BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
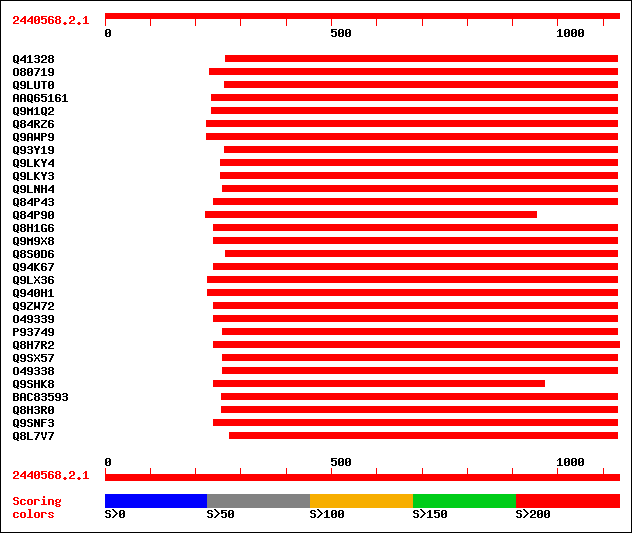
Query= 2440568.2.1
(1135 letters)
Database: /db/trembl-ebi/tmp/swall
1,302,759 sequences; 415,658,970 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q41328|Q41328 Pto kinase interactor 1. 504 e-142
sptr|O80719|O80719 Putative PTO kinase (Hypothetical protein). 501 e-141
sptr|Q9LUT0|Q9LUT0 Serine/threonine protein kinase-like protein ... 495 e-139
sptrnew|AAQ65161|AAQ65161 At3g62220. 490 e-137
sptr|Q9M1Q2|Q9M1Q2 Serine/threonine protein kinase-like protein. 490 e-137
sptr|Q84RZ6|Q84RZ6 OSJNBa0011P19.1 protein. 484 e-135
sptr|Q9AWP9|Q9AWP9 Putative Pto kinase interactor 1. 484 e-135
sptr|Q93Y19|Q93Y19 Similar to Pto kinase interactor 1 gb|AAC6180... 470 e-131
sptr|Q9LKY4|Q9LKY4 Pti1 kinase-like protein. 468 e-131
sptr|Q9LKY3|Q9LKY3 Pti1 kinase-like protein. 467 e-130
sptr|Q9LNH4|Q9LNH4 F21D18.6. 456 e-127
sptr|Q84P43|Q84P43 Protein kinase Pti1. 455 e-127
sptr|Q84P90|Q84P90 Hypothetical protein (Fragment). 453 e-126
sptr|Q8H1G6|Q8H1G6 Putative kinase interactor. 452 e-126
sptr|Q9M9X8|Q9M9X8 Putative protein kinase. 452 e-126
sptr|Q8S0D6|Q8S0D6 Hypothetical protein. 452 e-126
sptr|Q94K67|Q94K67 Hypothetical protein. 451 e-126
sptr|Q9LX36|Q9LX36 Protein kinase-like protein. 450 e-125
sptr|Q940H1|Q940H1 Protein kinase-like protein (At3g59350). 450 e-125
sptr|Q9ZW72|Q9ZW72 Hypothetical protein. 449 e-125
sptr|O49339|O49339 Hypothetical protein (Putative PTI1-like prot... 446 e-124
sptr|P93749|P93749 Hypothetical protein. 428 e-119
sptr|Q8H7R2|Q8H7R2 Putative protein kinase. 420 e-116
sptr|Q9SX57|Q9SX57 F11A17.22 protein. 412 e-114
sptr|O49338|O49338 Hypothetical protein. 408 e-113
sptr|Q9SHK8|Q9SHK8 F12K11.1. 397 e-109
sptrnew|BAC83593|BAC83593 Hypothetical protein P0625E02.106. 274 2e-72
sptr|Q8H3R0|Q8H3R0 Hypothetical protein. 273 4e-72
sptr|Q9SNF3|Q9SNF3 EST AU030604(E51294) corresponds to a region ... 271 2e-71
sptr|Q8L7V7|Q8L7V7 AT4g02010/T10M13_2. 267 2e-70
>sptr|Q41328|Q41328 Pto kinase interactor 1.
Length = 354
Score = 504 bits (1298), Expect = e-142
Identities = 249/289 (86%), Positives = 267/289 (92%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E++++T FG +ALIGEGS+GRVY GVL++GR+AA+KKLDS+KQPD+EFLAQVSMVSRLK
Sbjct: 60 ELKDITDNFGSKALIGEGSYGRVYHGVLKSGRAAAIKKLDSSKQPDREFLAQVSMVSRLK 119
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
ENVVELLGYC DG RVLAYE+A GSLHD+LHGRKGVKGAQPGPVLSW+QRVKIAVGA
Sbjct: 120 DENVVELLGYCVDGGFRVLAYEYAPNGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVGA 179
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
AKGLEYLHEKAQPHIIHRDIKSSN+LLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 180 AKGLEYLHEKAQPHIIHRDIKSSNILLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 239
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV
Sbjct: 240 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 299
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLL 266
+QCVD+RL DYPP ALCVQYEADFRPNMSIVVKALQPLL
Sbjct: 300 KQCVDARLNTDYPPKAIAKMAAVAALCVQYEADFRPNMSIVVKALQPLL 348
>sptr|O80719|O80719 Putative PTO kinase (Hypothetical protein).
Length = 365
Score = 501 bits (1291), Expect = e-141
Identities = 247/301 (82%), Positives = 265/301 (88%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E++E T FG +LIGEGS+GRVY GVL N +A+KKLDSNKQPD EFLAQVSMVSRLK
Sbjct: 65 ELKEATDDFGSNSLIGEGSYGRVYYGVLNNDLPSAIKKLDSNKQPDNEFLAQVSMVSRLK 124
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
H+N V+LLGYC DG R+L+YEFA GSLHD+LHGRKGVKGAQPGPVLSW QRVKIAVGA
Sbjct: 125 HDNFVQLLGYCVDGNSRILSYEFANNGSLHDILHGRKGVKGAQPGPVLSWYQRVKIAVGA 184
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA PHIIHRDIKSSNVLLF+DDVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 185 ARGLEYLHEKANPHIIHRDIKSSNVLLFEDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 244
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQL++KSDVYSFGVVLLELLTGRKPVDH LPRGQQSLVTWATP+LSEDKV
Sbjct: 245 GYHAPEYAMTGQLNAKSDVYSFGVVLLELLTGRKPVDHRLPRGQQSLVTWATPKLSEDKV 304
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNPGE 233
+QCVD+RLGGDYPP ALCVQYEADFRPNMSIVVKALQPLLNA A A G
Sbjct: 305 KQCVDARLGGDYPPKAVAKLAAVAALCVQYEADFRPNMSIVVKALQPLLNARAVAPGEGV 364
Query: 232 H 230
H
Sbjct: 365 H 365
>sptr|Q9LUT0|Q9LUT0 Serine/threonine protein kinase-like protein
(Hypothetical protein).
