BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
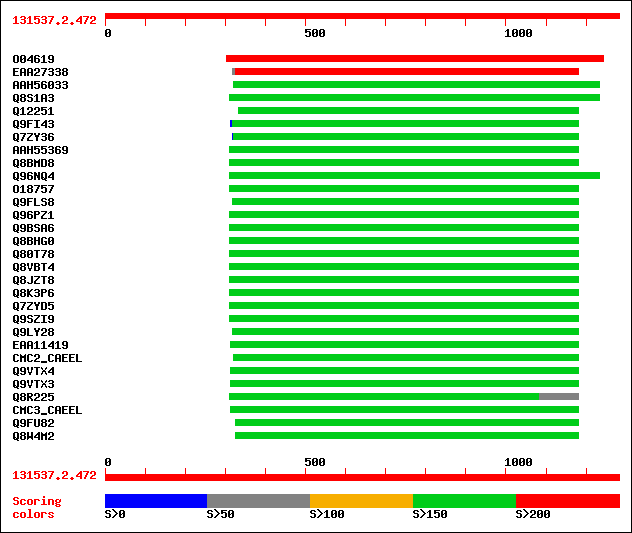
Query= 131537.2.472
(1282 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O04619|O04619 Putative carrier protein (AT4G01100/F2N1_16). 504 e-142
sptrnew|EAA27338|EAA27338 Hypothetical protein. 214 2e-54
sptrnew|AAH56033|AAH56033 Hypothetical protein. 190 3e-47
sptr|Q8S1A3|Q8S1A3 Putative mitochondrial carrier. 189 7e-47
sptr|Q12251|Q12251 Hypothetical 35.9 kDa protein YPR011C. 189 7e-47
sptr|Q9FI43|Q9FI43 Calcium-binding transporter-like protein. 189 7e-47
sptr|Q7ZY36|Q7ZY36 Similar to hypothetical protein LOC229731 (Fr... 187 2e-46
sptrnew|AAH55369|AAH55369 Calcium-binding transporter. 184 2e-45
sptr|Q8BMD8|Q8BMD8 Peroxisomal CA-dependent solute carrier homolog. 184 2e-45
sptr|Q96NQ4|Q96NQ4 Hypothetical protein FLJ30339. 184 2e-45
sptr|O18757|O18757 Peroxisomal Ca-dependent solute carrier. 182 9e-45
sptr|Q9FLS8|Q9FLS8 Peroxisomal Ca-dependent solute carrier-like ... 181 2e-44
sptr|Q96PZ1|Q96PZ1 Hypothetical protein KIAA1896 (Fragment). 179 6e-44
sptr|Q9BSA6|Q9BSA6 Hypothetical protein (Fragment). 179 6e-44
sptr|Q8BHG0|Q8BHG0 Weakly similar to peroxisomal CA-dependent so... 179 8e-44
sptr|Q80T78|Q80T78 MKIAA1896 protein (Fragment). 179 8e-44
sptr|Q8VBT4|Q8VBT4 Hypothetical 40.6 kDa protein (Similar to KIA... 179 8e-44
sptr|Q8JZT8|Q8JZT8 Similar to peroxisomal Ca-dependent solute ca... 179 8e-44
sptr|Q8K3P6|Q8K3P6 Peroxisomal Ca-dependent solute carrier-like ... 179 1e-43
sptr|Q7ZYD5|Q7ZYD5 Similar to hypothetical protein MGC36388. 178 2e-43
sptr|Q9SZI9|Q9SZI9 Putative mitochondrial carrier protein. 177 2e-43
sptr|Q9LY28|Q9LY28 Peroxisomal Ca-dependent solute carrier-like ... 177 2e-43
sptrnew|EAA11419|EAA11419 ENSANGP00000009995 (Fragment). 177 3e-43
sw|Q20799|CMC2_CAEEL Probable calcium-binding mitochondrial carr... 171 2e-41
sptr|Q9VTX4|Q9VTX4 CG32103 protein. 170 5e-41
sptr|Q9VTX3|Q9VTX3 CG32103 protein (RE56970p). 170 5e-41
sptr|Q8R225|Q8R225 Hypothetical 25.4 kDa protein (Fragment). 170 5e-41
sw|Q19529|CMC3_CAEEL Probable calcium-binding mitochondrial carr... 169 8e-41
sptr|Q9FU82|Q9FU82 Putative peroxisomal Ca-dependent solute carr... 168 1e-40
sptr|Q8W4M2|Q8W4M2 Hypothetical 39.8 kDa protein. 168 2e-40
>sptr|O04619|O04619 Putative carrier protein (AT4G01100/F2N1_16).
