BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
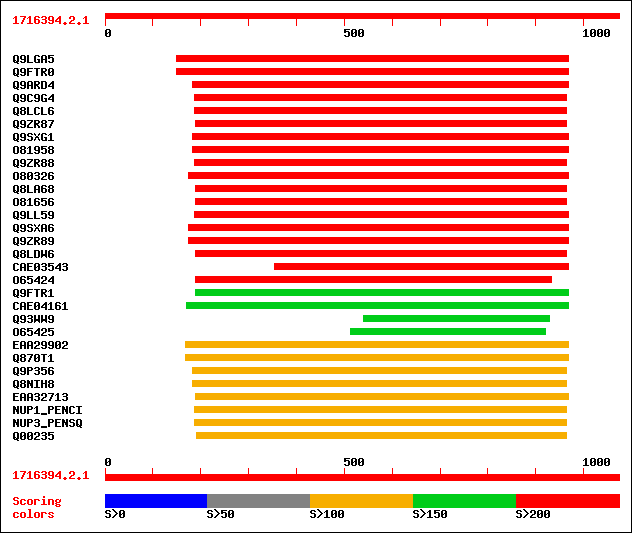
Query= 1716394.2.1
(1074 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9LGA5|Q9LGA5 ESTs D48949(S15541). 508 e-143
sptr|Q9FTR0|Q9FTR0 Putative bifunctional nuclease. 501 e-141
sptr|Q9ARD4|Q9ARD4 Putative nuclease. 335 7e-91
sptr|Q9C9G4|Q9C9G4 Putative bifunctional nuclease. 333 2e-90
sptr|Q8LCL6|Q8LCL6 Putative bifunctional nuclease. 332 4e-90
sptr|Q9ZR87|Q9ZR87 Bifunctional nuclease. 330 2e-89
sptr|Q9SXG1|Q9SXG1 Nuclease I. 330 3e-89
sptr|O81958|O81958 Endonuclease. 325 5e-88
sptr|Q9ZR88|Q9ZR88 Bifunctional nuclease (Fragment). 317 2e-85
sptr|O80326|O80326 Endonuclease precursor. 305 7e-82
sptr|Q8LA68|Q8LA68 Endonuclease, putative. 298 9e-80
sptr|O81656|O81656 Senescence-associated protein 6. 296 3e-79
sptr|Q9LL59|Q9LL59 CEL I mismatch endonuclease. 296 3e-79
sptr|Q9SXA6|Q9SXA6 Bifunctional nuclease bfn1 (Putative bifuncti... 295 6e-79
sptr|Q9ZR89|Q9ZR89 Bifunctional nuclease bfn1. 293 3e-78
sptr|Q8LDW6|Q8LDW6 Putative bifunctional nuclease. 288 1e-76
sptrnew|CAE03543|CAE03543 OSJNBa0060D06.9 protein. 239 4e-62
sptr|O65424|O65424 Hypothetical 41.8 kDa protein (Putative bifun... 232 8e-60
sptr|Q9FTR1|Q9FTR1 Putative bifunctional nuclease. 196 4e-49
sptrnew|CAE04161|CAE04161 OSJNBb0034I13.4 protein. 186 4e-46
sptr|Q93WW9|Q93WW9 S1-type endonuclease (Fragment). 157 2e-37
sptr|O65425|O65425 Hypothetical 51.9 kDa protein (Putative bifun... 153 4e-36
sptrnew|EAA29902|EAA29902 Hypothetical protein. 137 2e-31
sptr|Q870T1|Q870T1 Probable nuclease S1. 137 2e-31
sptr|Q9P356|Q9P356 Nuclease Le1. 130 2e-29
sptr|Q8NIH8|Q8NIH8 Nuclease Le3. 127 3e-28
sptrnew|EAA32713|EAA32713 Hypothetical protein. 115 1e-24
sw|P24289|NUP1_PENCI Nuclease P1 (EC 3.1.30.1) (Endonuclease P1)... 115 1e-24
sw|P24504|NUP3_PENSQ Nuclease PA3 (EC 3.1.3.6) (Endonuclease PA3... 114 3e-24
sptr|Q00235|Q00235 Nuclease S1 precursor. 107 3e-22
>sptr|Q9LGA5|Q9LGA5 ESTs D48949(S15541).
Length = 308
Score = 508 bits (1309), Expect = e-143
Identities = 240/276 (86%), Positives = 257/276 (93%), Gaps = 3/276 (1%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGK+GHI+VCKIAEKYLSEKAAAAV+ LLPESAGGELSTVCPWAD+VR+HY+W+ PLHY
Sbjct: 33 AWGKQGHIIVCKIAEKYLSEKAAAAVEELLPESAGGELSTVCPWADEVRFHYYWSRPLHY 92
Query: 788 ANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQ-KTSYNLTESLMFLAHFV 612
ANTPQVCNFKYSRDCHNSR QQGMCVVGAINNYTDQLYSYG K+SYNLTESLMFLAHFV
Sbjct: 93 ANTPQVCNFKYSRDCHNSRHQQGMCVVGAINNYTDQLYSYGDSKSSYNLTESLMFLAHFV 152
Query: 611 GDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALK 432
GDVHQPLHVGFE+DEGGNTI VHWYRRK NLHHVWD SII+TA+KDFYN+S+DTMVEALK
Sbjct: 153 GDVHQPLHVGFEEDEGGNTIKVHWYRRKENLHHVWDNSIIETAMKDFYNRSLDTMVEALK 212
Query: 431 MNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYP 252
MNLTDGWS+DI HWENC NK TCANDYAIESIH SCNYAYKDVEQDITLGDDYF+SRYP
Sbjct: 213 MNLTDGWSEDISHWENCGNKKETCANDYAIESIHLSCNYAYKDVEQDITLGDDYFYSRYP 272
Query: 251 IVEKRLAQAGIRLALVLNRIFGGGEADD--IPLQVQ 150
IVEKRLAQAGIRLAL+LNRIFG + D IPLQVQ
Sbjct: 273 IVEKRLAQAGIRLALILNRIFGEDKPDGNVIPLQVQ 308
>sptr|Q9FTR0|Q9FTR0 Putative bifunctional nuclease.
Length = 310
Score = 501 bits (1289), Expect = e-141
Identities = 239/278 (85%), Positives = 256/278 (92%), Gaps = 5/278 (1%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGK+GHI+VCKIAEKYLSEKAAAAV+ LLPESAGGELSTVCPWAD+VR+HY+W+ PLHY
Sbjct: 33 AWGKQGHIIVCKIAEKYLSEKAAAAVEELLPESAGGELSTVCPWADEVRFHYYWSRPLHY 92
Query: 788 ANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQ-KTSYNLTESLMFLAHFV 612
ANTPQVCNFKYSRDCHNSR QQGMCVVGAINNYTDQLYSYG K+SYNLTESLMFLAHFV
Sbjct: 93 ANTPQVCNFKYSRDCHNSRHQQGMCVVGAINNYTDQLYSYGDSKSSYNLTESLMFLAHFV 152
Query: 611 GDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALK 432
GDVHQPLHVGFE+DEGGNTI VHWYRRK NLHHVWD SII+TA+KDFYN+S+DTMVEALK
Sbjct: 153 GDVHQPLHVGFEEDEGGNTIKVHWYRRKENLHHVWDNSIIETAMKDFYNRSLDTMVEALK 212
Query: 431 MNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGD--DYFFSR 258
MNLTDGWS+DI HWENC NK TCANDYAIESIH SCNYAYKDVEQDITLG DYF+SR
Sbjct: 213 MNLTDGWSEDISHWENCGNKKETCANDYAIESIHLSCNYAYKDVEQDITLGGEYDYFYSR 272
Query: 257 YPIVEKRLAQAGIRLALVLNRIFGGGEADD--IPLQVQ 150
YPIVEKRLAQAGIRLAL+LNRIFG + D IPLQVQ
Sbjct: 273 YPIVEKRLAQAGIRLALILNRIFGEDKPDGNVIPLQVQ 310
>sptr|Q9ARD4|Q9ARD4 Putative nuclease.