Length = 364
Score = 495 bits (1275), Expect = e-139
Identities = 240/290 (82%), Positives = 266/290 (91%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+R++T +G ++LIGEGS+GRV+ G+L++G++AA+KKLDS+KQPDQEFLAQVSMVSRL+
Sbjct: 61 ELRDITDNYGSKSLIGEGSYGRVFYGILKSGKAAAIKKLDSSKQPDQEFLAQVSMVSRLR 120
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
ENVV LLGYC DG LRVLAYE+A GSLHD+LHGRKGVKGAQPGPVLSW QRVKIAVGA
Sbjct: 121 QENVVALLGYCVDGPLRVLAYEYAPNGSLHDILHGRKGVKGAQPGPVLSWHQRVKIAVGA 180
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA PH+IHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 181 ARGLEYLHEKANPHVIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 240
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTG LS+KSDVYSFGVVLLELLTGRKPVDHTLPRGQQS+VTWATP+LSEDKV
Sbjct: 241 GYHAPEYAMTGTLSTKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSVVTWATPKLSEDKV 300
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLN 263
+QCVD+RL G+YPP ALCVQYEADFRPNMSIVVKALQPLLN
Sbjct: 301 KQCVDARLNGEYPPKAVAKLAAVAALCVQYEADFRPNMSIVVKALQPLLN 350
>sptrnew|AAQ65161|AAQ65161 At3g62220.
Length = 361
Score = 490 bits (1262), Expect = e-137
Identities = 242/299 (80%), Positives = 262/299 (87%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+ E T FG +LIGEGS+ RVY GVL+NG+ AA+KKLDSNKQP++EFLAQVSMVSRLK
Sbjct: 61 ELIEATNDFGTNSLIGEGSYARVYHGVLKNGQRAAIKKLDSNKQPNEEFLAQVSMVSRLK 120
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
H N VELLGY DG R+L +EFA GSLHD+LHGRKGVKGA+PGP+LSW QRVKIAVGA
Sbjct: 121 HVNFVELLGYSVDGNSRILVFEFAQNGSLHDILHGRKGVKGAKPGPLLSWHQRVKIAVGA 180
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA PH+IHRDIKSSNVL+FD+DVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 181 ARGLEYLHEKANPHVIHRDIKSSNVLIFDNDVAKIADFDLSNQAPDMAARLHSTRVLGTF 240
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLS+KSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 241 GYHAPEYAMTGQLSAKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 300
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNPG 236
+QCVDSRLGGDYPP ALCVQYEADFRPNMSIVVKALQPLLNA G
Sbjct: 301 KQCVDSRLGGDYPPKAVAKLAAVAALCVQYEADFRPNMSIVVKALQPLLNARTGPAGEG 359
>sptr|Q9M1Q2|Q9M1Q2 Serine/threonine protein kinase-like protein.
Length = 361
Score = 490 bits (1262), Expect = e-137
Identities = 242/299 (80%), Positives = 262/299 (87%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+ E T FG +LIGEGS+ RVY GVL+NG+ AA+KKLDSNKQP++EFLAQVSMVSRLK
Sbjct: 61 ELIEATNDFGTNSLIGEGSYARVYHGVLKNGQRAAIKKLDSNKQPNEEFLAQVSMVSRLK 120
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
H N VELLGY DG R+L +EFA GSLHD+LHGRKGVKGA+PGP+LSW QRVKIAVGA
Sbjct: 121 HVNFVELLGYSVDGNSRILVFEFAQNGSLHDILHGRKGVKGAKPGPLLSWHQRVKIAVGA 180
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA PH+IHRDIKSSNVL+FD+DVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 181 ARGLEYLHEKANPHVIHRDIKSSNVLIFDNDVAKIADFDLSNQAPDMAARLHSTRVLGTF 240
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLS+KSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 241 GYHAPEYAMTGQLSAKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 300
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNPG 236
+QCVDSRLGGDYPP ALCVQYEADFRPNMSIVVKALQPLLNA G
Sbjct: 301 KQCVDSRLGGDYPPKAVAKLAAVAALCVQYEADFRPNMSIVVKALQPLLNARTGPAGEG 359
>sptr|Q84RZ6|Q84RZ6 OSJNBa0011P19.1 protein.
Length = 371
Score = 484 bits (1246), Expect = e-135
Identities = 235/308 (76%), Positives = 268/308 (87%), Gaps = 5/308 (1%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
EI+ +TK F +ALIGEGS+ RV+ GVLR+GR +AVKKLDS+KQPDQEFL QVS VSRLK
Sbjct: 63 EIKGITKNFSSDALIGEGSYARVFFGVLRDGRRSAVKKLDSSKQPDQEFLVQVSAVSRLK 122
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HEN+++L+GYCA G++RVLAYE+A GSLHD+LHG+KGVKGAQPGP LSW QRVKIA+ A
Sbjct: 123 HENIIQLIGYCAGGSIRVLAYEYAPRGSLHDILHGKKGVKGAQPGPALSWMQRVKIALSA 182
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
AKGLE+LHEKA+P ++HRDIKSSN++LFD+DVAK+ DFD+SNQ+PDMAARLHSTRVLGTF
Sbjct: 183 AKGLEFLHEKAEPRVVHRDIKSSNIMLFDNDVAKVGDFDVSNQSPDMAARLHSTRVLGTF 242
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLS+KSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV
Sbjct: 243 GYHAPEYAMTGQLSTKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 302
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAH-----ARA 248
+QCVD RL GDYPP ALCVQYEADFRPNMSIVVKAL PLLN+ A
Sbjct: 303 KQCVDPRLEGDYPPKAVAKMAAVAALCVQYEADFRPNMSIVVKALNPLLNSRPNNRPASF 362
Query: 247 TNPGEHAG 224
T+ GE +G
Sbjct: 363 TDAGERSG 370
>sptr|Q9AWP9|Q9AWP9 Putative Pto kinase interactor 1.
