaps = 2/284 (0%)
Frame = +2
Query: 11 PLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXDSK 187
PLWI+F R MVI L QEL+V+ +KTIDGRGA V I A IT + K
Sbjct: 167 PLWIIFQRSMVITLTQELMVSSDKTIDGRGANVQIRDGAGITMQFVNNVIIHGLRIKNIK 226
Query: 188 AHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITVS 367
A +GG+IRDS H G+RTR N+WIDH+S+S C DGL+DV+ GSTA+T+S
Sbjct: 227 ARNGGLIRDSFDHIGVRTRSDGDAISVFGSSNIWIDHISLSDCEDGLVDVIQGSTAVTIS 286
Query: 368 NSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHW 547
N H T H+ VMLFGAS+ D +MQ+TVAFNHFG+GL+QRMPRCR+GFFHV+NNDYTHW
Sbjct: 287 NCHMTKHNDVMLFGASDTYQDDKIMQITVAFNHFGQGLIQRMPRCRWGFFHVLNNDYTHW 346
Query: 548 IMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAFF 727
IMYAIGG+ PTI+SQGNRFIAP + AK VT R+Y P + +W W+S+GD MNGA F
Sbjct: 347 IMYAIGGSSAPTILSQGNRFIAPHNNAAKTVTHRDYAPESVWSKWQWRSEGDHFMNGATF 406
Query: 728 NESGGQ-NERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+SG + + + +HG +LTRF+G L C+VG+PC
Sbjct: 407 IQSGPPIKNLPFKKGFLMKPRHGSQANRLTRFSGALNCVVGRPC 450
>sw|O64510|PEL6_ARATH Probable pectate lyase 6 precursor (EC 4.2.2.2).
Length = 455
Score = 316 bits (809), Expect = 5e-85
Identities = 160/291 (54%), Positives = 198/291 (68%), Gaps = 7/291 (2%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
DRPLWI+FAR M+I+L+QELI+ ++KTIDGRGA+++I A +T
Sbjct: 165 DRPLWIIFARSMIIKLQQELIITNDKTIDGRGAKIYITGGAGLTLQFVRNVIIHNIHIKQ 224
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K +GG+I DS++H+GLRT NVWIDHVSM+ CSDG+ID + GSTAIT
Sbjct: 225 IKRGAGGLIIDSEQHFGLRTVSDGDGINIFGATNVWIDHVSMTDCSDGMIDAIMGSTAIT 284
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SNSHFTDHD VMLFG +N D MQ+TVAFNHFG+ L QRMPR R+G HVVNNDYT
Sbjct: 285 ISNSHFTDHDEVMLFGGTNKDVIDKKMQITVAFNHFGKRLKQRMPRVRFGLVHVVNNDYT 344
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGGNMNPTIISQGNRFIAP ++K+VTKREYTPY ++K W W+S+ D +NGA
Sbjct: 345 HWEMYAIGGNMNPTIISQGNRFIAPPIEDSKQVTKREYTPYPEWKSWNWQSEKDYFLNGA 404
Query: 722 FFNESGGQNERKYDRFDFIPAKH------GRYVGQLTRFAGPLKCIVGQPC 856
+F +SG N + IP K G V +LT+ AG L C G+ C
Sbjct: 405 YFVQSGKANAWSATPKNPIPRKFAIRPQPGTKVRRLTKDAGTLGCKPGKSC 455
>sw|Q940Q1|PEL1_ARATH Probable pectate lyase 1 precursor (EC
4.2.2.2) (Pectate lyase A1).
Length = 431
Score = 308 bits (790), Expect = 8e-83
Identities = 154/285 (54%), Positives = 185/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
DRPLWIVF RDMVI+L+QELIVN KTIDGRGA VHI IT D
Sbjct: 145 DRPLWIVFKRDMVIQLKQELIVNSFKTIDGRGANVHIANGGCITIQFVTNVIVHGLHIHD 204
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S+ H+G RT +VWIDH S+S C+DGL+D V GSTAIT
Sbjct: 205 CKPTGNAMVRSSETHFGWRTMADGDAISIFGSSHVWIDHNSLSHCADGLVDAVMGSTAIT 264
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN+H T H+ VML G S+ +D MQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 265 ISNNHLTHHNEVMLLGHSDSYMRDKAMQVTIAYNHFGVGLIQRMPRCRHGYFHVVNNDYT 324
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ NPTI SQGNR+ AP +P AKEVTKR TP +K W W+S+GD++ NGA
Sbjct: 325 HWEMYAIGGSANPTINSQGNRYAAPKNPFAKEVTKRVDTPASHWKGWNWRSEGDLLQNGA 384
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F SG Y R + AK VG +T AG L C G+ C
Sbjct: 385 YFTSSGAAASGSYARASSLSAKSSSLVGHITSDAGALPCRRGRQC 429
>sw|Q9M8Z8|PEL8_ARATH Probable pectate lyase 8 precursor (EC
4.2.2.2).