Length = 352
Score = 504 bits (1299), Expect = e-142
Identities = 251/316 (79%), Positives = 277/316 (87%), Gaps = 2/316 (0%)
Frame = -1
Query: 1243 TICKSLXXXXXXXXXXXXXXAPLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLF 1064
+ICKSL APLER+KILLQVQNPH+IKY+GTVQGLK+IWRTEGLRGLF
Sbjct: 37 SICKSLFAGGVAGGVSRTAVAPLERMKILLQVQNPHNIKYSGTVQGLKHIWRTEGLRGLF 96
Query: 1063 KGNGTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAM 884
KGNGTNCARIVPNSAVKFFSYEQAS GIL+ YRQ+TG E+AQLTPLLRLGAGA AGIIAM
Sbjct: 97 KGNGTNCARIVPNSAVKFFSYEQASNGILYMYRQRTGNENAQLTPLLRLGAGATAGIIAM 156
Query: 883 SATYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGL 704
SATYPMDMVRGR+TVQT SPYQYRG+ HAL TV REEG RALY+GWLPSVIGVVPYVGL
Sbjct: 157 SATYPMDMVRGRLTVQTANSPYQYRGIAHALATVLREEGPRALYRGWLPSVIGVVPYVGL 216
Query: 703 NFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVG 524
NF+VYESLKDWL++ N +GL +NEL VVTRL CGA+AGT+GQT+AYPLDVIRRRMQMVG
Sbjct: 217 NFSVYESLKDWLVKENPYGLVENNELTVVTRLTCGAIAGTVGQTIAYPLDVIRRRMQMVG 276
Query: 523 WSHADSIVTGQGKE--ALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVT 350
W A +IVTG+G+ +L+Y GM+DAFRKTVRHEG GALYKGLVPNSVKVVPSIAIAFVT
Sbjct: 277 WKDASAIVTGEGRSTASLEYTGMVDAFRKTVRHEGFGALYKGLVPNSVKVVPSIAIAFVT 336
Query: 349 YEVVKDVLGVEMRISD 302
YE+VKDVLGVE RISD
Sbjct: 337 YEMVKDVLGVEFRISD 352
>sptrnew|EAA27338|EAA27338 Hypothetical protein.
Length = 338
Score = 214 bits (545), Expect = 2e-54
Identities = 129/293 (44%), Positives = 176/293 (60%), Gaps = 8/293 (2%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTV-QGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFS 1004
PLERLKIL QVQ+ Y +V + L +WR EG RG GNGTNC RIVP SAV+F S
Sbjct: 56 PLERLKILYQVQSSGREAYKLSVGKALAKMWREEGWRGFMAGNGTNCIRIVPYSAVQFGS 115
Query: 1003 YEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQT--- 833
Y + I + + G+ LTPL RL G AGI +++ TYP+D+VR R+++QT
Sbjct: 116 YNFYKRNI---FERHPGDS---LTPLSRLTCGGLAGITSVTFTYPLDIVRTRLSIQTASF 169
Query: 832 ---DKSPYQYRGMFHALGTVYREEG-FRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLL 665
+ P + GM+ L +YR EG F ALY+G +P+V GV PYVGLNF VYE ++ +L
Sbjct: 170 AELGERPRKMPGMWETLVKMYRTEGGFPALYRGIVPTVAGVAPYVGLNFMVYEHVRQYLT 229
Query: 664 QTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGK 485
L + V +L GA++G + QT YP DV+RRR Q+ + ++G G
Sbjct: 230 ------LDGEQNPSAVRKLLAGAISGAVAQTCTYPFDVLRRRFQI-------NTMSGMG- 275
Query: 484 EALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVL 326
QY G+ DA R V EG+ LYKG+VPN +KV PS+A ++++YEV +D L
Sbjct: 276 --YQYKGIFDAVRVIVTEEGIRGLYKGIVPNLLKVAPSMASSWLSYEVCRDFL 326
Score = 87.4 bits (215), Expect = 4e-16
Identities = 60/200 (30%), Positives = 95/200 (47%), Gaps = 1/200 (0%)
Frame = -1
Query: 913 AGACAGIIAMSATYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPS 734
AG AG ++ + P++ ++ VQ+ + AL ++REEG+R G +
Sbjct: 42 AGGVAGAVSRTVVSPLERLKILYQVQSSGREAYKLSVGKALAKMWREEGWRGFMAGNGTN 101
Query: 733 VIGVVPYVGLNFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLD 554
I +VPY + F Y K N F + L ++RL CG +AG T YPLD
Sbjct: 102 CIRIVPYSAVQFGSYNFYK-----RNIFERHPGDSLTPLSRLTCGGLAGITSVTFTYPLD 156
Query: 553 VIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHE-GVGALYKGLVPNSVKVV 377
++R R+ + S A+ G+ + GM + K R E G ALY+G+VP V
Sbjct: 157 IVRTRLSIQTASFAE-----LGERPRKMPGMWETLVKMYRTEGGFPALYRGIVPTVAGVA 211
Query: 376 PSIAIAFVTYEVVKDVLGVE 317