Length = 371
Score = 484 bits (1246), Expect = e-135
Identities = 235/308 (76%), Positives = 268/308 (87%), Gaps = 5/308 (1%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
EI+ +TK F +ALIGEGS+ RV+ GVLR+GR +AVKKLDS+KQPDQEFL QVS VSRLK
Sbjct: 63 EIKGITKNFSSDALIGEGSYARVFFGVLRDGRRSAVKKLDSSKQPDQEFLVQVSAVSRLK 122
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HEN+++L+GYCA G++RVLAYE+A GSLHD+LHG+KGVKGAQPGP LSW QRVKIA+ A
Sbjct: 123 HENIIQLIGYCAGGSIRVLAYEYAPRGSLHDILHGKKGVKGAQPGPALSWMQRVKIALSA 182
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
AKGLE+LHEKA+P ++HRDIKSSN++LFD+DVAK+ DFD+SNQ+PDMAARLHSTRVLGTF
Sbjct: 183 AKGLEFLHEKAEPRVVHRDIKSSNIMLFDNDVAKVGDFDVSNQSPDMAARLHSTRVLGTF 242
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLS+KSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV
Sbjct: 243 GYHAPEYAMTGQLSTKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 302
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAH-----ARA 248
+QCVD RL GDYPP ALCVQYEADFRPNMSIVVKAL PLLN+ A
Sbjct: 303 KQCVDPRLEGDYPPKAVAKMAAVAALCVQYEADFRPNMSIVVKALNPLLNSRPNNRPASF 362
Query: 247 TNPGEHAG 224
T+ GE +G
Sbjct: 363 TDAGERSG 370
>sptr|Q93Y19|Q93Y19 Similar to Pto kinase interactor 1 gb|AAC61805.1.
Length = 363
Score = 470 bits (1210), Expect = e-131
Identities = 228/290 (78%), Positives = 256/290 (88%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+R++T +G + LIGEGS+GRV+ GVL++G +A +KKLDS+KQPDQEFL+Q+SMVSRL+
Sbjct: 60 ELRDITDNYGSKTLIGEGSYGRVFYGVLKSGGAADIKKLDSSKQPDQEFLSQISMVSRLR 119
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
H+NV L+GYC DG LRVLAYEFA GSLHD LHG+KG KGA GPV++W QRVKIAVGA
Sbjct: 120 HDNVTALMGYCVDGPLRVLAYEFAPKGSLHDTLHGKKGAKGALRGPVMTWQQRVKIAVGA 179
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEK P +IHRDIKSSNVLLFDDDVAKI DFDLS+QAPDMAARLHSTRVLGTF
Sbjct: 180 ARGLEYLHEKVSPQVIHRDIKSSNVLLFDDDVAKIGDFDLSDQAPDMAARLHSTRVLGTF 239
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTG LSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 240 GYHAPEYAMTGTLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 299
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLN 263
+QCVD+RL G+YPP ALCVQYEA+FRPNMSIVVKALQPLLN
Sbjct: 300 KQCVDARLLGEYPPKAVGKLAAVAALCVQYEANFRPNMSIVVKALQPLLN 349
>sptr|Q9LKY4|Q9LKY4 Pti1 kinase-like protein.
Length = 360
Score = 468 bits (1203), Expect = e-131
Identities = 226/293 (77%), Positives = 255/293 (87%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E++ +T FG + IGEG++G+VY L+NGR+ +KKLDS+ QP+ EFL+QVS+VSRLK
Sbjct: 59 ELKPLTDNFGSKCFIGEGAYGKVYQATLKNGRAVVIKKLDSSNQPEHEFLSQVSIVSRLK 118
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HENVVEL+ YC DG R LAYE+A GSLHD+LHGRKGVKGAQPGPVLSW+QRVKIAVGA
Sbjct: 119 HENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVGA 178
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA+ HIIHR IKSSN+LLFDDDVAKIADFDLSNQAPD AARLHSTRVLGTF
Sbjct: 179 ARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKIADFDLSNQAPDAAARLHSTRVLGTF 238
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQL+SKSDVYSFGV+LLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 239 GYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 298
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHA 254
+QCVD RL G+YP ALCVQYEA+FRPNMSI+VKALQPLLN +
Sbjct: 299 KQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKALQPLLNTRS 351
>sptr|Q9LKY3|Q9LKY3 Pti1 kinase-like protein.
Length = 360
Score = 467 bits (1201), Expect = e-130
Identities = 225/293 (76%), Positives = 255/293 (87%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E++ +T FG + IGEG++G+VY L+NG + +KKLDS+ QP+QEFL+QVS+VSRLK
Sbjct: 59 ELKSLTDNFGSKYFIGEGAYGKVYQATLKNGHAVVIKKLDSSNQPEQEFLSQVSIVSRLK 118
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HENVVEL+ YC DG R LAYE+A GSLHD+LHGRKGVKGAQPGPVLSW+QRVKIAVGA
Sbjct: 119 HENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVGA 178
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEKA+ HIIHR IKSSN+LLFDDDVAK+ADFDLSNQAPD AARLHSTRVLGTF
Sbjct: 179 ARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKVADFDLSNQAPDAAARLHSTRVLGTF 238
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQL+SKSDVYSFGV+LLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 239 GYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 298
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHA 254
+QCVD RL G+YP ALCVQYEA+FRPNMSI+VKALQPLLN +
Sbjct: 299 KQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKALQPLLNTRS 351
>sptr|Q9LNH4|Q9LNH4 F21D18.6.