Length = 416
Score = 308 bits (790), Expect = 8e-83
Identities = 151/285 (52%), Positives = 185/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWI+F RDMVI L+QELI+N KTIDGRG VHI A +T D
Sbjct: 132 DEPLWIIFKRDMVITLKQELIMNSFKTIDGRGVNVHIANGACLTIQYVTNIIVHGIHVHD 191
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S HYG R+ ++WIDH S+S+C+DGL+D V STAIT
Sbjct: 192 CKPTGNAMVRSSPSHYGFRSMADGDAISIFGSSHIWIDHNSLSNCADGLVDAVMSSTAIT 251
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
VSN+ FT H+ VML G S+ +D VMQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 252 VSNNFFTHHNEVMLLGHSDSYTRDKVMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYT 311
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNRF+AP +P AKEVTKREYT +K W W+S+GD+ +NGA
Sbjct: 312 HWEMYAIGGSAGPTINSQGNRFLAPVNPFAKEVTKREYTGESKWKHWNWRSEGDLFLNGA 371
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK VG +T ++G L C G+ C
Sbjct: 372 FFTRSGAGAGANYARASSLSAKSSSLVGTMTSYSGALNCRAGRRC 416
>sw|P15722|PE59_LYCES Probable pectate lyase P59 precursor (EC
4.2.2.2).
Length = 449
Score = 306 bits (784), Expect = 4e-82
Identities = 153/285 (53%), Positives = 187/285 (65%), Gaps = 3/285 (1%)
Frame = +2
Query: 11 PLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXDSK 187
PLWI+F R M I L QE+I+ +KTID RG VHI A IT D
Sbjct: 165 PLWIIFKRGMNIRLHQEMIMQSDKTIDARGVNVHITKGAGITLQYIKNVIIHGLHIHDIV 224
Query: 188 AHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITVS 367
+GGM+RD+ H G+RT+ +WIDHVSM C DGLID V GST IT+S
Sbjct: 225 EGNGGMVRDAVDHIGIRTKSDGDGISIFGASYIWIDHVSMQRCYDGLIDAVEGSTGITIS 284
Query: 368 NSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHW 547
N HFTDH+ VMLFGAS+ S D VMQ+T+AFNHFG+ L+QRMPRCR+G+ HVVNNDYTHW
Sbjct: 285 NGHFTDHNEVMLFGASDSSSIDQVMQITLAFNHFGKRLIQRMPRCRWGYIHVVNNDYTHW 344
Query: 548 IMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAFF 727
MYAIGG+M+PTII QGNRFIAP D K+VTKREY P + +W W+S+G++ MNGA+F
Sbjct: 345 NMYAIGGSMHPTIIHQGNRFIAPPDIFKKQVTKREYNPESVWMQWTWRSEGNLFMNGAYF 404
Query: 728 NESGGQ--NERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
ESG + + D +D I A V +TRFAG L C G+PC
Sbjct: 405 TESGDPEWSSKHKDLYDGISAAPAEDVTWMTRFAGVLGCKPGKPC 449
>sptr|Q7XRM8|Q7XRM8 OSJNBa0095E20.8 protein.
Length = 472
Score = 305 bits (782), Expect = 7e-82
Identities = 152/285 (53%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWIVF RDMVI L+QELI+N KTIDGRGA VHI A IT D
Sbjct: 188 DEPLWIVFKRDMVITLKQELIMNSFKTIDGRGANVHIANGACITIQYVTNVIIHGLHIHD 247
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
+ M+R S HYG RT ++W+DH S+S+C+DGLID + GSTAIT
Sbjct: 248 CRPTGNAMVRSSPSHYGWRTMADGDAVSIFGASHIWVDHCSLSNCADGLIDAIMGSTAIT 307
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
VSN++FT H+ VML G S+ +D MQVT+AFNHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 308 VSNNYFTHHNEVMLLGHSDSYVKDKAMQVTIAFNHFGEGLIQRMPRCRHGYFHVVNNDYT 367
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNR++AP +P AKEVTKR T +K W W+S+GD+++NGA
Sbjct: 368 HWEMYAIGGSAEPTINSQGNRYLAPTNPFAKEVTKRVETAQTIWKGWNWRSEGDLLLNGA 427
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK VG +T AG L C G C
Sbjct: 428 FFTPSGAGASASYSRASSLGAKSSSMVGTITSGAGALSCRGGSAC 472
>sw|Q9LJ42|PL10_ARATH Probable pectate lyase 10 precursor (EC
4.2.2.2).