Length = 289
Score = 335 bits (859), Expect = 7e-91
Identities = 152/262 (58%), Positives = 197/262 (75%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGKEGH M CKIA+ +L+++A+A V+ LLP SA GEL+ VC WAD R+ Y W+SPLH+
Sbjct: 23 AWGKEGHYMTCKIADGFLTKEASAGVKDLLPSSANGELAEVCSWADSQRFRYRWSSPLHF 82
Query: 788 ANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFLAHFVG 609
A+TP+ C F Y+RDCH+++G + CVVGAINNYT L YN TESLMFLAHFVG
Sbjct: 83 ADTPKDCKFSYARDCHDTKGNKDACVVGAINNYTAALQD--PSNPYNRTESLMFLAHFVG 140
Query: 608 DVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALKM 429
DVHQPLH G +D GGNTI V WYRRK+NLHHVWDV +I+ A+KD Y + DTM++A++
Sbjct: 141 DVHQPLHCGRVEDLGGNTILVRWYRRKSNLHHVWDVDVIEQAMKDLYGRDQDTMIKAIQR 200
Query: 428 NLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYPI 249
N+T+ WS + WE C+++ TCA+ YA ES +C+ AYK V+QD TLGD+Y+F P+
Sbjct: 201 NITEDWSSEEKQWEACRSRTKTCADKYAEESAVLACD-AYKGVKQDSTLGDEYYFKALPV 259
Query: 248 VEKRLAQAGIRLALVLNRIFGG 183
V+KR+AQ G+RLA +LNRIF G
Sbjct: 260 VQKRIAQGGVRLAAILNRIFSG 281
>sptr|Q9C9G4|Q9C9G4 Putative bifunctional nuclease.
Length = 290
Score = 333 bits (855), Expect = 2e-90
Identities = 151/264 (57%), Positives = 198/264 (75%), Gaps = 4/264 (1%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHYA 786
WGKEGH ++CKIA+ L E AA AV+ LLPESA G+LS++C WAD+V++ YHW+SPLHY
Sbjct: 28 WGKEGHEIICKIAQTRLDETAAKAVKELLPESAEGDLSSLCLWADRVKFRYHWSSPLHYI 87
Query: 785 NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSY----GQKTSYNLTESLMFLAH 618
NTP C+++Y+RDC + G++G CV GAI NYT QL SY ++ YNLTE+L+F++H
Sbjct: 88 NTPDACSYQYNRDCKDESGEKGRCVAGAIYNYTTQLLSYKTAASSQSQYNLTEALLFVSH 147
Query: 617 FVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEA 438
F+GD+HQPLHV + D+GGNTI VHWY RKANLHH+WD +II+TA D YN +++ MV+A
Sbjct: 148 FMGDIHQPLHVSYASDKGGNTIEVHWYTRKANLHHIWDSNIIETAEADLYNSALEGMVDA 207
Query: 437 LKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSR 258
LK N+T W+D + WE C K A C + YA E I +C++AYK V + TL D+YF+SR
Sbjct: 208 LKKNITTEWADQVKRWETCTKKTA-CPDIYASEGIQAACDWAYKGVTEGDTLEDEYFYSR 266
Query: 257 YPIVEKRLAQAGIRLALVLNRIFG 186
PIV +RLAQ G+RLA LNRIFG
Sbjct: 267 LPIVYQRLAQGGVRLAATLNRIFG 290
>sptr|Q8LCL6|Q8LCL6 Putative bifunctional nuclease.
Length = 290
Score = 332 bits (852), Expect = 4e-90
Identities = 150/264 (56%), Positives = 198/264 (75%), Gaps = 4/264 (1%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHYA 786
WGKEGH ++CKIA+ L E AA AV+ LLPESA G+LS++C WAD+V++ YHW+SPLHY
Sbjct: 28 WGKEGHEIICKIAQTRLDETAAKAVKELLPESAEGDLSSLCLWADRVKFRYHWSSPLHYI 87
Query: 785 NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSY----GQKTSYNLTESLMFLAH 618
NTP C+++Y+RDC + G++G CV GAI NYT QL SY ++ YNLTE+L+F++H
Sbjct: 88 NTPDACSYQYNRDCKDESGEKGRCVAGAIYNYTTQLLSYKTAASSQSQYNLTEALLFVSH 147
Query: 617 FVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEA 438
F+GD+HQPLHV + D+GGNTI VHWY RKANLHH+WD +II+TA D YN +++ MV+A
Sbjct: 148 FMGDIHQPLHVSYASDKGGNTIEVHWYTRKANLHHIWDSNIIETAEADLYNSALEGMVDA 207
Query: 437 LKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSR 258
LK N+T W+D + W+ C K A C + YA E I +C++AYK V + TL D+YF+SR
Sbjct: 208 LKKNITTEWADQVKRWDTCTKKTA-CPDIYASEGIQAACDWAYKGVTEGDTLEDEYFYSR 266
Query: 257 YPIVEKRLAQAGIRLALVLNRIFG 186
PIV +RLAQ G+RLA LNRIFG
Sbjct: 267 LPIVYQRLAQGGVRLAATLNRIFG 290
>sptr|Q9ZR87|Q9ZR87 Bifunctional nuclease.
Length = 328
Score = 330 bits (846), Expect = 2e-89
Identities = 151/266 (56%), Positives = 194/266 (72%), Gaps = 7/266 (2%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRW--HYHWASPLH 792
WGKEGH CKIA+ +LSE+A AV+ LLPE+A G+L++VC W D+++W +HW S LH
Sbjct: 27 WGKEGHYATCKIAQSFLSEEALNAVKELLPETAEGDLASVCSWPDEIKWMHKWHWTSELH 86
Query: 791 YANTPQV-CNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYS-YGQKTS---YNLTESLMF 627
Y +TP CN+ Y RDCH+S G + CV GAI NYT+QL + Y S YNLTE+LMF
Sbjct: 87 YVDTPDFRCNYDYCRDCHDSSGVKDRCVTGAIYNYTEQLITGYNASNSVVKYNLTEALMF 146
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
L+H++GDVHQPLHV F DEGGNTI V WY+RK NLHH+WD +I++A+K FY+K +D M
Sbjct: 147 LSHYIGDVHQPLHVSFTSDEGGNTIIVRWYKRKTNLHHIWDTDMIESAMKTFYDKDIDIM 206
Query: 446 VEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYF 267
+ A++ N+TD WS+DI W NC + C + +A ESI YSCNYAY++ TLGD+YF
Sbjct: 207 ISAIEKNITDRWSNDISSWVNCTSGEEVCPDPWASESIKYSCNYAYRNATPGSTLGDEYF 266
Query: 266 FSRYPIVEKRLAQAGIRLALVLNRIF 189
+SR PIVE RLAQ G+RLA LNRIF
Sbjct: 267 YSRLPIVEMRLAQGGVRLAATLNRIF 292
>sptr|Q9SXG1|Q9SXG1 Nuclease I.