Length = 732
Score = 456 bits (1174), Expect = e-127
Identities = 220/282 (78%), Positives = 249/282 (88%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+R++T +G + LIGEGS+GRV+ GVL++G +AA+KKLDS+KQPDQEFL+Q+SMVSRL+
Sbjct: 60 ELRDITDNYGSKTLIGEGSYGRVFYGVLKSGGAAAIKKLDSSKQPDQEFLSQISMVSRLR 119
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
H+NV L+GYC DG LRVLAYEFA GSLHD LHG+KG KGA GPV++W QRVKIAVGA
Sbjct: 120 HDNVTALMGYCVDGPLRVLAYEFAPKGSLHDTLHGKKGAKGALRGPVMTWQQRVKIAVGA 179
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
A+GLEYLHEK P +IHRDIKSSNVLLFDDDVAKI DFDLS+QAPDMAARLHSTRVLGTF
Sbjct: 180 ARGLEYLHEKVSPQVIHRDIKSSNVLLFDDDVAKIGDFDLSDQAPDMAARLHSTRVLGTF 239
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTG LSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATP+LSEDKV
Sbjct: 240 GYHAPEYAMTGTLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 299
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVV 287
+QCVD+RL G+YPP ALCVQYEA+FRPNMSIV+
Sbjct: 300 KQCVDARLLGEYPPKAVGKLAAVAALCVQYEANFRPNMSIVL 341
Score = 412 bits (1060), Expect = e-114
Identities = 200/292 (68%), Positives = 242/292 (82%), Gaps = 1/292 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+ ++T+ F E L+G+GS+GRV+ GVL++G+ AA+KKL KQPDQEFL+QVSMVSRL
Sbjct: 428 ELEDITENFSSEVLVGKGSYGRVFYGVLKSGKEAAIKKLYPTKQPDQEFLSQVSMVSRLH 487
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HENVV L+ YC DG LRVLAYEFAT G+LHD+LHG+ GV GA GPV++W +RVKIA+GA
Sbjct: 488 HENVVALMAYCVDGPLRVLAYEFATYGTLHDVLHGQTGVIGALQGPVMTWQRRVKIALGA 547
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRV-LGT 596
A+GLEYLH+K P +IHRDIK+SN+LLFDDD+AKI DFDL +QAP+MA RLHS R+ LG
Sbjct: 548 ARGLEYLHKKVNPQVIHRDIKASNILLFDDDIAKIGDFDLYDQAPNMAGRLHSCRMALGA 607
Query: 595 FGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDK 416
H PE+AMTG L++KSDVYSFGVVLLELLTGRKPVD TLPRGQQ+LVTWATP+LS+DK
Sbjct: 608 SRSHCPEHAMTGILTTKSDVYSFGVVLLELLTGRKPVDRTLPRGQQNLVTWATPKLSKDK 667
Query: 415 VRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNA 260
V+QCVD+RL G+YPP A CV Y+ DFRP+MSIVVKALQPLLN+
Sbjct: 668 VKQCVDARLLGEYPPKAVAKLAAVSARCVHYDPDFRPDMSIVVKALQPLLNS 719
>sptr|Q84P43|Q84P43 Protein kinase Pti1.
Length = 366
Score = 455 bits (1171), Expect = e-127
Identities = 227/300 (75%), Positives = 254/300 (84%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQ--EFLAQVSMVSR 959
E++E T FG +ALIGEGS+GRVY L NG++ AVKKLD + +P+ EFL QVSMVSR
Sbjct: 63 ELKEKTDNFGSKALIGEGSYGRVYYATLNNGKAVAVKKLDVSSEPESNNEFLTQVSMVSR 122
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LK +N VE+ GYC +G LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W QRV+IAV
Sbjct: 123 LKDDNFVEMHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 182
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK QP IIHRDI+SSNVL+F+D AKIADF+LSNQAPDMAARLHSTRVLG
Sbjct: 183 DAARGLEYLHEKVQPPIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLG 242
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 243 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 302
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QCVD +L G+YPP ALCVQYEA+FRPNMSIVVKALQPLL + A A P
Sbjct: 303 KVKQCVDPKLKGEYPPKGVAKLGAVAALCVQYEAEFRPNMSIVVKALQPLLKSPAPAPAP 362
>sptr|Q84P90|Q84P90 Hypothetical protein (Fragment).
Length = 242
Score = 453 bits (1165), Expect = e-126
Identities = 227/244 (93%), Positives = 230/244 (94%)
Frame = -1
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HE+VVELLGYC DG LRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSW+QRVKIAVGA
Sbjct: 1 HEHVVELLGYCVDGNLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWAQRVKIAVGA 60
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 593
AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF
Sbjct: 61 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGTF 120
Query: 592 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 413
GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV
Sbjct: 121 GYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDKV 180
Query: 412 RQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNPGE 233
RQCVDSRLGGDYPP ALCVQYEADFRPNMSIVVKALQPLLN ARATNPGE
Sbjct: 181 RQCVDSRLGGDYPPKAVAKFAAVAALCVQYEADFRPNMSIVVKALQPLLN--ARATNPGE 238
Query: 232 HAGS 221
+AGS
Sbjct: 239 NAGS 242
>sptr|Q8H1G6|Q8H1G6 Putative kinase interactor.
Length = 361
Score = 452 bits (1164), Expect = e-126
Identities = 225/300 (75%), Positives = 253/300 (84%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E++E T+ FG +ALIGEGS+GRVY L +G + A+KKLD + D EFL+QVSMVSR
Sbjct: 60 EVKEKTENFGSKALIGEGSYGRVYYATLNDGVAVALKKLDVAPEAETDTEFLSQVSMVSR 119
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKHEN+++LLG+C DG LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W RVKIAV
Sbjct: 120 LKHENLIQLLGFCVDGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWITRVKIAV 179
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK+QP +IHRDI+SSNVLLF+D AKIADF+LSNQAPD AARLHSTRVLG
Sbjct: 180 EAARGLEYLHEKSQPPVIHRDIRSSNVLLFEDYKAKIADFNLSNQAPDNAARLHSTRVLG 239
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 240 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 299
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QC+D +L DYPP ALCVQYEA+FRPNMSIVVKALQPLL A A P
Sbjct: 300 KVKQCIDPKLKADYPPKAVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLKPPAAAPAP 359
>sptr|Q9M9X8|Q9M9X8 Putative protein kinase.
Length = 377
Score = 452 bits (1164), Expect = e-126
Identities = 225/300 (75%), Positives = 253/300 (84%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E++E T+ FG +ALIGEGS+GRVY L +G + A+KKLD + D EFL+QVSMVSR
Sbjct: 76 EVKEKTENFGSKALIGEGSYGRVYYATLNDGVAVALKKLDVAPEAETDTEFLSQVSMVSR 135
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKHEN+++LLG+C DG LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W RVKIAV
Sbjct: 136 LKHENLIQLLGFCVDGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWITRVKIAV 195
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK+QP +IHRDI+SSNVLLF+D AKIADF+LSNQAPD AARLHSTRVLG
Sbjct: 196 EAARGLEYLHEKSQPPVIHRDIRSSNVLLFEDYKAKIADFNLSNQAPDNAARLHSTRVLG 255
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 256 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 315
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QC+D +L DYPP ALCVQYEA+FRPNMSIVVKALQPLL A A P
Sbjct: 316 KVKQCIDPKLKADYPPKAVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLKPPAAAPAP 375
>sptr|Q8S0D6|Q8S0D6 Hypothetical protein.