Length = 440
Score = 305 bits (782), Expect = 7e-82
Identities = 151/285 (52%), Positives = 188/285 (65%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
DRPLWIVF RDMVI L QELI+N KTIDGRG V I A IT D
Sbjct: 156 DRPLWIVFKRDMVITLTQELIMNSFKTIDGRGVNVAIAGGACITIQYVTNIIIHGINVHD 215
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
+ M+R S HYG RT ++WIDH S+S+C+DGLID + GSTAIT
Sbjct: 216 CRRTGNAMVRSSPSHYGWRTMADGDAISIFGSSHIWIDHNSLSNCADGLIDAIMGSTAIT 275
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T H+ VML G S+ +D +MQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 276 ISNNYMTHHNEVMLMGHSDSYTRDKLMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYT 335
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW+MYAIGG+ NPTI SQGNRF+AP +P AKEVTKR + ++K+W W+SQGD+M+NGA
Sbjct: 336 HWVMYAIGGSANPTINSQGNRFLAPGNPFAKEVTKRVGSWQGEWKQWNWRSQGDLMLNGA 395
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F +SG Y R + AK V LT +G LKC +G C
Sbjct: 396 YFTKSGAAAPASYARASSLGAKPASVVSMLTYSSGALKCRIGMRC 440
>sw|Q9LFP5|PL19_ARATH Putative pectate lyase 19 precursor (EC
4.2.2.2).
Length = 472
Score = 304 bits (779), Expect = 1e-81
Identities = 152/285 (53%), Positives = 187/285 (65%), Gaps = 3/285 (1%)
Frame = +2
Query: 11 PLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXDSK 187
PLWI+F DM I L QEL++N +KTID RGA VH+ A IT
Sbjct: 188 PLWIIFKNDMSIRLNQELLINSHKTIDARGANVHVAHGAGITMQFVKNVIIHGLHIHHIS 247
Query: 188 AHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITVS 367
SGGMIRDS H+G+RTR N+W+DH+SMS C DGLID + GST IT+S
Sbjct: 248 ESSGGMIRDSVDHFGMRTRADGDGLSIYGSSNIWLDHISMSKCQDGLIDAIVGSTGITIS 307
Query: 368 NSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHW 547
NSHFT H+ VML GA N + D MQVTVA+NHFG+GLVQRMPR R+GF HVVNNDYTHW
Sbjct: 308 NSHFTHHNDVMLLGAQNTNEADKHMQVTVAYNHFGKGLVQRMPRIRWGFVHVVNNDYTHW 367
Query: 548 IMYAIGGNMNPTIISQGNRFIA-PDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAF 724
+YAIGG+ PTI+S GNRFIA P P+ +EVTKR+Y ++K W W+S DV MNGA+
Sbjct: 368 ELYAIGGSQGPTILSHGNRFIAPPHKPHYREVTKRDYASEDEWKHWNWRSDKDVFMNGAY 427
Query: 725 FNESGG-QNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
F +SG Q + + R I K+G V +LT++AG L C VG+ C
Sbjct: 428 FRQSGNPQYKCAHTRQQMIKPKNGLAVSKLTKYAGALDCRVGRRC 472
>sw|Q9C5M8|PL18_ARATH Probable pectate lyase 18 precursor (EC
4.2.2.2) (Pectate lyase A10).
Length = 408
Score = 304 bits (778), Expect = 2e-81
Identities = 151/285 (52%), Positives = 182/285 (63%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDM I+L++ELI+N KTIDGRGA VHI IT D
Sbjct: 124 DEPLWIIFQRDMTIQLKEELIMNSFKTIDGRGASVHISGGPCITIQYVTNIIIHGIHIHD 183
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S RH+G RT +VW+DH S S+C DGLID + GSTAIT
Sbjct: 184 CKQGGNAMVRSSPRHFGWRTISDGDGVSIFGGSHVWVDHCSFSNCEDGLIDAIMGSTAIT 243
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN+H T HD VML G S+ +D MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 244 LSNNHMTHHDKVMLLGHSDTYSRDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 303
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ NPTI SQGNRF+AP+ +KEVTK E P ++K W W+S GD+++NGA
Sbjct: 304 HWEMYAIGGSANPTINSQGNRFLAPNIRFSKEVTKHEDAPESEWKRWNWRSSGDLLLNGA 363
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SGG Y + + AK VG LT +G L C G C
Sbjct: 364 FFTPSGGAASSSYAKASSLGAKPSSLVGPLTSTSGALNCRKGSRC 408
>sw|Q944R1|PL15_ARATH Probable pectate lyase 15 precursor (EC 4.2.2.2)
(Pectate lyase A11).
Length = 470
Score = 303 bits (777), Expect = 3e-81
Identities = 152/285 (53%), Positives = 185/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
+ PLWIVF RDMVIEL+QELI+N KTID RG+ VHI A IT D
Sbjct: 186 EEPLWIVFKRDMVIELKQELIMNSFKTIDARGSNVHIANGACITIQFITNVIIHGLHIHD 245
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S H+G RT ++WIDH S+S C+DGL+D V GSTAIT
Sbjct: 246 CKPTGNAMVRSSPSHFGWRTMADGDAVSIFGSSHIWIDHNSLSHCADGLVDAVMGSTAIT 305
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
VSN+HFT H+ VML G S+ +D +MQVT+A+NHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 306 VSNNHFTHHNEVMLLGHSDSYTKDKLMQVTIAYNHFGEGLVQRMPRCRHGYFHVVNNDYT 365
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNR+ AP D AKEVTKR T ++K+W W+S+GD+++NGA
Sbjct: 366 HWEMYAIGGSAEPTINSQGNRYAAPMDRFAKEVTKRVETDASEWKKWNWRSEGDLLLNGA 425
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK V +T AG L C G+PC
Sbjct: 426 FFRPSGAGASASYGRASSLAAKPSSMVDTITSTAGALGCRKGRPC 470
>sw|O24554|PEL_ZINEL Pectate lyase precursor (EC 4.2.2.2) (ZePel).