P + + F+ YE V+ L ++
Sbjct: 212 PYVGLNFMVYEHVRQYLTLD 231
Score = 82.0 bits (201), Expect = 2e-14
Identities = 51/157 (32%), Positives = 84/157 (53%), Gaps = 2/157 (1%)
Frame = -1
Query: 1129 KYNGTVQGLKYIWRTEG-LRGLFKGNGTNCARIVPNSAVKFFSYEQASKGILWAYRQQTG 953
K G + L ++RTEG L++G A + P + F YE + Y G
Sbjct: 178 KMPGMWETLVKMYRTEGGFPALYRGIVPTVAGVAPYVGLNFMVYEHVRQ-----YLTLDG 232
Query: 952 EEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP-YQYRGMFHALGTVYR 776
E++ + + +L AGA +G +A + TYP D++R R + T YQY+G+F A+ +
Sbjct: 233 EQNP--SAVRKLLAGAISGAVAQTCTYPFDVLRRRFQINTMSGMGYQYKGIFDAVRVIVT 290
Query: 775 EEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLL 665
EEG R LYKG +P+++ V P + ++ YE +D+L+
Sbjct: 291 EEGIRGLYKGIVPNLLKVAPSMASSWLSYEVCRDFLV 327
>sptrnew|AAH56033|AAH56033 Hypothetical protein.
Length = 473
Score = 190 bits (483), Expect = 3e-47
Identities = 111/306 (36%), Positives = 174/306 (56%), Gaps = 1/306 (0%)
Frame = -1
Query: 1234 KSLXXXXXXXXXXXXXXAPLERLKILLQVQNPHSIKYNGTV-QGLKYIWRTEGLRGLFKG 1058
K L APL+RLK+++QV H K N + GLK + + G+R L++G
Sbjct: 196 KQLMAGGMAGAVSRTGTAPLDRLKVMMQV---HGSKGNSNIITGLKQMVKEGGIRSLWRG 252
Query: 1057 NGTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSA 878
NG N +I P +A+KF++YEQ Y++ E +L R AG+ AG A ++
Sbjct: 253 NGVNVIKIAPETAMKFWAYEQ--------YKKLFTSESGKLGTAERFVAGSLAGATAQTS 304
Query: 877 TYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNF 698
YPM++++ R+ V QY GMF + ++EG RA YKG++P+++G++PY G++
Sbjct: 305 IYPMEVLKTRLAVGRTG---QYSGMFDCAKKIMQKEGIRAFYKGYIPNILGIIPYAGIDL 361
Query: 697 AVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWS 518
A+YE+LK++ LQ ++ AN V+ LGCG + T GQ +YPL +IR RMQ
Sbjct: 362 AIYETLKNYWLQNHAKDSANPG---VLVLLGCGTASSTCGQLASYPLALIRTRMQ----- 413
Query: 517 HADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVV 338
+ + +G L G+ FRK V EG LY+G+ PN +KV+P+++I++V YE +
Sbjct: 414 ---AQASIEGAPQLNMGGL---FRKIVAKEGFLGLYRGIGPNFLKVLPAVSISYVVYEKM 467
Query: 337 KDVLGV 320
K LG+
Sbjct: 468 KVQLGI 473
>sptr|Q8S1A3|Q8S1A3 Putative mitochondrial carrier.
Length = 340
Score = 189 bits (480), Expect = 7e-47
Identities = 118/321 (36%), Positives = 170/321 (52%), Gaps = 13/321 (4%)
Frame = -1
Query: 1234 KSLXXXXXXXXXXXXXXAPLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGN 1055
K L APLER+KILLQ + H + G +Q L+ +W+ EG+RG +KGN
Sbjct: 31 KELIAGGAAGAFAKTAVAPLERVKILLQTRT-HGFQSLGILQSLRKLWQYEGIRGFYKGN 89
Query: 1054 GTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSAT 875
G + RIVP +A+ + +YEQ IL + G P++ L AG+ AG A+ T
Sbjct: 90 GASVLRIVPYAALHYMTYEQYRCWILNNFAPSVGTG-----PVVDLLAGSAAGGTAVLCT 144
Query: 874 YPMDMVRGRITVQTD-------------KSPYQYRGMFHALGTVYREEGFRALYKGWLPS 734
YP+D+ R ++ Q + P Y G+ TVY+E G RALY+G P+
Sbjct: 145 YPLDLARTKLAYQVSNVGQPGNALGNAGRQP-AYGGIKDVFKTVYKEGGARALYRGVGPT 203
Query: 733 VIGVVPYVGLNFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLD 554
+IG++PY GL F +YE LK + D + VV +L CGA+AG GQT+ YPLD
Sbjct: 204 LIGILPYAGLKFYIYEDLKS--------RVPEDYKRSVVLKLSCGALAGLFGQTLTYPLD 255
Query: 553 VIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVP 374
V+RR+MQ+ + +A + G +R +G L+ GL N VKVVP
Sbjct: 256 VVRRQMQV------QNKQPHNANDAFRIRGTFQGLALIIRCQGWRQLFAGLSLNYVKVVP 309
Query: 373 SIAIAFVTYEVVKDVLGVEMR 311
S+AI F TY+++K++L V R
Sbjct: 310 SVAIGFTTYDMMKNLLRVPPR 330
>sptr|Q12251|Q12251 Hypothetical 35.9 kDa protein YPR011C.