Length = 359
Score = 452 bits (1162), Expect = e-126
Identities = 221/290 (76%), Positives = 249/290 (85%), Gaps = 1/290 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNK-QPDQEFLAQVSMVSRL 956
E+ E T FG ALIGEGS+GRVY VL +G AVKKLDSN+ +P EFL QV++VSRL
Sbjct: 65 ELVEKTDDFGSSALIGEGSYGRVYYAVLDSGTKIAVKKLDSNENEPTSEFLTQVALVSRL 124
Query: 955 KHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVG 776
KHEN V++LGYC +G LR++AYEFATMGSLHD+LHGRKGV+GAQPGP L W QRV+IAV
Sbjct: 125 KHENFVDMLGYCTEGNLRLVAYEFATMGSLHDVLHGRKGVQGAQPGPALDWMQRVRIAVD 184
Query: 775 AAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGT 596
AAKGLEYLHEK QP I+HRDI+SSN+LLF+D AK+ADF+LSNQAPDMAARLHSTRVLGT
Sbjct: 185 AAKGLEYLHEKVQPSIVHRDIRSSNILLFEDFKAKVADFNLSNQAPDMAARLHSTRVLGT 244
Query: 595 FGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDK 416
FGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRL+EDK
Sbjct: 245 FGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLTEDK 304
Query: 415 VRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLL 266
V+QC+D RL G+YPP ALCVQYEA+FRPNMSIVVKAL PLL
Sbjct: 305 VKQCIDPRLNGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALSPLL 354
>sptr|Q94K67|Q94K67 Hypothetical protein.
Length = 361
Score = 451 bits (1160), Expect = e-126
Identities = 224/300 (74%), Positives = 252/300 (84%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E++E T+ FG +ALIGEGS+GRVY L +G + A+KKLD + D EFL+QVSMVSR
Sbjct: 60 EVKEKTENFGSKALIGEGSYGRVYYATLNDGVAVALKKLDVAPEAETDTEFLSQVSMVSR 119
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKHEN+++LLG+C DG LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W RVKIAV
Sbjct: 120 LKHENLIQLLGFCVDGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWITRVKIAV 179
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK+QP +IHRDI+SSNVLLF+D AKIADF+LSNQAPD AARLHSTRVLG
Sbjct: 180 EAARGLEYLHEKSQPPVIHRDIRSSNVLLFEDYKAKIADFNLSNQAPDNAARLHSTRVLG 239
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 240 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 299
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QC+D +L DYPP ALCVQYEA+FRPNMSIV KALQPLL A A P
Sbjct: 300 KVKQCIDPKLKADYPPKAVAKLAAVAALCVQYEAEFRPNMSIVAKALQPLLKPPAAAPAP 359
>sptr|Q9LX36|Q9LX36 Protein kinase-like protein.
Length = 403
Score = 450 bits (1158), Expect = e-125
Identities = 220/304 (72%), Positives = 254/304 (83%), Gaps = 2/304 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQ--EFLAQVSMVSR 959
E++E T FG ++LIGEGS+GR Y L++G++ AVKKLD+ +P+ EFL QVS VS+
Sbjct: 100 ELKEKTDNFGSKSLIGEGSYGRAYYATLKDGKAVAVKKLDNAAEPESNVEFLTQVSRVSK 159
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKH+N VEL GYC +G R+LAYEFATMGSLHD+LHGRKGV+GAQPGP L W QRV+IAV
Sbjct: 160 LKHDNFVELFGYCVEGNFRILAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 219
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK QP +IHRDI+SSNVLLF+D AKIADF+LSNQ+PDMAARLHSTRVLG
Sbjct: 220 DAARGLEYLHEKVQPAVIHRDIRSSNVLLFEDFKAKIADFNLSNQSPDMAARLHSTRVLG 279
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 280 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 339
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QCVD +L G+YPP ALCVQYE++FRPNMSIVVKALQPLL + A P
Sbjct: 340 KVKQCVDPKLKGEYPPKAVAKLAAVAALCVQYESEFRPNMSIVVKALQPLLRSSTAAAVP 399
Query: 238 GEHA 227
+ A
Sbjct: 400 VQEA 403
>sptr|Q940H1|Q940H1 Protein kinase-like protein (At3g59350).
Length = 366
Score = 450 bits (1158), Expect = e-125
Identities = 220/304 (72%), Positives = 254/304 (83%), Gaps = 2/304 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQ--EFLAQVSMVSR 959
E++E T FG ++LIGEGS+GR Y L++G++ AVKKLD+ +P+ EFL QVS VS+
Sbjct: 63 ELKEKTDNFGSKSLIGEGSYGRAYYATLKDGKAVAVKKLDNAAEPESNVEFLTQVSRVSK 122
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKH+N VEL GYC +G R+LAYEFATMGSLHD+LHGRKGV+GAQPGP L W QRV+IAV
Sbjct: 123 LKHDNFVELFGYCVEGNFRILAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 182
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK QP +IHRDI+SSNVLLF+D AKIADF+LSNQ+PDMAARLHSTRVLG
Sbjct: 183 DAARGLEYLHEKVQPAVIHRDIRSSNVLLFEDFKAKIADFNLSNQSPDMAARLHSTRVLG 242
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 243 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 302
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QCVD +L G+YPP ALCVQYE++FRPNMSIVVKALQPLL + A P
Sbjct: 303 KVKQCVDPKLKGEYPPKAVAKLAAVAALCVQYESEFRPNMSIVVKALQPLLRSSTAAAVP 362
Query: 238 GEHA 227
+ A
Sbjct: 363 VQEA 366
>sptr|Q9ZW72|Q9ZW72 Hypothetical protein.