Length = 401
Score = 303 bits (776), Expect = 3e-81
Identities = 149/285 (52%), Positives = 185/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDMVI+LRQEL++N +KTIDGRG VHI IT D
Sbjct: 117 DEPLWIIFKRDMVIQLRQELVMNSHKTIDGRGVNVHIGNGPCITIHYASNIIIHGIHIHD 176
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K G IR+S H G T+ ++WIDH S+S+C DGLID ++GSTAIT
Sbjct: 177 CKQAGNGNIRNSPHHSGWWTQSDGDGISIFASKDIWIDHNSLSNCHDGLIDAIHGSTAIT 236
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T HD VML G S+ QD MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 237 ISNNYMTHHDKVMLLGHSDSYTQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 296
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNRF+AP+ KEVTK E P ++K W W+S+GD+M+NGA
Sbjct: 297 HWEMYAIGGSASPTIYSQGNRFLAPNTRFDKEVTKHENAPESEWKNWNWRSEGDLMLNGA 356
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F ESGG+ + R + + V +TR AG L C G C
Sbjct: 357 YFRESGGRAASSFARASSLSGRPSTLVASMTRSAGALVCRKGSRC 401
>sw|Q93Z25|PL22_ARATH Probable pectate lyase 22 precursor (EC
4.2.2.2).
Length = 432
Score = 302 bits (774), Expect = 6e-81
Identities = 147/287 (51%), Positives = 187/287 (65%), Gaps = 3/287 (1%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDM I+L++ELI+N KT+DGRGA VHI IT D
Sbjct: 146 DEPLWIIFKRDMTIQLKEELIMNSFKTLDGRGASVHISGGPCITIQYVTNIIIHGLHIHD 205
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K +RDS HYG RT +VW+DH S+S+C+DGLID + GSTAIT
Sbjct: 206 CKQGGNTYVRDSPEHYGYRTVSDGDGVSIFGGSHVWVDHCSLSNCNDGLIDAIRGSTAIT 265
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T H+ VML G S+ QD MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 266 ISNNYLTHHNKVMLLGHSDTYEQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 325
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ NPTI SQGNRF+APDD ++KEVTK E P +++ W W+S+GD+++NGA
Sbjct: 326 HWEMYAIGGSANPTINSQGNRFLAPDDSSSKEVTKHEDAPEDEWRNWNWRSEGDLLLNGA 385
Query: 722 FFNESGG--QNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y + + A+ +VG++T +G L C G C
Sbjct: 386 FFTYSGAGPAKSSSYSKASSLAARPSSHVGEITIASGALSCKRGSHC 432
>sptrnew|AAQ84042|AAQ84042 Pectate lyase.
Length = 418
Score = 302 bits (774), Expect = 6e-81
Identities = 148/285 (51%), Positives = 182/285 (63%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDM I+L++ELI+N KTIDGRGA VHI IT D
Sbjct: 134 DEPLWIIFQRDMTIQLKEELIMNSFKTIDGRGASVHIAGGPCITIQFVTNIIIHGLHIHD 193
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S RH+G RT +VW+DH S+S+C DGL+D + GSTAIT
Sbjct: 194 CKQGGNAMVRSSPRHFGWRTVSDGDGVSIFGGSHVWVDHCSLSNCKDGLVDAIYGSTAIT 253
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T HD VML G S+ D MQ+T+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 254 ISNNYMTHHDKVMLLGHSDSYTNDKNMQITIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 313
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNRF APD ++KEVTK E P ++K W W+S+GD+M+NGA
Sbjct: 314 HWEMYAIGGSADPTINSQGNRFAAPDIRSSKEVTKHEDAPESEWKNWNWRSEGDLMLNGA 373
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK VG +T +G L C G C
Sbjct: 374 FFTASGAGASSSYARASSLGAKPSSLVGAITTASGALSCRKGSRC 418
>sptr|Q93XJ1|Q93XJ1 Pectate lyase.