Length = 290
Score = 330 bits (845), Expect = 3e-89
Identities = 154/263 (58%), Positives = 197/263 (74%), Gaps = 1/263 (0%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGKEGH M CKIA+ +L+ +A+AAV+ LLP A GEL+ VC WAD+ R+ Y W+SPLH+
Sbjct: 23 AWGKEGHYMTCKIADGFLTSEASAAVKDLLPSWANGELAEVCAWADRQRFRYRWSSPLHF 82
Query: 788 ANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFLAHFVG 609
A+TP CNF Y+RDCH+++G + +CVVGAINNYT L + Y+ TESLMFLAHFVG
Sbjct: 83 ADTPGDCNFSYARDCHDTKGNKDVCVVGAINNYTAALED--PSSPYDPTESLMFLAHFVG 140
Query: 608 DVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALKM 429
DVHQPLH G DD GGNTI + WYRRK+NLHHVWD +I A+KDF+N+ DTM+EA++
Sbjct: 141 DVHQPLHCGHVDDLGGNTIKLRWYRRKSNLHHVWDSDVITQAMKDFFNRDQDTMIEAIQR 200
Query: 428 NLTDGWSDDIVHWENCKNK-HATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYP 252
N+TD WS + WE C ++ TCA YA ES +C+ AY+ VEQ TLGDDY+F P
Sbjct: 201 NITDDWSSEEKQWEACGSRTKITCAEKYAKESALLACD-AYEGVEQGDTLGDDYYFRALP 259
Query: 251 IVEKRLAQAGIRLALVLNRIFGG 183
+VEKR+AQ G+RLA++LN+IF G
Sbjct: 260 VVEKRIAQGGVRLAVILNQIFSG 282
>sptr|O81958|O81958 Endonuclease.
Length = 288
Score = 325 bits (834), Expect = 5e-88
Identities = 148/262 (56%), Positives = 190/262 (72%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGKEGH M CKIA+ +L+ +A V+ALLP A GEL+ VC WAD R+ Y W+ LH+
Sbjct: 23 AWGKEGHYMTCKIADGFLTSEALTGVKALLPSWANGELAEVCSWADSQRFRYRWSRSLHF 82
Query: 788 ANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFLAHFVG 609
A+TP C F Y+RDCH+++G + +CVVGAINNYT L + +N TESLMFLAHFVG
Sbjct: 83 ADTPGDCKFSYARDCHDTKGNKNVCVVGAINNYTAALQD--SSSPFNPTESLMFLAHFVG 140
Query: 608 DVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALKM 429
DVHQP+H G DD GGNTI + WYRRK+NLHHVWD +I +KDF++K D M+E+++
Sbjct: 141 DVHQPMHCGHVDDLGGNTIKLRWYRRKSNLHHVWDSDVITQTMKDFFDKDQDAMIESIQR 200
Query: 428 NLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYPI 249
N+TD WS + WE C++K TCA YA ES +C+ AY+ VEQD TLGD+Y+F P+
Sbjct: 201 NITDDWSSEEKQWETCRSKTTTCAEKYAQESAVLACD-AYEGVEQDDTLGDEYYFKALPV 259
Query: 248 VEKRLAQAGIRLALVLNRIFGG 183
V+KRLAQ G+RLA +LNRIF G
Sbjct: 260 VQKRLAQGGLRLAAILNRIFSG 281
>sptr|Q9ZR88|Q9ZR88 Bifunctional nuclease (Fragment).
Length = 280
Score = 317 bits (812), Expect = 2e-85
Identities = 146/262 (55%), Positives = 187/262 (71%), Gaps = 2/262 (0%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHYA 786
WG +GH + CKIA+ LS+ A AV +LLPE A G+L+++C WAD V++ YHW+S LHY
Sbjct: 19 WGVDGHFITCKIAQGRLSQTAVDAVNSLLPEYAEGDLASLCSWADHVKFRYHWSSALHYI 78
Query: 785 NTPQ-VCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS-YNLTESLMFLAHFV 612
+TP +C ++Y RDC + G G CV GAI NYT QL YG++TS YNLTE+L+FL+HF+
Sbjct: 79 DTPDNLCTYQYRRDCKDEDGVMGRCVAGAIMNYTTQLLDYGKQTSQYNLTEALLFLSHFM 138
Query: 611 GDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALK 432
GD+HQPLHVGF D GGNTI VHW+ RKA LHHVWD SII+TA + FY +++ +++A++
Sbjct: 139 GDIHQPLHVGFTSDRGGNTIDVHWFTRKAVLHHVWDDSIIETAEERFYGSNVENLIDAIE 198
Query: 431 MNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYP 252
N+T+ W D + WENC TC N YA E I +CN+AYK V L DDYF SR P
Sbjct: 199 TNITNVWGDQVKAWENCSANQKTCPNIYATEGIKAACNWAYKGVTNGSVLEDDYFLSRLP 258
Query: 251 IVEKRLAQAGIRLALVLNRIFG 186
IV RLAQ G+RLA LNRIFG
Sbjct: 259 IVNWRLAQGGVRLAANLNRIFG 280
>sptr|O80326|O80326 Endonuclease precursor.
Length = 303
Score = 305 bits (781), Expect = 7e-82
Identities = 143/272 (52%), Positives = 191/272 (70%), Gaps = 7/272 (2%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR--WHYHWASPL 795
AW KEGH+M C+IA++ LS AA AVQ LLP+ G LS +C W DQ+R + Y W SPL
Sbjct: 25 AWSKEGHVMTCQIAQELLSPDAAHAVQMLLPDYVKGNLSALCVWPDQIRHWYRYRWTSPL 84
Query: 794 HYANTPQ-VCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMF 627
H+ +TP C+F Y+RDCH+S G MCV GAI N+T QL Y TS YN+TE+L+F
Sbjct: 85 HFIDTPDDACSFDYTRDCHDSNGMVDMCVAGAIKNFTSQLSHYQHGTSDRRYNMTEALLF 144
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
++HF+GD+HQP+HVGF DEGGNTI + W+R K+NLHHVWD II TA + Y+K M+++
Sbjct: 145 VSHFMGDIHQPMHVGFTTDEGGNTIDLRWFRHKSNLHHVWDREIILTAASELYDKDMESL 204
Query: 446 VEALKMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDY 270
+A++ N T G WSDD+ W++C + + C N YA ESI +C + Y+ VE TL DDY
Sbjct: 205 QKAIQANFTHGLWSDDVNSWKDC-DDISNCVNKYAKESIALACKWGYEGVEAGETLSDDY 263
Query: 269 FFSRYPIVEKRLAQAGIRLALVLNRIFGGGEA 174
F SR PIV KR+AQ G+RL+++LNR+FG +
Sbjct: 264 FDSRMPIVMKRIAQGGVRLSMILNRVFGSSSS 295
>sptr|Q8LA68|Q8LA68 Endonuclease, putative.