Length = 326
Score = 189 bits (480), Expect = 7e-47
Identities = 116/294 (39%), Positives = 169/294 (57%), Gaps = 11/294 (3%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
P ER+KILLQVQ+ + G ++ ++ EG +GLF+GNG NC RI P SAV+F Y
Sbjct: 42 PFERVKILLQVQSSTTSYNRGIFSSIRQVYHEEGTKGLFRGNGLNCIRIFPYSAVQFVVY 101
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQT---- 833
E A K L+ G+E QLT RL +GA G ++ ATYP+D+++ R+++QT
Sbjct: 102 E-ACKKKLFHVNGNNGQE--QLTNTQRLFSGALCGGCSVVATYPLDLIKTRLSIQTANLS 158
Query: 832 ------DKSPYQYRGMFHALGTVYR-EEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKD 674
KS + G++ L YR E G R LY+G P+ +GVVPYV LNFAVYE L++
Sbjct: 159 SLNRSKAKSISKPPGIWQLLSETYRLEGGLRGLYRGVWPTSLGVVPYVALNFAVYEQLRE 218
Query: 673 WLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTG 494
+ + ++ + + L+ +L GA++G + QT+ YP D++RRR Q+ + G
Sbjct: 219 FGVNSSDAQPSWKSNLY---KLTIGAISGGVAQTITYPFDLLRRRFQV--------LAMG 267
Query: 493 QGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKD 332
+ +Y + DA R EGV YKGL N KVVPS A++++ YEVV D
Sbjct: 268 GNELGFRYTSVWDALVTIGRAEGVSGYYKGLAANLFKVVPSTAVSWLVYEVVCD 321
>sptr|Q9FI43|Q9FI43 Calcium-binding transporter-like protein.
Length = 487
Score = 189 bits (480), Expect = 7e-47
Identities = 106/288 (36%), Positives = 164/288 (56%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+LLQ+Q + + +K IW+ G+RG F+GNG N ++ P SA+KF++Y
Sbjct: 228 PLDRLKVLLQIQKTDA----RIREAIKLIWKQGGVRGFFRGNGLNIVKVAPESAIKFYAY 283
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
E I + GE+ A + +RL AG AG +A ++ YP+D+V+ R+ T ++
Sbjct: 284 ELFKNAI----GENMGEDKADIGTTVRLFAGGMAGAVAQASIYPLDLVKTRLQTYTSQAG 339
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
+ + EG RA YKG PS++G++PY G++ A YE+LKD L +T +
Sbjct: 340 VAVPRLGTLTKDILVHEGPRAFYKGLFPSLLGIIPYAGIDLAAYETLKD-LSRTY---IL 395
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
D E + +LGCG ++G +G T YPL V+R RMQ A + ++G
Sbjct: 396 QDAEPGPLVQLGCGTISGALGATCVYPLQVVRTRMQA---ERARTSMSG----------- 441
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
FR+T+ EG ALYKGL+PN +KVVP+ +I ++ YE +K L ++
Sbjct: 442 --VFRRTISEEGYRALYKGLLPNLLKVVPAASITYMVYEAMKKSLELD 487
Score = 37.4 bits (85), Expect = 0.47
Identities = 26/96 (27%), Positives = 42/96 (43%)
Frame = -1
Query: 601 GAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGV 422
G +AG +T PLD ++ +Q+ D+ + +A + + GV
Sbjct: 215 GGIAGAASRTATAPLDRLKVLLQI---QKTDARIR-------------EAIKLIWKQGGV 258
Query: 421 GALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEM 314
++G N VKV P AI F YE+ K+ +G M
Sbjct: 259 RGFFRGNGLNIVKVAPESAIKFYAYELFKNAIGENM 294
>sptr|Q7ZY36|Q7ZY36 Similar to hypothetical protein LOC229731
(Fragment).