Length = 406
Score = 449 bits (1154), Expect = e-125
Identities = 222/300 (74%), Positives = 252/300 (84%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQ--EFLAQVSMVSR 959
E++E T+ FG +ALIGEGS+GRVY +G++ AVKKLD+ +P+ EFL QVS VSR
Sbjct: 103 ELKEKTQNFGSKALIGEGSYGRVYYANFNDGKAVAVKKLDNASEPETNVEFLTQVSKVSR 162
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LK +N V+LLGYC +G LRVLAYEFATM SLHD+LHGRKGV+GAQPGP L W QRV++AV
Sbjct: 163 LKSDNFVQLLGYCVEGNLRVLAYEFATMRSLHDILHGRKGVQGAQPGPTLEWMQRVRVAV 222
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AAKGLEYLHEK QP +IHRDI+SSNVL+F+D AKIADF+LSNQAPDMAARLHSTRVLG
Sbjct: 223 DAAKGLEYLHEKVQPAVIHRDIRSSNVLIFEDFKAKIADFNLSNQAPDMAARLHSTRVLG 282
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 283 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 342
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QCVD +L G+YPP ALCVQYEA+FRPNMSIVVKALQPLL + A P
Sbjct: 343 KVKQCVDPKLKGEYPPKAVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLRSATAAAPP 402
>sptr|O49339|O49339 Hypothetical protein (Putative PTI1-like protein
tyrosine kinase) (EC 2.7.1.112).
Length = 366
Score = 446 bits (1146), Expect = e-124
Identities = 221/300 (73%), Positives = 251/300 (83%), Gaps = 2/300 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E++E T FG ++LIGEGS+GRVY L +G++ A+KKLD + + EFL QVSMVSR
Sbjct: 63 EVKEKTDNFGSKSLIGEGSYGRVYYATLNDGKAVALKKLDVAPEAETNTEFLNQVSMVSR 122
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKHEN+++L+GYC D LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W RVKIAV
Sbjct: 123 LKHENLIQLVGYCVDENLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWLTRVKIAV 182
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK QP +IHRDI+SSNVLLF+D AK+ADF+LSNQAPD AARLHSTRVLG
Sbjct: 183 EAARGLEYLHEKVQPPVIHRDIRSSNVLLFEDYQAKVADFNLSNQAPDNAARLHSTRVLG 242
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPRLSED
Sbjct: 243 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 302
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATNP 239
KV+QCVD +L G+YPP ALCVQYE++FRPNMSIVVKALQPLL A A P
Sbjct: 303 KVKQCVDPKLKGEYPPKSVAKLAAVAALCVQYESEFRPNMSIVVKALQPLLKPPAPAPAP 362
>sptr|P93749|P93749 Hypothetical protein.
Length = 365
Score = 428 bits (1101), Expect = e-119
Identities = 206/293 (70%), Positives = 250/293 (85%), Gaps = 2/293 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E+ + FG++ALIGEGS+GRV+ G + G + A+KKLD S+++PD +F +Q+S+VSR
Sbjct: 65 ELNRMAGNFGNKALIGEGSYGRVFCGKFK-GEAVAIKKLDASSSEEPDSDFTSQLSVVSR 123
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKH++ VELLGYC + R+L Y+FAT GSLHD+LHGRKGV+GA+PGPVL+W+QRVKIA
Sbjct: 124 LKHDHFVELLGYCLEANNRILIYQFATKGSLHDVLHGRKGVQGAEPGPVLNWNQRVKIAY 183
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
GAAKGLE+LHEK QP I+HRD++SSNVLLFDD VAK+ADF+L+N + D AARLHSTRVLG
Sbjct: 184 GAAKGLEFLHEKVQPPIVHRDVRSSNVLLFDDFVAKMADFNLTNASSDTAARLHSTRVLG 243
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
TFGYHAPEYAMTGQ++ KSDVYSFGVVLLELLTGRKPVDHT+P+GQQSLVTWATPRLSED
Sbjct: 244 TFGYHAPEYAMTGQITQKSDVYSFGVVLLELLTGRKPVDHTMPKGQQSLVTWATPRLSED 303
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNA 260
KV+QC+D +L D+PP ALCVQYEADFRPNM+IVVKALQPLLN+
Sbjct: 304 KVKQCIDPKLNNDFPPKAVAKLAAVAALCVQYEADFRPNMTIVVKALQPLLNS 356
>sptr|Q8H7R2|Q8H7R2 Putative protein kinase.
Length = 364
Score = 420 bits (1079), Expect = e-116
Identities = 207/301 (68%), Positives = 241/301 (80%), Gaps = 2/301 (0%)
Frame = -1
Query: 1135 GEIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQE--FLAQVSMVS 962
GE+ +T FG ALIGEGS+GR+Y VL +G A+KKLD + D E F AQ+SMVS
Sbjct: 64 GELNNITGHFGQSALIGEGSYGRIYRAVLTSGEPVAIKKLDPSVSSDSEADFSAQLSMVS 123
Query: 961 RLKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIA 782
RLK+E + L+GY D R+L Y+FAT GSLHD+LHG+KGV+ A PGP L+WSQRVK+A
Sbjct: 124 RLKNEYFIRLMGYYLDANRRILVYQFATHGSLHDILHGKKGVRDAAPGPALNWSQRVKVA 183
Query: 781 VGAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVL 602
GAA+GLEYLHEKAQP I+HRD++SSNVLLFD +K+ADF+L+ Q PD AARLHSTRVL
Sbjct: 184 YGAARGLEYLHEKAQPPIVHRDVRSSNVLLFDGYESKLADFNLTTQPPDGAARLHSTRVL 243
Query: 601 GTFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSE 422
GTFGYHAPEYAMTGQL+ KSDVYSFGV+LLELLTGRKPVDHT+P+ QQSLVTWATPRLSE
Sbjct: 244 GTFGYHAPEYAMTGQLNQKSDVYSFGVILLELLTGRKPVDHTMPKRQQSLVTWATPRLSE 303
Query: 421 DKVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARATN 242
DKVRQCVD +LG DYPP ALCVQYE+DFRPNM+IVVKALQPLL+ A A
Sbjct: 304 DKVRQCVDPKLGDDYPPKAVAKMAAVAALCVQYESDFRPNMTIVVKALQPLLSKPAGAGG 363
Query: 241 P 239
P
Sbjct: 364 P 364
>sptr|Q9SX57|Q9SX57 F11A17.22 protein.