Length = 296
Score = 298 bits (763), Expect = 9e-80
Identities = 140/267 (52%), Positives = 184/267 (68%), Gaps = 8/267 (2%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESA-GGELSTVCPWADQVRW--HYHWASPL 795
WGK+GH VCK+AE + + AAV+ LLPES GG L+ C W D+++ + W S L
Sbjct: 21 WGKDGHYTVCKLAEGFFEDDTIAAVKKLLPESVDGGGLADFCSWPDEIKKLSQWQWTSTL 80
Query: 794 HYANTPQV-CNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS----YNLTESLM 630
HY NTP+ CN++Y RDCH++ + CV GAI NYT+QL S + + YNLTE+L+
Sbjct: 81 HYVNTPEYRCNYEYCRDCHDTHKHRDWCVTGAIFNYTNQLMSASENSQNIVHYNLTEALL 140
Query: 629 FLAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDT 450
FL+H++GDVHQPLH GF D GGNTI V+WY K+NLHHVWD IID+A++ +YN S+
Sbjct: 141 FLSHYMGDVHQPLHTGFLGDLGGNTIIVNWYHNKSNLHHVWDNMIIDSALETYYNSSLPH 200
Query: 449 MVEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDY 270
M++AL+ L +GWS+D+ W++C C N YA ESI +C YAY++ TLGD+Y
Sbjct: 201 MIQALQAKLKNGWSNDVPSWKSCHFHQKACPNLYASESIDLACKYAYRNATPGTTLGDEY 260
Query: 269 FFSRYPIVEKRLAQAGIRLALVLNRIF 189
F SR P+VEKRLAQ GIRLA LNRIF
Sbjct: 261 FLSRLPVVEKRLAQGGIRLAATLNRIF 287
>sptr|O81656|O81656 Senescence-associated protein 6.
Length = 298
Score = 296 bits (759), Expect = 3e-79
Identities = 142/266 (53%), Positives = 183/266 (68%), Gaps = 7/266 (2%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPLH 792
W KEGHI+ C+IA+ L +AA V+ LLP G+LS +C W DQ+R W+ Y W+SPLH
Sbjct: 23 WSKEGHIVTCRIAQDLLEPEAAETVRNLLPHYVDGDLSALCTWPDQIRHWYKYRWSSPLH 82
Query: 791 YANTPQ-VCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMFL 624
+ +TP C+F YSRDCH+ +G + MCV GA++NYT QL Y TS YNLTESL+FL
Sbjct: 83 FIDTPDDACSFDYSRDCHDPKGAEDMCVAGAVHNYTTQLMHYRDGTSDRRYNLTESLLFL 142
Query: 623 AHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMV 444
+HF+GD+HQP+HVGF DEGGNTI + W+R K+NLHHVWD II TA+ D+Y K +D
Sbjct: 143 SHFMGDIHQPMHVGFTSDEGGNTINLRWFRHKSNLHHVWDREIILTALADYYGKDLDAFQ 202
Query: 443 EALKMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYF 267
+ L+ N T G WSDD W C + +C +A ESI +C + YK V TL D+YF
Sbjct: 203 QDLQNNFTTGIWSDDTSSWGECDDLF-SCPKKWASESISLACKWGYKGVTPGETLSDEYF 261
Query: 266 FSRYPIVEKRLAQAGIRLALVLNRIF 189
SR PIV KR+AQ G+RLA+VLNR+F
Sbjct: 262 NSRMPIVMKRIAQGGVRLAMVLNRVF 287
>sptr|Q9LL59|Q9LL59 CEL I mismatch endonuclease.
Length = 296
Score = 296 bits (758), Expect = 3e-79
Identities = 140/268 (52%), Positives = 189/268 (70%), Gaps = 7/268 (2%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPL 795
AW KEGH+M C+IA+ L +AA AV+ LLP+ A G LS++C W DQ+R W+ Y W S L
Sbjct: 22 AWSKEGHVMTCQIAQDLLEPEAAHAVKMLLPDYANGNLSSLCVWPDQIRHWYKYRWTSSL 81
Query: 794 HYANTP-QVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMF 627
H+ +TP Q C+F Y RDCH+ G + MCV GAI N+T QL + TS YN+TE+L+F
Sbjct: 82 HFIDTPDQACSFDYQRDCHDPHGGKDMCVAGAIQNFTSQLGHFRHGTSDRRYNMTEALLF 141
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
L+HF+GD+HQP+HVGF D GGN+I + W+R K+NLHHVWD II TA D++ K M ++
Sbjct: 142 LSHFMGDIHQPMHVGFTSDMGGNSIDLRWFRHKSNLHHVWDREIILTAAADYHGKDMHSL 201
Query: 446 VEALKMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDY 270
++ ++ N T+G W D+ W+ C + +TCAN YA ESI +CN+ YKDVE TL D Y
Sbjct: 202 LQDIQRNFTEGSWLQDVESWKEC-DDISTCANKYAKESIKLACNWGYKDVESGETLSDKY 260
Query: 269 FFSRYPIVEKRLAQAGIRLALVLNRIFG 186
F +R PIV KR+AQ GIRL+++LNR+ G
Sbjct: 261 FNTRMPIVMKRIAQGGIRLSMILNRVLG 288
>sptr|Q9SXA6|Q9SXA6 Bifunctional nuclease bfn1 (Putative
bifunctional nuclease bfn1).
Length = 305
Score = 295 bits (756), Expect = 6e-79
Identities = 139/272 (51%), Positives = 189/272 (69%), Gaps = 7/272 (2%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPL 795
+W KEGHI+ C+IA+ L A V+ LLP+ G+LS +C W DQ+R W+ Y W S L
Sbjct: 28 SWSKEGHILTCRIAQNLLEAGPAHVVENLLPDYVKGDLSALCVWPDQIRHWYKYRWTSHL 87
Query: 794 HYANTP-QVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMF 627
HY +TP Q C+++YSRDCH+ G + MCV GAI N+T QL YG+ TS YN+TE+L+F
Sbjct: 88 HYIDTPDQACSYEYSRDCHDQHGLKDMCVDGAIQNFTSQLQHYGEGTSDRRYNMTEALLF 147
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
L+HF+GD+HQP+HVGF DEGGNTI + WY+ K+NLHHVWD II TA+K+ Y+K++D +
Sbjct: 148 LSHFMGDIHQPMHVGFTSDEGGNTIDLRWYKHKSNLHHVWDREIILTALKENYDKNLDLL 207
Query: 446 VEALKMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDY 270
E L+ N+T+G W DD+ W C N C + YA ESI +C + YK V+ TL ++Y
Sbjct: 208 QEDLEKNITNGLWHDDLSSWTEC-NDLIACPHKYASESIKLACKWGYKGVKSGETLSEEY 266
Query: 269 FFSRYPIVEKRLAQAGIRLALVLNRIFGGGEA 174
F +R PIV KR+ Q G+RLA++LNR+F A
Sbjct: 267 FNTRLPIVMKRIVQGGVRLAMILNRVFSDDHA 298
>sptr|Q9ZR89|Q9ZR89 Bifunctional nuclease bfn1.