Length = 535
Score = 187 bits (476), Expect = 2e-46
Identities = 108/288 (37%), Positives = 169/288 (58%), Gaps = 1/288 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTV-QGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFS 1004
PL+RLK+++QV H K N + GLK + + G+R L++GNG N +I P +A+KF++
Sbjct: 276 PLDRLKVMMQV---HGTKGNSNIITGLKQMVKEGGVRSLWRGNGVNVIKIAPETAMKFWA 332
Query: 1003 YEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKS 824
YEQ Y++ E +L R AG+ AG A ++ YPM++++ R+ V
Sbjct: 333 YEQ--------YKKLFTSESGKLGTAERFIAGSLAGATAQTSIYPMEVLKTRLAVGKTG- 383
Query: 823 PYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGL 644
QY GMF + ++EG A YKG++P+++G++PY G++ A+YE+LK++ LQ +
Sbjct: 384 --QYSGMFDCAKKIMQKEGILAFYKGYIPNILGIIPYAGIDLAIYETLKNYWLQNYAKDS 441
Query: 643 ANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNG 464
AN V+ LGCG V+ T GQ +YPL +IR RMQ + + +G L G
Sbjct: 442 ANPG---VLVLLGCGTVSSTCGQLASYPLALIRTRMQ--------AQASIEGAPQLNMGG 490
Query: 463 MIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGV 320
+ FRK V EG LY G+ PN +KV+P+++I++V YE +K LG+
Sbjct: 491 L---FRKIVAKEGFFGLYTGIAPNFLKVLPAVSISYVVYEKMKIQLGI 535
Score = 43.5 bits (101), Expect = 0.007
Identities = 27/95 (28%), Positives = 45/95 (47%)
Frame = -1
Query: 601 GAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGV 422
G +AG + +T PLD ++ MQ+ G +I+TG ++ V+ GV
Sbjct: 263 GGMAGAVSRTGTAPLDRLKVMMQVHGTKGNSNIITG--------------LKQMVKEGGV 308
Query: 421 GALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
+L++G N +K+ P A+ F YE K + E
Sbjct: 309 RSLWRGNGVNVIKIAPETAMKFWAYEQYKKLFTSE 343
>sptrnew|AAH55369|AAH55369 Calcium-binding transporter.
Length = 475
Score = 184 bits (468), Expect = 2e-45
Identities = 104/290 (35%), Positives = 166/290 (57%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+++QV S+ G G + + + G+R L++GNGTN +I P +AVKF++Y
Sbjct: 214 PLDRLKVMMQVHGSKSMNIFG---GFRQMVKEGGIRSLWRGNGTNVIKIAPETAVKFWAY 270
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ Y++ EE +L R +G+ AG A + YPM++++ R+ V
Sbjct: 271 EQ--------YKKLLTEEGQKLGTFERFISGSMAGATAQTFIYPMEVLKTRLAVAKTG-- 320
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY G++ + + EGF A YKG++P+++G++PY G++ AVYE LK + L +
Sbjct: 321 -QYSGIYGCAKKILKHEGFGAFYKGYIPNLLGIIPYAGIDLAVYELLKSYWLDNFAKDSV 379
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
N V+ L CGA++ T GQ +YPL ++R RMQ + T +G L M
Sbjct: 380 NPG---VMVLLSCGALSSTCGQLASYPLALVRTRMQ--------AQATVEGAPQL---SM 425
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
+ F++ V EGV LY+G+ PN +KV+P++ I++V YE +K LGV +
Sbjct: 426 VGLFQRIVSKEGVSGLYRGITPNFMKVLPAVGISYVVYENMKQTLGVAQK 475
Score = 45.1 bits (105), Expect = 0.002
Identities = 29/99 (29%), Positives = 48/99 (48%)
Frame = -1
Query: 613 RLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVR 434
+L G VAG + +T PLD ++ MQ+ G +++ G FR+ V+
Sbjct: 197 QLLAGGVAGAVSRTSTAPLDRLKVMMQV------------HGSKSMNIFG---GFRQMVK 241
Query: 433 HEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
G+ +L++G N +K+ P A+ F YE K +L E
Sbjct: 242 EGGIRSLWRGNGTNVIKIAPETAVKFWAYEQYKKLLTEE 280
>sptr|Q8BMD8|Q8BMD8 Peroxisomal CA-dependent solute carrier homolog.