Length = 409
Score = 301 bits (771), Expect = 1e-80
Identities = 151/285 (52%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWIVFARDMVI+LR+ELI+N KTIDGRGA VHI IT D
Sbjct: 125 DEPLWIVFARDMVIKLREELIMNSFKTIDGRGASVHIAGGPCITIQYVTNIIIHGVNIHD 184
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K +RDS HYG RT +VW+DH S+S+C+DGLID ++GSTAIT
Sbjct: 185 CKRGGNAHVRDSPSHYGWRTVSDGDGVSIFGGSHVWVDHCSLSNCNDGLIDAIHGSTAIT 244
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T H+ VML G S+ QD MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 245 ISNNYLTHHNKVMLLGHSDSYKQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 304
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNRF+AP+D KEVTK E P +K W W+S+GD+++NGA
Sbjct: 305 HWKMYAIGGSADPTINSQGNRFLAPNDRFNKEVTKHEDAPQSAWKGWNWRSEGDLLLNGA 364
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y + + A+ V +T AG L C G C
Sbjct: 365 FFTASGAGASSSYAKASSLGARSSSLVSSITAGAGSLVCKKGSRC 409
>sw|P24396|9612_LYCES Style development-specific protein 9612
precursor.
Length = 404
Score = 300 bits (769), Expect = 2e-80
Identities = 149/287 (51%), Positives = 184/287 (64%), Gaps = 3/287 (1%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDMVI+L+QEL++N KTIDGRGA VHI IT D
Sbjct: 118 DEPLWIIFKRDMVIQLKQELVMNSYKTIDGRGASVHISGGPCITIHHTSNIIIHGINIHD 177
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K G IRDS H G N+W+DH S+S+C DGLID ++GSTAIT
Sbjct: 178 CKQSGNGNIRDSPNHSGWWDVSDGDGISIFGGKNIWVDHCSLSNCHDGLIDAIHGSTAIT 237
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++FT HD VML G S+ QD MQVTVAFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 238 ISNNYFTHHDKVMLLGHSDSFTQDKGMQVTVAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 297
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNRF+AP++ KEVTK E P ++ W W+S+GD+M+NGA
Sbjct: 298 HWEMYAIGGSAAPTINSQGNRFLAPNEKYRKEVTKHEDAPESQWRSWNWRSEGDLMLNGA 357
Query: 722 FFNE--SGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F + +G + Y R + A+ VG +T AGP+ C G C
Sbjct: 358 YFRQTGAGASSSSTYARASSLSARPSSLVGSITTNAGPVNCKKGSRC 404
>sptr|Q9SDW3|Q9SDW3 Pectate lyase 2.
Length = 454
Score = 300 bits (767), Expect = 4e-80
Identities = 149/285 (52%), Positives = 182/285 (63%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWI F DM I L++ELI+N KTIDGRG VHI A IT D
Sbjct: 170 DVPLWITFKHDMEITLKEELIMNSFKTIDGRGVNVHIANGACITIQYITNVIIHGLHIHD 229
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S HYG RT ++W+DH S+S+C+DGL+D V GSTAIT
Sbjct: 230 CKPTGNAMVRSSPSHYGWRTMADGDAVSIFGSSHIWVDHCSLSNCADGLVDAVMGSTAIT 289
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
VSN++FT H+ VML G ++ +D++MQVT+AFNHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 290 VSNNYFTHHNEVMLLGHTDSYARDSIMQVTIAFNHFGEGLIQRMPRCRHGYFHVVNNDYT 349
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ NPTI SQGNR++AP +P AKEVTKR T +K W W+S+GD+++NGA
Sbjct: 350 HWEMYAIGGSANPTINSQGNRYLAPTNPFAKEVTKRVDTDQSTWKNWNWRSEGDLLLNGA 409
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R AK V LT AG L C VG C
Sbjct: 410 FFTPSGAGASASYARASSFGAKPSSLVDTLTSDAGVLSCQVGTRC 454
>sptr|Q94FT5|Q94FT5 Pectate lyase (Fragment).
Length = 368
Score = 300 bits (767), Expect = 4e-80
Identities = 148/285 (51%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWIVF RDMVI+L+QELI+N KTIDGRG VHI A IT D
Sbjct: 84 DEPLWIVFKRDMVIQLKQELIMNSFKTIDGRGVNVHIANGACITIQFVTNVIVHGLHIHD 143
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S H+G RT ++W+DH S+S+C+DGL+D V GSTAIT
Sbjct: 144 CKPTGNAMVRSSPSHFGWRTMADGDAISIFGSSHIWVDHNSLSNCADGLVDAVMGSTAIT 203
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN+H T H+ VML G S+ +D MQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 204 ISNNHLTHHNEVMLLGHSDSYTRDKQMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYT 263
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNR+ AP +P AKEVTKR T ++ W W+S+GD+++NGA
Sbjct: 264 HWEMYAIGGSADPTINSQGNRYAAPTNPFAKEVTKRVETSQTQWRGWNWRSEGDLLLNGA 323
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK VG +T AG L C G+ C
Sbjct: 324 FFTPSGAGASAVYARASSLGAKSSAMVGTITASAGALGCRRGRTC 368
>sw|Q9FXD8|PEL5_ARATH Probable pectate lyase 5 precursor (EC
4.2.2.2).