Length = 305
Score = 293 bits (750), Expect = 3e-78
Identities = 139/272 (51%), Positives = 188/272 (69%), Gaps = 7/272 (2%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPL 795
+W KEGHI+ C+IA+ L A V+ LLP+ G+LS +C W DQ+R W+ Y W S L
Sbjct: 28 SWSKEGHILTCRIAQNLLEAGPAHVVENLLPDYVKGDLSALCVWPDQIRHWYKYRWTSHL 87
Query: 794 HYANTP-QVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMF 627
HY +TP Q C+++YSRDCH+ G + MCV GAI N+T QL YG+ TS YN+TE+L+F
Sbjct: 88 HYIDTPDQACSYEYSRDCHDQHGLKDMCVDGAIQNFTSQLQHYGEGTSDRRYNMTEALLF 147
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
L+HF+GD+HQP+HVGF DEGGNTI + WY+ K+NLHHVWD II TA+K+ Y+K++D +
Sbjct: 148 LSHFMGDIHQPMHVGFTSDEGGNTIDLRWYKHKSNLHHVWDREIILTALKENYDKNLDLL 207
Query: 446 VEALKMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDY 270
E L+ N+T+G W DD+ W C N C + YA ESI +C + YK V+ TL ++Y
Sbjct: 208 QEDLEKNITNGLWHDDLSSWTEC-NDLIACPHKYASESIKLACKWGYKGVKSGETLSEEY 266
Query: 269 FFSRYPIVEKRLAQAGIRLALVLNRIFGGGEA 174
F +R PIV KR+ Q G+RLA++LNR F A
Sbjct: 267 FNTRLPIVMKRIVQGGVRLAMILNRDFSDDHA 298
>sptr|Q8LDW6|Q8LDW6 Putative bifunctional nuclease.
Length = 294
Score = 288 bits (736), Expect = 1e-76
Identities = 138/266 (51%), Positives = 177/266 (66%), Gaps = 7/266 (2%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRW--HYHWASPLH 792
WG GH VCKIA+ Y E AV+ LLPESA GEL+ VC W D+++ + W S LH
Sbjct: 25 WGDAGHYAVCKIAQSYFEEDTVVAVKKLLPESANGELAAVCSWPDEIKKLPQWRWTSALH 84
Query: 791 YANTPQV-CNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS----YNLTESLMF 627
+A+TP CN++YSRDC + CV GAI NYT+QL S + + YNLTE+LMF
Sbjct: 85 FADTPDYKCNYEYSRDC-----PKDWCVTGAIFNYTNQLMSTSENSQSIVHYNLTEALMF 139
Query: 626 LAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTM 447
L+H++GD+HQPLH GF D GGN I VHWY ++ NLH VWD II++A++ +YN S+ M
Sbjct: 140 LSHYMGDIHQPLHEGFIGDLGGNKIKVHWYNQETNLHRVWDDMIIESALETYYNSSLPRM 199
Query: 446 VEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYF 267
+ L+ L +GWS+D+ WE+C+ C N YA ESI +C YAY++ TLGD YF
Sbjct: 200 IHELQAKLKNGWSNDVPSWESCQLNQTACPNPYASESIDLACKYAYRNATAGTTLGDYYF 259
Query: 266 FSRYPIVEKRLAQAGIRLALVLNRIF 189
SR P+VEKRLAQ GIRLA LNRIF
Sbjct: 260 VSRLPVVEKRLAQGGIRLAGTLNRIF 285
>sptrnew|CAE03543|CAE03543 OSJNBa0060D06.9 protein.
Length = 234
Score = 239 bits (611), Expect = 4e-62
Identities = 116/210 (55%), Positives = 149/210 (70%), Gaps = 5/210 (2%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHYHWASPLHY 789
AWGKEGH MVCKIAE +L+++AA AV+ LLP AGGEL+ C WAD R+ + L
Sbjct: 26 AWGKEGHYMVCKIAEGFLTKEAATAVKELLPGWAGGELAETCSWADTERFRGNRLPELSV 85
Query: 788 A----NTPQVCNFKYSR-DCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFL 624
T + + Y DCHN+ G++ MCVVGAINNYT+ L + Y+ TESLMFL
Sbjct: 86 HAMIDKTLRTIMYGYGYGDCHNTNGEKDMCVVGAINNYTNALED--SSSPYDPTESLMFL 143
Query: 623 AHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMV 444
AHFVGDVHQPLH G D GGNTI VHWY RK+NLHHVWDV++I+TA+K+FYN+ + TM+
Sbjct: 144 AHFVGDVHQPLHCGHVKDLGGNTIIVHWYTRKSNLHHVWDVNVIETALKEFYNEDVSTMI 203
Query: 443 EALKMNLTDGWSDDIVHWENCKNKHATCAN 354
+A+KMN+TD WS + WE C+++ TCA+
Sbjct: 204 KAIKMNITDEWSTEEKQWETCRSRTKTCAD 233
>sptr|O65424|O65424 Hypothetical 41.8 kDa protein (Putative
bifunctional nuclease).
Length = 362
Score = 232 bits (591), Expect = 8e-60
Identities = 114/251 (45%), Positives = 151/251 (60%), Gaps = 3/251 (1%)
Frame = -2
Query: 932 IAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRW--HYHWASPLHYANTPQV-CNF 762
+ + Y E AV+ LLPESA GEL+ VC W D+++ + W S LH+A+TP CN+
Sbjct: 136 LRKSYFEEDTVVAVKKLLPESANGELAAVCSWPDEIKKLPQWRWTSALHFADTPDYKCNY 195
Query: 761 KYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFLAHFVGDVHQPLHVG 582
+YS +NLTE+LMFL+H++GD+HQPLH G
Sbjct: 196 EYS---------------------------------HNLTEALMFLSHYMGDIHQPLHEG 222
Query: 581 FEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEALKMNLTDGWSDD 402
F D GGN I VHWY ++ NLH VWD II++A++ +YN S+ M+ L+ L +GWS+D
Sbjct: 223 FIGDLGGNKIKVHWYNQETNLHRVWDDMIIESALETYYNSSLPRMIHELQAKLKNGWSND 282
Query: 401 IVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSRYPIVEKRLAQAG 222
+ WE+C+ C N YA ESI +C YAY++ TLGD YF SR P+VEKRLAQ G
Sbjct: 283 VPSWESCQLNQTACPNPYASESIDLACKYAYRNATAGTTLGDYYFVSRLPVVEKRLAQGG 342
Query: 221 IRLALVLNRIF 189
IRLA LNRIF
Sbjct: 343 IRLAGTLNRIF 353
Score = 116 bits (290), Expect = 6e-25
Identities = 57/106 (53%), Positives = 68/106 (64%), Gaps = 28/106 (26%)
Frame = -2
Query: 653 YNLTESLMFLAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHH------------- 513
YNLTE+LMFL+HF+GD+HQPLHVGF DEGGNTITV WYRRK NLHH
Sbjct: 16 YNLTEALMFLSHFIGDIHQPLHVGFLGDEGGNTITVRWYRRKTNLHHVSVCYRMLKEKVI 75
Query: 512 ---------------VWDVSIIDTAIKDFYNKSMDTMVEALKMNLT 420
VWD II++A+K +YNKS+ M+EAL+ NLT
Sbjct: 76 FPDWINYSYDLPMMKVWDNMIIESALKTYYNKSLPLMIEALQANLT 121
>sptr|Q9FTR1|Q9FTR1 Putative bifunctional nuclease.