Length = 364
Score = 412 bits (1060), Expect = e-114
Identities = 200/292 (68%), Positives = 242/292 (82%), Gaps = 1/292 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNKQPDQEFLAQVSMVSRLK 953
E+ ++T+ F E L+G+GS+GRV+ GVL++G+ AA+KKL KQPDQEFL+QVSMVSRL
Sbjct: 60 ELEDITENFSSEVLVGKGSYGRVFYGVLKSGKEAAIKKLYPTKQPDQEFLSQVSMVSRLH 119
Query: 952 HENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVGA 773
HENVV L+ YC DG LRVLAYEFAT G+LHD+LHG+ GV GA GPV++W +RVKIA+GA
Sbjct: 120 HENVVALMAYCVDGPLRVLAYEFATYGTLHDVLHGQTGVIGALQGPVMTWQRRVKIALGA 179
Query: 772 AKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRV-LGT 596
A+GLEYLH+K P +IHRDIK+SN+LLFDDD+AKI DFDL +QAP+MA RLHS R+ LG
Sbjct: 180 ARGLEYLHKKVNPQVIHRDIKASNILLFDDDIAKIGDFDLYDQAPNMAGRLHSCRMALGA 239
Query: 595 FGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSEDK 416
H PE+AMTG L++KSDVYSFGVVLLELLTGRKPVD TLPRGQQ+LVTWATP+LS+DK
Sbjct: 240 SRSHCPEHAMTGILTTKSDVYSFGVVLLELLTGRKPVDRTLPRGQQNLVTWATPKLSKDK 299
Query: 415 VRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNA 260
V+QCVD+RL G+YPP A CV Y+ DFRP+MSIVVKALQPLLN+
Sbjct: 300 VKQCVDARLLGEYPPKAVAKLAAVSARCVHYDPDFRPDMSIVVKALQPLLNS 351
>sptr|O49338|O49338 Hypothetical protein.
Length = 338
Score = 408 bits (1049), Expect = e-113
Identities = 203/293 (69%), Positives = 237/293 (80%), Gaps = 2/293 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLD--SNKQPDQEFLAQVSMVSR 959
E+ E T FG +LIGEGS+GRVY L +G++ A+KKLD + + EFL+QVSMVSR
Sbjct: 39 EVNEQTDNFGPNSLIGEGSYGRVYYATLNDGKAVALKKLDLAPEDETNTEFLSQVSMVSR 98
Query: 958 LKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAV 779
LKHEN+++L+GYC D LRVLAYEFATMGSLHD+LHGRKGV+ A PGP L W RVKIAV
Sbjct: 99 LKHENLIQLVGYCVDENLRVLAYEFATMGSLHDILHGRKGVQDALPGPTLDWITRVKIAV 158
Query: 778 GAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLG 599
AA+GLEYLHEK QP +IHRDI+SSN+LLFDD AKIADF+LSNQ+PD AARL STRVLG
Sbjct: 159 EAARGLEYLHEKVQPQVIHRDIRSSNILLFDDYQAKIADFNLSNQSPDNAARLQSTRVLG 218
Query: 598 TFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSED 419
+FGY++PEYAMTG+L+ KSDVY FGVVLLELLTGRKPVDHT+PRGQQSLVTWATP+LSED
Sbjct: 219 SFGYYSPEYAMTGELTHKSDVYGFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPKLSED 278
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNA 260
V +CVD +L G+Y P ALCVQYE++ RP MS VVKALQ LL A
Sbjct: 279 TVEECVDPKLKGEYSPKSVAKLAAVAALCVQYESNCRPKMSTVVKALQQLLIA 331
>sptr|Q9SHK8|Q9SHK8 F12K11.1.
Length = 246
Score = 397 bits (1021), Expect = e-109
Identities = 195/244 (79%), Positives = 213/244 (87%)
Frame = -1
Query: 970 MVSRLKHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRV 791
MVSRLKHEN+++LLG+C DG LRVLAYEFATMGSLHD+LHGRKGV+GAQPGP L W RV
Sbjct: 1 MVSRLKHENLIQLLGFCVDGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWITRV 60
Query: 790 KIAVGAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHST 611
KIAV AA+GLEYLHEK+QP +IHRDI+SSNVLLF+D AKIADF+LSNQAPD AARLHST
Sbjct: 61 KIAVEAARGLEYLHEKSQPPVIHRDIRSSNVLLFEDYKAKIADFNLSNQAPDNAARLHST 120
Query: 610 RVLGTFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPR 431
RVLGTFGYHAPEYAMTGQL+ KSDVYSFGVVLLELLTGRKPVDHT+PRGQQSLVTWATPR
Sbjct: 121 RVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPR 180
Query: 430 LSEDKVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHAR 251
LSEDKV+QC+D +L DYPP ALCVQYEA+FRPNMSIVVKALQPLL A
Sbjct: 181 LSEDKVKQCIDPKLKADYPPKAVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLKPPAA 240
Query: 250 ATNP 239
A P
Sbjct: 241 APAP 244
>sptrnew|BAC83593|BAC83593 Hypothetical protein P0625E02.106.
Length = 479
Score = 274 bits (700), Expect = 2e-72
Identities = 150/294 (51%), Positives = 192/294 (65%), Gaps = 2/294 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNK-QPDQEFLAQVSMVSRL 956
E+ TK F + L+GEG FGRVY G + NG+ AVK+LD N Q ++EFL +V M+S L
Sbjct: 71 ELAVATKNFRKDCLLGEGGFGRVYKGQMENGQVIAVKQLDRNGLQGNREFLVEVLMLSLL 130
Query: 955 KHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVG 776
H N+V L+GYCADG R+L YE+ +GSL + LH R G +P L W+ R+KIAVG
Sbjct: 131 HHPNLVRLIGYCADGDQRLLVYEYMLLGSLENHLHDRP--PGKKP---LDWNARMKIAVG 185
Query: 775 AAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGT 596
AAKGLEYLH+KA P +I+RD KSSN+LL +D K++DF L+ P STRV+GT
Sbjct: 186 AAKGLEYLHDKANPPVIYRDFKSSNILLGEDYYPKLSDFGLAKLGPVGDKTHVSTRVMGT 245
Query: 595 FGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSE-D 419
+GY APEYAMTGQL+ KSDVYSFGVV LEL+TGRK +DHT P G+Q+LV WA P +
Sbjct: 246 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDHTQPAGEQNLVAWARPLFRDRR 305
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAH 257
K Q D L G YP ++C+Q A RP ++ +V AL L + H
Sbjct: 306 KFCQMADPSLQGCYPKRGLYQALAVASMCLQENATSRPLIADIVTALSYLASNH 359
>sptr|Q8H3R0|Q8H3R0 Hypothetical protein.