Length = 408
Score = 299 bits (766), Expect = 5e-80
Identities = 149/285 (52%), Positives = 185/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWI+FARDMVI+L++ELI+N KTIDGRGA VHI A IT D
Sbjct: 124 DEPLWIIFARDMVIKLKEELIMNSFKTIDGRGASVHIAGGACITVQYVTNIIIHGVNIHD 183
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K +RDS HYG RT +VW+DH S+S+C+DGLID ++GSTAIT
Sbjct: 184 CKRKGNAYVRDSPSHYGWRTASDGDAVSIFGGSHVWVDHCSLSNCADGLIDAIHGSTAIT 243
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ + H+ VML G S+ +D MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 244 ISNNYLSHHNKVMLLGHSDSYTRDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 303
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNRF+AP+D KEVTK E P +K+W W+S+GD+ +NGA
Sbjct: 304 HWQMYAIGGSAAPTINSQGNRFLAPNDHVFKEVTKYEDAPRSKWKKWNWRSEGDLFLNGA 363
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SGG Y + + A+ V +T AG L C G C
Sbjct: 364 FFTPSGGGASSSYAKASSLSARPSSLVASVTSNAGALFCRKGSRC 408
>sw|Q9LTZ0|PL11_ARATH Putative pectate lyase 11 precursor (EC
4.2.2.2).
Length = 409
Score = 299 bits (765), Expect = 6e-80
Identities = 146/285 (51%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
+ PLWI+F RDMVI L++ELI+ KTIDGRG+ VHI + D
Sbjct: 125 EEPLWIIFKRDMVIRLKKELIITSFKTIDGRGSSVHITDGPCLKIHYATNIIIHGINIHD 184
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K SGGMI+D H G + +VWIDH S+S+C DGLID ++GSTAIT
Sbjct: 185 CKPGSGGMIKDGPHHTGWWMQSDGDAVAIFGGKHVWIDHCSLSNCDDGLIDAIHGSTAIT 244
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN+H T HD VML G S+ QD MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 245 ISNNHMTHHDKVMLLGHSDSYTQDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 304
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNRF+AP+ KEVTK E P +++W W+S+GD+++NGA
Sbjct: 305 HWEMYAIGGSASPTIYSQGNRFLAPNTRFNKEVTKHEDAPESKWRDWNWRSEGDMLLNGA 364
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F ESG + Y R + A+ VG +T AG L C G+ C
Sbjct: 365 YFRESGAEAPSTYARASSLSARPSSLVGSITTTAGTLSCRRGRRC 409
>sw|Q9SVQ6|PL14_ARATH Putative pectate lyase 14 precursor (EC
4.2.2.2).
Length = 418
Score = 299 bits (765), Expect = 6e-80
Identities = 148/285 (51%), Positives = 183/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
D PLWIVF RDMVI L QELI+N KTIDGRG VHI A +T D
Sbjct: 134 DEPLWIVFKRDMVITLSQELIMNSFKTIDGRGVNVHIAGGACLTVQYVTNIIIHGINIHD 193
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K M+R S+ HYG RT ++WIDH S+SSC+DGLID + GSTAIT
Sbjct: 194 CKRTGNAMVRSSESHYGWRTMADGDGISIFGSSHIWIDHNSLSSCADGLIDAIMGSTAIT 253
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ T H+ +L G ++ +D +MQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 254 ISNNYLTHHNEAILLGHTDSYTRDKMMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYT 313
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ NPTI SQGNRF+AP + AKEVTKR ++ W W+SQGD+M+NGA
Sbjct: 314 HWEMYAIGGSANPTINSQGNRFLAPGNRFAKEVTKRVGAGKGEWNNWNWRSQGDLMLNGA 373
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
+F SG Y R + AK VG LT +G LKC +G C
Sbjct: 374 YFTSSGAGASANYARASSLAAKSSSLVGMLTSSSGALKCRIGTLC 418
>sptr|Q9SDW4|Q9SDW4 Pectate lyase 1.
Length = 407
Score = 297 bits (761), Expect = 2e-79
Identities = 147/285 (51%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
+ PLWI+F RDMVI+L++ELI+N +KTIDGRGA VHI IT D
Sbjct: 123 EEPLWIIFKRDMVIQLKEELIMNSHKTIDGRGASVHISGGPCITIQYVTNIIIHGVHIHD 182
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K +RDS HYG RT +VW+DH ++S+C DGLID ++GSTAIT
Sbjct: 183 CKQGGNAYVRDSPGHYGWRTVSDGDGVSIFGGSHVWVDHCTLSNCHDGLIDAIHGSTAIT 242
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++ + HD VML G S++ D MQVT+AFNHFG LVQRMPRCR+G+FHVVNNDYT
Sbjct: 243 ISNNYLSHHDKVMLLGHSDELTSDKSMQVTIAFNHFGEDLVQRMPRCRHGYFHVVNNDYT 302
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ PTI SQGNRF+AP+D AKEVTKRE ++K+W W+S+GD M+NGA
Sbjct: 303 HWEMYAIGGSAAPTINSQGNRFLAPNDRFAKEVTKREDAQESEWKKWNWRSEGDQMLNGA 362
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y + + A+ VG +T AG L C G C
Sbjct: 363 FFTPSGAGASSSYAKASSLGARSSSLVGTITVSAGVLSCKKGSRC 407
>sw|Q9SRH4|PEL7_ARATH Probable pectate lyase 7 precursor (EC 4.2.2.2).