Length = 311
Score = 196 bits (499), Expect = 4e-49
Identities = 117/289 (40%), Positives = 148/289 (51%), Gaps = 29/289 (10%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYL------------SEKAAAAVQALLPESA---GGELSTVCPWA 834
AWG GH++VC+IA+ E AA V+ PE G P
Sbjct: 27 AWGIHGHLIVCQIAQARTFFRPAERRGRRGGEGPAAVVRRGQPEQPLLLGRRRQAQVPLV 86
Query: 833 DQVRWHYHWASPLHYANTPQVCNFKYSRDCHNSRGQQGMCVVGA--------------IN 696
H H PL + F+ +R S G +N
Sbjct: 87 GAAPLHRHPGPPLQL----HLRQFETARTRIASGGGASPAPSTTTPPSSSPMMQHPLLLN 142
Query: 695 NYTDQLYSYGQKTSYNLTESLMFLAHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLH 516
+ D + +S NLT++L+FLAHFVGD+HQPLHVGF D+GGNTI VHWY RK LH
Sbjct: 143 SQADDKFLI-LSSSDNLTQALLFLAHFVGDIHQPLHVGFTSDKGGNTIDVHWYTRKTVLH 201
Query: 515 HVWDVSIIDTAIKDFYNKSMDTMVEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIES 336
HVWD +II TA D+Y + + V+AL N+T WS + WE C TC + YA ES
Sbjct: 202 HVWDDNIIKTAENDYYGEGVAEFVDALMQNITGEWSQRVPGWEECSKNQTTCPDTYASES 261
Query: 335 IHYSCNYAYKDVEQDITLGDDYFFSRYPIVEKRLAQAGIRLALVLNRIF 189
I +C++AYKDV +D L D YF SR P+V RLAQ G+RLA LNRIF
Sbjct: 262 IAAACDWAYKDVTEDSLLEDAYFGSRLPVVNLRLAQGGVRLAATLNRIF 310
>sptrnew|CAE04161|CAE04161 OSJNBb0034I13.4 protein.
Length = 252
Score = 186 bits (473), Expect = 4e-46
Identities = 108/269 (40%), Positives = 148/269 (55%), Gaps = 3/269 (1%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPL 795
+W KEGH++ C+IA+ L AA AV+ LL E A G+LS +C W DQVR W+ Y W SPL
Sbjct: 29 SWSKEGHMLTCRIAQDLLEPAAAHAVRNLLTEEADGDLSALCVWPDQVRHWYKYRWTSPL 88
Query: 794 HYANTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFLAHF 615
H+ +T ++ +S QG+ + + YN+TE+L+FL+HF
Sbjct: 89 HFIDT-------LTKPAASST--QGIAMARMVRRI------------YNMTEALLFLSHF 127
Query: 614 VGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMVEAL 435
+GDVHQ VWD +I TAI +FY K MD + L
Sbjct: 128 MGDVHQ----------------------------VWDREMILTAIAEFYGKDMDAFQKDL 159
Query: 434 KMNLTDG-WSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKDVEQDITLGDDYFFSR 258
N T G WSDD+ W +C++ +C+ YA ESI+ +C +AY DV + TL DDYF SR
Sbjct: 160 VHNFTTGTWSDDVSSWGDCEDL-LSCSTKYATESINLACKWAYNDVREGETLSDDYFGSR 218
Query: 257 YPIVEKRLAQAGIRLALVLNRIFGGGEAD 171
PIV +R+AQ G+RLA+ LNR+FG D
Sbjct: 219 LPIVTRRIAQGGVRLAMFLNRLFGEHNRD 247
>sptr|Q93WW9|Q93WW9 S1-type endonuclease (Fragment).
Length = 136
Score = 157 bits (398), Expect = 2e-37
Identities = 73/136 (53%), Positives = 97/136 (71%), Gaps = 6/136 (4%)
Frame = -2
Query: 929 AEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVR-WH-YHWASPLHYANTPQ-VCNFK 759
A+ L AA V+ LLP GG+LS +C W DQ+R W+ Y W+SPLH+ +TP C+F
Sbjct: 1 AQDLLQPDAAEVVKNLLPHYVGGDLSALCTWPDQIRHWYKYRWSSPLHFIDTPDNACSFD 60
Query: 758 YSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS---YNLTESLMFLAHFVGDVHQPLH 588
Y+RDCH+ +GQ+ MCV GA+ NYT QL + +S YNL+ESL+FL+HF+GD+HQP+H
Sbjct: 61 YTRDCHDPKGQEDMCVAGAVRNYTTQLLHNREGSSDRRYNLSESLLFLSHFMGDIHQPMH 120
Query: 587 VGFEDDEGGNTITVHW 540
VGF DEGGNTI + W
Sbjct: 121 VGFTSDEGGNTIDLRW 136
>sptr|O65425|O65425 Hypothetical 51.9 kDa protein (Putative
bifunctional nuclease).
Length = 454
Score = 153 bits (387), Expect = 4e-36
Identities = 73/144 (50%), Positives = 97/144 (67%), Gaps = 8/144 (5%)
Frame = -2
Query: 920 YLSEKAAAAVQALLPESA-GGELSTVCPWADQVRW--HYHWASPLHYANTPQV-CNFKYS 753
+ + AAV+ LLPES GG L+ C W D+++ + W S LHY NTP+ CN++Y
Sbjct: 4 FFEDDTIAAVKKLLPESVDGGGLADFCSWPDEIKKLSQWQWTSTLHYVNTPEYRCNYEYC 63
Query: 752 RDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTS----YNLTESLMFLAHFVGDVHQPLHV 585
RDCH++ + CV GAI NYT+QL S + + YNLTE+L+FL+H++GDVHQPLH
Sbjct: 64 RDCHDTHKHKDWCVTGAIFNYTNQLMSASENSQNIVHYNLTEALLFLSHYMGDVHQPLHT 123
Query: 584 GFEDDEGGNTITVHWYRRKANLHH 513
GF D GGNTI V+WY K+NLHH
Sbjct: 124 GFLGDLGGNTIIVNWYHNKSNLHH 147
>sptrnew|EAA29902|EAA29902 Hypothetical protein.