Length = 475
Score = 184 bits (468), Expect = 2e-45
Identities = 104/290 (35%), Positives = 166/290 (57%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+++QV S+ G G + + + G+R L++GNGTN +I P +AVKF++Y
Sbjct: 214 PLDRLKVMMQVHGSKSMNIFG---GFRQMVKEGGIRSLWRGNGTNVIKIAPETAVKFWAY 270
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ Y++ EE +L R +G+ AG A + YPM++++ R+ V
Sbjct: 271 EQ--------YKKLLTEEGQKLGTFERFISGSMAGATAQTFIYPMEVLKTRLAVAKTG-- 320
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY G++ + + EGF A YKG++P+++G++PY G++ AVYE LK + L +
Sbjct: 321 -QYSGIYGCAKKILKHEGFGAFYKGYIPNLLGIIPYAGIDLAVYELLKSYWLDNFAKDSV 379
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
N V+ L CGA++ T GQ +YPL ++R RMQ + T +G L M
Sbjct: 380 NPG---VMVLLSCGALSSTCGQLASYPLALVRTRMQ--------AQATVEGAPQL---SM 425
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
+ F++ V EGV LY+G+ PN +KV+P++ I++V YE +K LGV +
Sbjct: 426 VGLFQRIVSKEGVSGLYRGITPNFMKVLPAVGISYVVYENMKQTLGVAQK 475
Score = 45.1 bits (105), Expect = 0.002
Identities = 29/99 (29%), Positives = 48/99 (48%)
Frame = -1
Query: 613 RLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVR 434
+L G VAG + +T PLD ++ MQ+ G +++ G FR+ V+
Sbjct: 197 QLLAGGVAGAVSRTSTAPLDRLKVMMQV------------HGSKSMNIFG---GFRQMVK 241
Query: 433 HEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
G+ +L++G N +K+ P A+ F YE K +L E
Sbjct: 242 EGGIRSLWRGNGTNVIKIAPETAVKFWAYEQYKKLLTEE 280
>sptr|Q96NQ4|Q96NQ4 Hypothetical protein FLJ30339.
Length = 384
Score = 184 bits (467), Expect = 2e-45
Identities = 108/308 (35%), Positives = 172/308 (55%)
Frame = -1
Query: 1234 KSLXXXXXXXXXXXXXXAPLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGN 1055
K L APL+RLK+ +QV + + N + GL+ + G+R L++GN
Sbjct: 103 KQLVAGAVAGAVSRTGTAPLDRLKVFMQVHASKTNRLN-ILGGLRSMVLEGGIRSLWRGN 161
Query: 1054 GTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSAT 875
G N +I P SA+KF +YEQ + IL G+++ L R AG+ AG A +
Sbjct: 162 GINVLKIAPESAIKFMAYEQIKRAIL-------GQQET-LHVQERFVAGSLAGATAQTII 213
Query: 874 YPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFA 695
YPM++++ R+T++ QY+G+ + EG RA Y+G+LP+V+G++PY G++ A
Sbjct: 214 YPMEVLKTRLTLRRTG---QYKGLLDCARRILEREGPRAFYRGYLPNVLGIIPYAGIDLA 270
Query: 694 VYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSH 515
VYE+LK+W LQ S A+ ++ L CG ++ T GQ +YPL ++R RMQ
Sbjct: 271 VYETLKNWWLQQYSHDSADPG---ILVLLACGTISSTCGQIASYPLALVRTRMQ------ 321
Query: 514 ADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVK 335
A + + G G + M+ R + EG+ LY+G+ PN +KV+P+++I++V YE +K
Sbjct: 322 AQASIEG-GPQL----SMLGLLRHILSQEGMRGLYRGIAPNFMKVIPAVSISYVVYENMK 376
Query: 334 DVLGVEMR 311
LGV R
Sbjct: 377 QALGVTSR 384
>sptr|O18757|O18757 Peroxisomal Ca-dependent solute carrier.
Length = 475
Score = 182 bits (462), Expect = 9e-45
Identities = 105/290 (36%), Positives = 168/290 (57%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+++QV S+ G G + + + G+R L++GNGTN +I P +AVKF+ Y
Sbjct: 214 PLDRLKVMMQVHGSKSMNIFG---GFRQMIKEGGVRSLWRGNGTNVIKIAPETAVKFWVY 270
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ Y++ EE ++ R +G+ AG A + YPM++++ R+ V
Sbjct: 271 EQ--------YKKLLTEEGQKIGTFERFISGSMAGATAQTFIYPMEVMKTRLAVGKTG-- 320
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY G++ + + EGF A YKG++P+++G++PY G++ AVYE LK L +
Sbjct: 321 -QYSGIYDCAKKILKYEGFGAFYKGYVPNLLGIIPYAGIDLAVYELLKSHWLDNFAKDSV 379
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
N V+ LGCGA++ T GQ +YPL ++R RMQ A +++ G A Q N M
Sbjct: 380 NPG---VLVLLGCGALSSTCGQLASYPLALVRTRMQ------AQAMLEG----APQLN-M 425
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
+ FR+ + EG+ LY+G+ PN +KV+P++ I++V YE +K LGV +
Sbjct: 426 VGLFRRIISKEGLPGLYRGITPNFMKVLPAVGISYVVYENMKQTLGVTQK 475
Score = 44.7 bits (104), Expect = 0.003
Identities = 28/99 (28%), Positives = 48/99 (48%)
Frame = -1
Query: 613 RLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVR 434
+L G +AG + +T PLD ++ MQ+ G +++ G FR+ ++
Sbjct: 197 QLLAGGIAGAVSRTSTAPLDRLKVMMQV------------HGSKSMNIFG---GFRQMIK 241
Query: 433 HEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
GV +L++G N +K+ P A+ F YE K +L E
Sbjct: 242 EGGVRSLWRGNGTNVIKIAPETAVKFWVYEQYKKLLTEE 280
>sptr|Q9FLS8|Q9FLS8 Peroxisomal Ca-dependent solute carrier-like
protein (Putative peroxisomal Ca-dependent solute carrier
protein).