Length = 475
Score = 295 bits (755), Expect = 9e-79
Identities = 148/285 (51%), Positives = 185/285 (64%), Gaps = 3/285 (1%)
Frame = +2
Query: 11 PLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXDSK 187
PLWIVF DM I L QEL++ +KTID RGA VHI + A IT
Sbjct: 191 PLWIVFKHDMSIRLSQELMITSDKTIDARGANVHIAYGAGITMQYVHNIIIHGLHVHHIV 250
Query: 188 AHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITVS 367
SGG+IRDS H+G R N+W+DH+SMS C DGLID + GSTAIT+S
Sbjct: 251 KSSGGLIRDSINHFGHRGEADGDGISIFGATNIWLDHISMSKCQDGLIDAIMGSTAITIS 310
Query: 368 NSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHW 547
NSHFT H+ VML GA N++ D MQVTVA+NHFG+GLVQRMPR R+GF HVVNNDYTHW
Sbjct: 311 NSHFTHHNDVMLLGAQNNNMDDKKMQVTVAYNHFGKGLVQRMPRVRWGFVHVVNNDYTHW 370
Query: 548 IMYAIGGNMNPTIISQGNRFIA-PDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAF 724
+YAIGG+ PTI+S GNRFIA P + +EVTKR+Y ++K W W+S+ DV MN A+
Sbjct: 371 ELYAIGGSQGPTILSHGNRFIAPPHKQHYREVTKRDYASESEWKNWNWRSEKDVFMNNAY 430
Query: 725 FNESGGQNER-KYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
F +SG + + + R I K+G V +LT++AG L C VG+ C
Sbjct: 431 FRQSGNPHFKCSHSRQQMIKPKNGMAVSKLTKYAGALDCRVGKAC 475
>sw|Q93WF1|PL20_ARATH Probable pectate lyase 20 precursor (EC
4.2.2.2).
Length = 417
Score = 295 bits (755), Expect = 9e-79
Identities = 147/285 (51%), Positives = 184/285 (64%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXD 181
+ PLWIVF RDMVI L++ELI+N KTIDGRG VHI A IT D
Sbjct: 133 EEPLWIVFKRDMVITLKEELIMNSFKTIDGRGVNVHIANGACITIQFVTNIIIHGIHIHD 192
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
+ M+R S HYG RT ++WIDH S+S+C+DGLID V STAIT
Sbjct: 193 CRPTGNAMVRSSPSHYGWRTMADGDGISIFGSSHIWIDHNSLSNCADGLIDAVMASTAIT 252
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN++FT H+ VML G S+ +D VMQVT+A+NHFG GL+QRMPRCR+G+FHVVNNDYT
Sbjct: 253 ISNNYFTHHNEVMLLGHSDTYTRDKVMQVTIAYNHFGEGLIQRMPRCRHGYFHVVNNDYT 312
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNR++AP + AKEVTKR+Y ++ W W+S+GD+ +NGA
Sbjct: 313 HWEMYAIGGSASPTINSQGNRYLAPRNRFAKEVTKRDYAGQWQWRHWNWRSEGDLFLNGA 372
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF SG Y R + AK VG +T AG L C G+ C
Sbjct: 373 FFTRSGSGLGASYARASSLAAKSSSLVGVITYNAGALNCRGGRRC 417
>sptr|Q9M505|Q9M505 Pectate lyase.
Length = 398
Score = 291 bits (744), Expect = 2e-77
Identities = 145/285 (50%), Positives = 181/285 (63%), Gaps = 1/285 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
D PLWI+F RDMVI+L+QEL++N KTIDGRGA VHI IT D
Sbjct: 114 DEPLWIIFKRDMVIKLKQELVMNSFKTIDGRGASVHIAGGPCITIHYASNIIIHGLHIHD 173
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAIT 361
K IR+S H G T ++W+DH S+S+C DGLID ++GSTAIT
Sbjct: 174 CKQGGNANIRNSPHHSGWWTVSDGDGVSIFGGRHIWVDHCSLSNCHDGLIDAIHGSTAIT 233
Query: 362 VSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYT 541
+SN+ T HD VML G S+ +D MQVT+AFNHFG GLVQRMPRCR+G+FHVVNNDYT
Sbjct: 234 ISNNFMTHHDKVMLLGHSDSYTEDKNMQVTIAFNHFGEGLVQRMPRCRHGYFHVVNNDYT 293
Query: 542 HWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGA 721
HW MYAIGG+ +PTI SQGNRF+AP+D K VTK E P +++ W W+S+GD+M+NGA
Sbjct: 294 HWEMYAIGGSADPTINSQGNRFLAPNDRFKKAVTKHEDAPESEWRHWNWRSEGDLMLNGA 353
Query: 722 FFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
FF +SG Y R + A+ VG +T +G L C G C
Sbjct: 354 FFLQSGAGASSSYARRSSLSARPSSLVGSITLGSGALGCRKGSRC 398
>sw|Q9LRM5|PEL9_ARATH Putative pectate lyase 9 precursor (EC 4.2.2.2).