Length = 324
Score = 137 bits (346), Expect = 2e-31
Identities = 97/288 (33%), Positives = 135/288 (46%), Gaps = 21/288 (7%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHY--HWASPL 795
AWG GHI V +A ++S A+ Q LL L+ V WAD +R+ W PL
Sbjct: 20 AWGGFGHITVAYVASNFVSNSTASYFQTLLRNDTSDYLANVATWADSIRYTKWGRWTGPL 79
Query: 794 HYANT----PQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMF 627
HY + P C Y RDC + CVV AI NYT +L ++ F
Sbjct: 80 HYIDAKDSPPDSCGIIYERDC-----KPEGCVVSAIQNYTSRLLDQSLHV-VERAQAAKF 133
Query: 626 LAHFVGDVHQPLHVGFED-DEGGNTITVHWYRRKANLHHVWDVSIIDTAIK--------- 477
+ HFVGD+HQPLH ED ++GGN I+V + ++ NLHHVWD SI + +
Sbjct: 134 VIHFVGDIHQPLHT--EDVEKGGNGISVFFDEKRFNLHHVWDSSIAEKIVTHKKQGVGRR 191
Query: 476 --DFYNKSMDTMVEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKD 303
K D + E ++ S D V K + + A +AIE + C D
Sbjct: 192 PFPAAKKWADALAEEIREGQYKANSSDWVKGLELK-EASEIALQWAIEGNEHVCTVVLPD 250
Query: 302 ---VEQDITLGDDYFFSRYPIVEKRLAQAGIRLALVLNRIFGGGEADD 168
+D LG YF + P+VE ++A+AG RLA L+ I G E ++
Sbjct: 251 GPEAIRDQELGGAYFEAAAPVVELQIAKAGYRLAAWLDLIVSGIEKNE 298
>sptr|Q870T1|Q870T1 Probable nuclease S1.
Length = 324
Score = 137 bits (346), Expect = 2e-31
Identities = 97/288 (33%), Positives = 135/288 (46%), Gaps = 21/288 (7%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHY--HWASPL 795
AWG GHI V +A ++S A+ Q LL L+ V WAD +R+ W PL
Sbjct: 20 AWGGFGHITVAYVASNFVSNSTASYFQTLLRNDTSDYLANVATWADSIRYTKWGRWTGPL 79
Query: 794 HYANT----PQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMF 627
HY + P C Y RDC + CVV AI NYT +L ++ F
Sbjct: 80 HYIDAKDSPPDSCGIIYERDC-----KPEGCVVSAIQNYTSRLLDQSLHV-VERAQAAKF 133
Query: 626 LAHFVGDVHQPLHVGFED-DEGGNTITVHWYRRKANLHHVWDVSIIDTAIK--------- 477
+ HFVGD+HQPLH ED ++GGN I+V + ++ NLHHVWD SI + +
Sbjct: 134 VIHFVGDIHQPLHT--EDVEKGGNGISVFFDEKRFNLHHVWDSSIAEKIVTHKKQGVGRR 191
Query: 476 --DFYNKSMDTMVEALKMNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKD 303
K D + E ++ S D V K + + A +AIE + C D
Sbjct: 192 PFPAAKKWADALAEEIREGQYKANSSDWVKGLELK-EASEIALQWAIEGNEHVCTVVLPD 250
Query: 302 ---VEQDITLGDDYFFSRYPIVEKRLAQAGIRLALVLNRIFGGGEADD 168
+D LG YF + P+VE ++A+AG RLA L+ I G E ++
Sbjct: 251 GPEAIRDQELGGAYFEAAAPVVELQIAKAGYRLAAWLDLIVSGIEKNE 298
>sptr|Q9P356|Q9P356 Nuclease Le1.
Length = 310
Score = 130 bits (328), Expect = 2e-29
Identities = 91/293 (31%), Positives = 129/293 (44%), Gaps = 32/293 (10%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWH--YHWASPLH 792
WG GH V +A K+LS KA + VQ+ L L PWAD VR+ + W++P H
Sbjct: 21 WGAIGHETVGYVAMKFLSPKALSFVQSSLGSEYSESLGPAAPWADDVRFEAAFSWSAPFH 80
Query: 791 YA----NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFL 624
+ N P C+ + RDC G C++ AI NYT ++ T E+L F+
Sbjct: 81 FVDAEDNPPTSCSVEQMRDC-----SSGTCILSAIANYTTRVVDISLSTEQR-QEALKFI 134
Query: 623 AHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDTMV 444
HF+GD+ QPLHV + GGN I + NLH VWD II+ +K Y+ S
Sbjct: 135 DHFIGDIGQPLHVE-AVEVGGNDINAVCNGERTNLHAVWDSGIINIFLKAQYSNSAIVWA 193
Query: 443 EALKM--------NLTDGWSDDIVHWENCKNKHATCAND------------------YAI 342
AL +LT W E N+ + +D +A
Sbjct: 194 NALAQRIQTGEFKSLTSTWLSCSSTTEPVNNRRRSIEDDINGLVSDATITPLECPLVWAR 253
Query: 341 ESIHYSCNYAYKDVEQDITLGDDYFFSRYPIVEKRLAQAGIRLALVLNRIFGG 183
ES Y C++ + Y PI+E+++A+ G RLA LN +F G
Sbjct: 254 ESNAYDCSFVFSYTGFSDLCTSSYATGAQPIIEEQIAKQGYRLAAWLNVLFDG 306
>sptr|Q8NIH8|Q8NIH8 Nuclease Le3.
Length = 298
Score = 127 bits (319), Expect = 3e-28
Identities = 94/283 (33%), Positives = 141/283 (49%), Gaps = 22/283 (7%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLP-ESAGGELSTVCPWADQVRWH--YHWASPL 795
WG +GH V IA K+L+ +A++ V+ L L PWAD+VR Y W++PL
Sbjct: 19 WGMKGHEAVGFIAMKFLAPEASSFVETSLSGPQYHSSLGLAAPWADEVRRQKGYAWSAPL 78
Query: 794 HYANT----PQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMF 627
H+ + P C+ RDC + C++ AI NYT ++ S + E+L F
Sbjct: 79 HFVDAEDQPPTECSVNQKRDCAGNG-----CILTAIANYTSRVVDTSLSDS-DRQEALKF 132
Query: 626 LAHFVGDVHQPLHV-GFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSMDT 450
+ HF+GD+ QPLHV G E GGN I V +K NLH VWD II+ + D Y+ S+
Sbjct: 133 IDHFIGDIGQPLHVEGIE--RGGNGIHVQCAGKKNNLHSVWDDGIINKLLDDKYDGSVIQ 190
Query: 449 MVEAL-----KMNLTDGWSD---DIVHWENCK----NKHATCANDYAIESIHYSCNYAYK 306
V +L +++ W + +C + C +A ES Y C++ +
Sbjct: 191 WVNSLIERIQQLDCLRAWIKLPLGVRRRADCPRASLDDTLNCPLVWAKESNAYDCSFVWT 250
Query: 305 -DVEQDITLGD-DYFFSRYPIVEKRLAQAGIRLALVLNRIFGG 183
D +D+ D DY+ PI+E ++A+ G RLA LN +F G
Sbjct: 251 YDSYEDLCSDDNDYYSGAVPIIELQIAKQGYRLAAWLNVLFDG 293
>sptrnew|EAA32713|EAA32713 Hypothetical protein.