Length = 478
Score = 181 bits (459), Expect = 2e-44
Identities = 111/288 (38%), Positives = 160/288 (55%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+ LQVQ + G V +K IWR + L G F+GNG N A++ P SA+KF +Y
Sbjct: 224 PLDRLKVALQVQRTNL----GVVPTIKKIWREDKLLGFFRGNGLNVAKVAPESAIKFAAY 279
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
E I G D + RL AG AG +A +A YPMD+V+ R+ QT S
Sbjct: 280 EMLKPII--------GGADGDIGTSGRLLAGGLAGAVAQTAIYPMDLVKTRL--QTFVSE 329
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
++ ++ +EG RA Y+G PS+IG++PY G++ A YE+LKD + + L
Sbjct: 330 VGTPKLWKLTKDIWIQEGPRAFYRGLCPSLIGIIPYAGIDLAAYETLKD---LSRAHFLH 386
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
+ E + +LGCG +G +G + YPL VIR RMQ ADS T G+E
Sbjct: 387 DTAEPGPLIQLGCGMTSGALGASCVYPLQVIRTRMQ------ADSSKTSMGQE------- 433
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
F KT+R EG+ Y+G+ PN KV+PS +I+++ YE +K L ++
Sbjct: 434 ---FLKTLRGEGLKGFYRGIFPNFFKVIPSASISYLVYEAMKKNLALD 478
Score = 38.9 bits (89), Expect = 0.16
Identities = 23/93 (24%), Positives = 42/93 (45%)
Frame = -1
Query: 601 GAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGV 422
G +AG + +T PLD ++ +Q+ + G++ +K R + +
Sbjct: 211 GGIAGAVSRTATAPLDRLKVALQVQRTN----------------LGVVPTIKKIWREDKL 254
Query: 421 GALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLG 323
++G N KV P AI F YE++K ++G
Sbjct: 255 LGFFRGNGLNVAKVAPESAIKFAAYEMLKPIIG 287
>sptr|Q96PZ1|Q96PZ1 Hypothetical protein KIAA1896 (Fragment).
Length = 568
Score = 179 bits (455), Expect = 6e-44
Identities = 108/292 (36%), Positives = 165/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYN--GTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N G V G + R G R L++GNG N +I P SA+KF
Sbjct: 305 PLDRLKVLMQV---HASRSNNMGIVGGFTQMIREGGARSLWRGNGINVLKIAPESAIKFM 361
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 362 AYEQIKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 410
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 411 KTGQYSGMLDCARRILAREGVAAFYKGYVPNMLGIIPYAGIDLAVYETLKNAWLQHYA-- 468
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 469 -VNSADPGVFVLLACGTMSSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 517
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F+ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 518 -MSSLFKHILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 568
>sptr|Q9BSA6|Q9BSA6 Hypothetical protein (Fragment).
Length = 311
Score = 179 bits (455), Expect = 6e-44
Identities = 108/292 (36%), Positives = 165/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYN--GTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N G V G + R G R L++GNG N +I P SA+KF
Sbjct: 48 PLDRLKVLMQV---HASRSNNMGIVGGFTQMIREGGARSLWRGNGINVLKIAPESAIKFM 104
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 105 AYEQIKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 153
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 154 KTGQYSGMLDCARRILAREGVAAFYKGYVPNMLGIIPYAGIDLAVYETLKNAWLQHYA-- 211
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 212 -VNSADPGVFVLLACGTMSSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 260
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F+ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 261 -MSSLFKHILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 311
>sptr|Q8BHG0|Q8BHG0 Weakly similar to peroxisomal CA-dependent solute
carrier.
Length = 502
Score = 179 bits (454), Expect = 8e-44
Identities = 106/292 (36%), Positives = 166/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGT--VQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N V G + R G + L++GNG N +I P SA+KF
Sbjct: 239 PLDRLKVLMQV---HASRSNNMCIVGGFTQMIREGGAKSLWRGNGINVLKIAPESAIKFM 295
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 296 AYEQMKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 344
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 345 KTGQYSGMLDCARRILAKEGVAAFYKGYIPNMLGIIPYAGIDLAVYETLKNTWLQRYA-- 402
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 403 -VNSADPGVFVLLACGTISSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 451
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F++ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 452 -MSSLFKQILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 502
>sptr|Q80T78|Q80T78 MKIAA1896 protein (Fragment).