Length = 452
Score = 289 bits (739), Expect = 6e-77
Identities = 148/283 (52%), Positives = 184/283 (65%), Gaps = 1/283 (0%)
Frame = +2
Query: 11 PLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMF-AQITXXXXXXXXXXXXXXXDSK 187
PLWI+F RDMVI+L+QELI+N KTID RGA VHI A IT D K
Sbjct: 170 PLWIIFKRDMVIKLKQELIMNSFKTIDARGANVHIANGACITIQNITNVIVHGLHIHDCK 229
Query: 188 AHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNVWIDHVSMSSCSDGLIDVVNGSTAITVS 367
+R S G R ++WIDH S+S+C+DGL+DVVNGSTAIT+S
Sbjct: 230 RTGNVTVRSSPSQAGFRGTADGDAINIFGSSHIWIDHNSLSNCTDGLVDVVNGSTAITIS 289
Query: 368 NSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDYTHW 547
N+HFT HD VML G ++ +D +MQVTVA+NHFG GL+QRMPRCR+G+FHVVNNDYTHW
Sbjct: 290 NNHFTHHDEVMLLGHNDSYTRDKMMQVTVAYNHFGEGLIQRMPRCRHGYFHVVNNDYTHW 349
Query: 548 IMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNGAFF 727
MYAIGG+ NPTI SQGNRF AP + +AKEVTKR T ++ EW W+S+ D+++NGAFF
Sbjct: 350 KMYAIGGSANPTINSQGNRFAAPKNHSAKEVTKRLDTKGNEWMEWNWRSEKDLLVNGAFF 409
Query: 728 NESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
S G+ + +PAK V +T AG L C G+PC
Sbjct: 410 TPS-GEGASGDSQTLSLPAKPASMVDAITASAGALSCRRGKPC 451
>sptr|Q43783|Q43783 Pectate lyase (EC 4.2.2.2) (Fragment).
Length = 398
Score = 280 bits (717), Expect = 2e-74
Identities = 142/286 (49%), Positives = 179/286 (62%), Gaps = 2/286 (0%)
Frame = +2
Query: 5 DRPLWIVFARDMVIELRQELIVNHNKTIDGRGAQVHIMFAQ-ITXXXXXXXXXXXXXXXD 181
+ PLWI+F RD+VI+L++ELI+N +KTIDGRGA VHI IT D
Sbjct: 113 EEPLWIIFKRDIVIQLKEELIMNSHKTIDGRGASVHISGGPCITIQYVTNIIIHGVHIHD 172
Query: 182 SKAHSGGMIRDSKRHYGLRTRXXXXXXXXXXXXNV-WIDHVSMSSCSDGLIDVVNGSTAI 358
K +RDS HYG RT W+DH ++ +C DGLID ++GSTAI
Sbjct: 173 CKQGGNAYVRDSPGHYGWRTVSDGDGVSIFGGQPPSWVDHCTLFNCHDGLIDAIHGSTAI 232
Query: 359 TVSNSHFTDHDHVMLFGASNDSPQDAVMQVTVAFNHFGRGLVQRMPRCRYGFFHVVNNDY 538
T+SN++ HD VML G S++ D MQVT+AFNHFG LVQRMPRCR+G+FHVVNNDY
Sbjct: 233 TISNNYLRHHDKVMLLGHSDELTSDKSMQVTIAFNHFGEDLVQRMPRCRHGYFHVVNNDY 292
Query: 539 THWIMYAIGGNMNPTIISQGNRFIAPDDPNAKEVTKREYTPYKDYKEWVWKSQGDVMMNG 718
THW MYAIGG+ PTI SQGNRF+AP+D AKEVTKRE ++K+W W+S+GD M+NG
Sbjct: 293 THWEMYAIGGSAAPTINSQGNRFLAPNDRFAKEVTKREDAQESEWKKWNWRSEGDQMLNG 352
Query: 719 AFFNESGGQNERKYDRFDFIPAKHGRYVGQLTRFAGPLKCIVGQPC 856
AFF SG + + + + VG +T AG L C G C
Sbjct: 353 AFFTPSGAGASSSHAKASSLGPRSSSLVGTITVSAGVLSCKKGSRC 398
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 677,361,405
Number of Sequences: 1395590
Number of extensions: 13920834
Number of successful extensions: 42025
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 39757
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 41840
length of database: 442,889,342
effective HSP length: 125
effective length of database: 268,440,592
effective search space used: 69526113328
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