Length = 306
Score = 115 bits (288), Expect = 1e-24
Identities = 84/292 (28%), Positives = 129/292 (44%), Gaps = 32/292 (10%)
Frame = -2
Query: 968 AWGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWH---YHWASP 798
AWGK GH V +A++YL+ VQ +L +++ + + WAD R+ W++
Sbjct: 19 AWGKLGHATVASVAQQYLTPNTVKQVQTILGDNSTSYMGNIASWADSFRYESAANAWSAG 78
Query: 797 LHYAN-----TPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESL 633
LH+ N P+ C+ DC CVV AI NYT+++ T+ ++L
Sbjct: 79 LHFVNGHDGPPPESCHLVLPEDCPPEG-----CVVSAIGNYTERV-QMKNITADQKAQAL 132
Query: 632 MFLAHFVGDVHQPLHV-GFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSM 456
F+ HF+GD+ QPLH GF EG N ITV + K NLH WD SI + + +
Sbjct: 133 KFIVHFLGDIAQPLHTEGF--GEGANNITVTFQGYKTNLHAAWDTSIPNAML------GI 184
Query: 455 DTMVEALKMNLTD--GWSDDIV------------------HWENCKNKHATCANDYAIES 336
A + D GW++++ H + A +A +
Sbjct: 185 SPPTSAANITSADFLGWANNLAAKINQGQYRKDVRRWLRYHSVATRKASERAAAAWAQDG 244
Query: 335 IHYSCNYAYKDVEQDIT---LGDDYFFSRYPIVEKRLAQAGIRLALVLNRIF 189
C+Y K + +G DY+ +VE+ + + GIRLA LN IF
Sbjct: 245 NEEVCHYVMKVPGNQLNGTEIGGDYYKGATEVVERSIIKGGIRLAGWLNLIF 296
>sw|P24289|NUP1_PENCI Nuclease P1 (EC 3.1.30.1) (Endonuclease P1)
(Deoxyribonuclease P1).
Length = 270
Score = 115 bits (287), Expect = 1e-24
Identities = 87/276 (31%), Positives = 127/276 (46%), Gaps = 16/276 (5%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHY--HWASPLH 792
WG GH V +A+ Y+S +AA+ Q +L S+ L+++ WAD+ R W++ LH
Sbjct: 1 WGALGHATVAYVAQHYVSPEAASWAQGILGSSSSSYLASIASWADEYRLTSAGKWSASLH 60
Query: 791 YA----NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFL 624
+ N P CN Y RDC +S C + AI NYT Q S +S N E+L FL
Sbjct: 61 FIDAEDNPPTNCNVDYERDCGSSG-----CSISAIANYT-QRVSDSSLSSENHAEALRFL 114
Query: 623 AHFVGDVHQPLHVGFEDDE----GGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSM 456
HF+GD+ QPLH DE GGN I V + NLH WD + I
Sbjct: 115 VHFIGDMTQPLH-----DEAYAVGGNKINVTFDGYHDNLHSDWDTYMPQKLIGGHALSDA 169
Query: 455 DTMVEALKMNLTDG-WSDDIVHWENCKN--KHATCANDYAIESIHYSCNYAYKDVEQDIT 285
++ + L N+ G ++ + W N + T A +A ++ C +
Sbjct: 170 ESWAKTLVQNIESGNYTAQAIGWIKGDNISEPITTATRWASDANALVCTVVMPHGAAALQ 229
Query: 284 LGD---DYFFSRYPIVEKRLAQAGIRLALVLNRIFG 186
GD Y+ S +E ++A+ G RLA +N I G
Sbjct: 230 TGDLYPTYYDSVIDTIELQIAKGGYRLANWINEIHG 265
>sw|P24504|NUP3_PENSQ Nuclease PA3 (EC 3.1.3.6) (Endonuclease PA3)
(Deoxyribonuclease PA3).
Length = 270
Score = 114 bits (284), Expect = 3e-24
Identities = 87/276 (31%), Positives = 126/276 (45%), Gaps = 16/276 (5%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHY--HWASPLH 792
WG GH V +A+ Y+S +AA+ Q +L S+ L+++ WAD+ R W++ LH
Sbjct: 1 WGALGHATVAYVAQHYVSPEAASWAQGILGSSSSSYLASIASWADEYRLTSAGKWSASLH 60
Query: 791 YA----NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFL 624
+ N P CN Y RDC +S C + AI NYT Q S +S N E+L FL
Sbjct: 61 FIDAEDNPPTNCNVDYERDCGSSG-----CSISAIANYT-QRVSDSSLSSENHAEALRFL 114
Query: 623 AHFVGDVHQPLHVGFEDDE----GGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSM 456
HF+GD+ QPLH DE GGN I V + NLH WD + I
Sbjct: 115 VHFIGDMTQPLH-----DEAYAVGGNKINVTFDGYHDNLHSDWDTYMPQKLIGGHALSDA 169
Query: 455 DTMVEALKMNLTDG-WSDDIVHWENCKN--KHATCANDYAIESIHYSCNYAYKDVEQDIT 285
++ + L N+ G ++ W N + T A +A ++ C +
Sbjct: 170 ESWAKTLVQNIESGNYTAQATGWIKGDNISEPITTATRWASDANALVCTVVMPHGAAALQ 229
Query: 284 LGD---DYFFSRYPIVEKRLAQAGIRLALVLNRIFG 186
GD Y+ S +E ++A+ G RLA +N I G
Sbjct: 230 TGDLYPTYYDSVIDTIELQIAKGGYRLANWINEIHG 265
>sptr|Q00235|Q00235 Nuclease S1 precursor.
Length = 287
Score = 107 bits (267), Expect = 3e-22
Identities = 82/276 (29%), Positives = 129/276 (46%), Gaps = 18/276 (6%)
Frame = -2
Query: 965 WGKEGHIMVCKIAEKYLSEKAAAAVQALLPESAGGELSTVCPWADQVRWHY--HWASPLH 792
WG GH V IA+ +++ + Q +L + + L+ V WAD ++ ++ P H
Sbjct: 21 WGNLGHETVAYIAQSFVASSTESFCQNILGDDSTSYLANVATWADTYKYTDAGEFSKPYH 80
Query: 791 YA----NTPQVCNFKYSRDCHNSRGQQGMCVVGAINNYTDQLYSYGQKTSYNLTESLMFL 624
+ N PQ C Y RDC G G C + AI NYT+ L +L F+
Sbjct: 81 FIDAQDNPPQSCGVDYDRDC----GSAG-CSISAIQNYTNILLE--SPNGSEALNALKFV 133
Query: 623 AHFVGDVHQPLHVGFEDDEGGNTITVHWYRRKANLHHVWDVSIIDTAIKDFYNKSM---- 456
H +GD+HQPLH + GGN I V + NLHH+WD ++ + A Y+ S+
Sbjct: 134 VHIIGDIHQPLH-DENLEAGGNGIDVTYDGETTNLHHIWDTNMPEEAAGG-YSLSVAKTY 191
Query: 455 -DTMVEALK----MNLTDGWSDDIVHWENCKNKHATCANDYAIESIHYSCNYAYKD---V 300
D + E +K + D W+D I + K+ +T + +A ++ Y C+ D
Sbjct: 192 ADLLTERIKTGTYSSKKDSWTDGI----DIKDPVST-SMIWAADANTYVCSTVLDDGLAY 246
Query: 299 EQDITLGDDYFFSRYPIVEKRLAQAGIRLALVLNRI 192
L +Y+ P+ E+ +A+AG RLA L+ I
Sbjct: 247 INSTDLSGEYYDKSQPVFEELIAKAGYRLAAWLDLI 282
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 677,382,919
Number of Sequences: 1272877
Number of extensions: 13900731
Number of successful extensions: 40544
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 37925
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 40360
length of database: 406,886,216
effective HSP length: 124
effective length of database: 249,049,468
effective search space used: 58028526044
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