Length = 477
Score = 273 bits (697), Expect = 4e-72
Identities = 149/294 (50%), Positives = 190/294 (64%), Gaps = 2/294 (0%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSNK-QPDQEFLAQVSMVSRL 956
E+ TK F + L+GEG FGRVY G + NG+ AVK+LD N Q ++EFL +V M+S L
Sbjct: 70 ELAVATKNFRKDCLLGEGGFGRVYKGQMENGQVIAVKQLDRNGLQGNREFLVEVLMLSLL 129
Query: 955 KHENVVELLGYCADGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIAVG 776
H N+V L+GYCADG R+L YE+ +GSL + LH G K L W+ R+KIAVG
Sbjct: 130 HHPNLVRLIGYCADGDQRLLVYEYMLLGSLENHLHRPPGKKP------LDWNARMKIAVG 183
Query: 775 AAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVLGT 596
AAKGLEYLH+KA P +I+RD KSSN+LL +D K++DF L+ P STRV+GT
Sbjct: 184 AAKGLEYLHDKANPPVIYRDFKSSNILLGEDYYPKLSDFGLAKLGPVGDKTHVSTRVMGT 243
Query: 595 FGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRLSE-D 419
+GY APEYAMTGQL+ KSDVYSFGVV LEL+TGRK +DHT P G+Q+LV WA P +
Sbjct: 244 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDHTQPAGEQNLVAWARPLFRDRR 303
Query: 418 KVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAH 257
K Q D L G YP ++C+Q A RP ++ +V AL L + H
Sbjct: 304 KFCQMADPSLQGCYPKRGLYQALAVASMCLQENATSRPLIADIVTALSYLASNH 357
>sptr|Q9SNF3|Q9SNF3 EST AU030604(E51294) corresponds to a region of
the predicted gene.
Length = 820
Score = 271 bits (692), Expect = 2e-71
Identities = 148/302 (49%), Positives = 198/302 (65%), Gaps = 4/302 (1%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSN-KQPDQEFLAQVSMVSRL 956
E++E T F +++GEG FGRV+ GVL +G + A+KKL S Q D+EFL +V M+SRL
Sbjct: 471 ELKEATNNFDPSSMLGEGGFGRVFKGVLTDGTAVAIKKLTSGGHQGDKEFLVEVEMLSRL 530
Query: 955 KHENVVELLGYCA--DGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIA 782
H N+V+L+GY + + + +L YE GSL LHG G ++P L W R++IA
Sbjct: 531 HHRNLVKLIGYYSNRESSQNLLCYELVPNGSLEAWLHGTLGA--SRP---LDWDTRMRIA 585
Query: 781 VGAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVL 602
+ AA+GL YLHE +QP +IHRD K+SN+LL DD AK++DF L+ QAP+ STRV+
Sbjct: 586 LDAARGLAYLHEDSQPCVIHRDFKASNILLEDDFHAKVSDFGLAKQAPEGCTNYLSTRVM 645
Query: 601 GTFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRL-S 425
GTFGY APEYAMTG L KSDVYS+GVVLLELLTGR+PVD + P GQ++LVTWA P L
Sbjct: 646 GTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLTGRRPVDMSQPSGQENLVTWARPILRD 705
Query: 424 EDKVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQPLLNAHARAT 245
+D + + D +LGG YP A CV EA RP M VV++L+ + + + +
Sbjct: 706 KDTLEELADPKLGGQYPKDDFVRVCTIAAACVSPEASQRPTMGEVVQSLKMVQRSEFQES 765
Query: 244 NP 239
P
Sbjct: 766 IP 767
>sptr|Q8L7V7|Q8L7V7 AT4g02010/T10M13_2.
Length = 725
Score = 267 bits (683), Expect = 2e-70
Identities = 146/290 (50%), Positives = 191/290 (65%), Gaps = 4/290 (1%)
Frame = -1
Query: 1132 EIREVTKGFGDEALIGEGSFGRVYLGVLRNGRSAAVKKLDSN-KQPDQEFLAQVSMVSRL 956
E++E T F +++GEG FG+VY G+L +G + A+KKL S Q D+EF ++ M+SRL
Sbjct: 372 ELKEATSNFESASILGEGGFGKVYRGILADGTAVAIKKLTSGGPQGDKEFQVEIDMLSRL 431
Query: 955 KHENVVELLGYCA--DGTLRVLAYEFATMGSLHDMLHGRKGVKGAQPGPVLSWSQRVKIA 782
H N+V+L+GY + D + +L YE GSL LHG G+ L W R+KIA
Sbjct: 432 HHRNLVKLVGYYSSRDSSQHLLCYELVPNGSLEAWLHGPLGLNCP-----LDWDTRMKIA 486
Query: 781 VGAAKGLEYLHEKAQPHIIHRDIKSSNVLLFDDDVAKIADFDLSNQAPDMAARLHSTRVL 602
+ AA+GL YLHE +QP +IHRD K+SN+LL ++ AK+ADF L+ QAP+ STRV+
Sbjct: 487 LDAARGLAYLHEDSQPSVIHRDFKASNILLENNFNAKVADFGLAKQAPEGRGNHLSTRVM 546
Query: 601 GTFGYHAPEYAMTGQLSSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLVTWATPRL-S 425
GTFGY APEYAMTG L KSDVYS+GVVLLELLTGRKPVD + P GQ++LVTW P L
Sbjct: 547 GTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLTGRKPVDMSQPSGQENLVTWTRPVLRD 606
Query: 424 EDKVRQCVDSRLGGDYPPXXXXXXXXXXALCVQYEADFRPNMSIVVKALQ 275
+D++ + VDSRL G YP A CV EA RP M VV++L+
Sbjct: 607 KDRLEELVDSRLEGKYPKEDFIRVCTIAAACVAPEASQRPTMGEVVQSLK 656
Database: /db/trembl-ebi/tmp/swall
Posted date: Nov 7, 2003 8:22 PM
Number of letters in database: 415,658,970
Number of sequences in database: 1,302,759
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 862,248,847
Number of Sequences: 1302759
Number of extensions: 18642902
Number of successful extensions: 75897
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 56967
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 61706
length of database: 415,658,970
effective HSP length: 124
effective length of database: 254,116,854
effective search space used: 64291564062
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