Length = 515
Score = 179 bits (454), Expect = 8e-44
Identities = 106/292 (36%), Positives = 166/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGT--VQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N V G + R G + L++GNG N +I P SA+KF
Sbjct: 252 PLDRLKVLMQV---HASRSNNMCIVGGFTQMIREGGAKSLWRGNGINVLKIAPESAIKFM 308
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 309 AYEQMKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 357
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 358 KTGQYSGMLDCARRILAKEGVAAFYKGYIPNMLGIIPYAGIDLAVYETLKNTWLQRYA-- 415
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 416 -VNSADPGVFVLLACGTISSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 464
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F++ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 465 -MSSLFKQILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 515
>sptr|Q8VBT4|Q8VBT4 Hypothetical 40.6 kDa protein (Similar to KIAA1896
protein).
Length = 366
Score = 179 bits (454), Expect = 8e-44
Identities = 106/292 (36%), Positives = 166/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGT--VQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N V G + R G + L++GNG N +I P SA+KF
Sbjct: 103 PLDRLKVLMQV---HASRSNNMCIVGGFTQMIREGGAKSLWRGNGINVLKIAPESAIKFM 159
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 160 AYEQMKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 208
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 209 KTGQYSGMLDCARRILAKEGVAAFYKGYIPNMLGIIPYAGIDLAVYETLKNTWLQRYA-- 266
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 267 -VNSADPGVFVLLACGTISSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 315
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F++ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 316 -MSSLFKQILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 366
>sptr|Q8JZT8|Q8JZT8 Similar to peroxisomal Ca-dependent solute
carrier-like protein.
Length = 514
Score = 179 bits (454), Expect = 8e-44
Identities = 106/292 (36%), Positives = 166/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGT--VQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N V G + R G + L++GNG N +I P SA+KF
Sbjct: 251 PLDRLKVLMQV---HASRSNNMCIVGGFTQMIREGGAKSLWRGNGINVLKIAPESAIKFM 307
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 308 AYEQMKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 356
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 357 KTGQYSGMLDCARRILAKEGVAAFYKGYIPNMLGIIPYAGIDLAVYETLKNTWLQRYA-- 414
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 415 -VNSADPGVFVLLACGTISSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 463
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F++ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 464 -MSSLFKQILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 514
>sptr|Q8K3P6|Q8K3P6 Peroxisomal Ca-dependent solute carrier-like
protein.
Length = 469
Score = 179 bits (453), Expect = 1e-43
Identities = 105/292 (35%), Positives = 166/292 (56%), Gaps = 2/292 (0%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGT--VQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFF 1007
PL+RLK+L+QV H+ + N + G + R G + L++GNG N +I P SA+KF
Sbjct: 206 PLDRLKVLMQV---HASRSNNMCIIGGFTQMIREGGAKSLWRGNGINVLKIAPESAIKFM 262
Query: 1006 SYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
+YEQ + + G + L RL AG+ AG IA S+ YPM++++ R+ + +
Sbjct: 263 AYEQMKRLV--------GSDQETLRIHERLVAGSLAGAIAQSSIYPMEVLKTRMAL---R 311
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFG 647
QY GM + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ +
Sbjct: 312 KTGQYSGMLDCAKRILAKEGVAAFYKGYIPNMLGIIPYAGIDLAVYETLKNTWLQRYA-- 369
Query: 646 LANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN 467
N + V L CG ++ T GQ +YPL ++R RMQ A + + G +
Sbjct: 370 -VNSADPGVFVLLACGTISSTCGQLASYPLALVRTRMQ------AQASIEGAPEVT---- 418
Query: 466 GMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
M F++ +R EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 419 -MSSLFKQILRTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKITLGVQSR 469
>sptr|Q7ZYD5|Q7ZYD5 Similar to hypothetical protein MGC36388.
Length = 514
Score = 178 bits (451), Expect = 2e-43
Identities = 106/290 (36%), Positives = 169/290 (58%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+L+QV S + + G ++ R G R L++GNG N +I P SA+KF +Y
Sbjct: 251 PLDRLKVLMQVHASRSNNMS-ILGGFTHMIREGGFRSLWRGNGINVIKIAPESAIKFMAY 309
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ K I+ + ++ G + R AG+ AG+IA S+ YPM++++ R+ + +
Sbjct: 310 EQI-KRIIGSNQETLGIHE-------RFVAGSLAGVIAQSSIYPMEVLKTRMAL---RKT 358
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY+G+ + +EG A YKG++P+++G++PY G++ AVYE+LK+ LQ + A
Sbjct: 359 GQYQGVLDCGKKILLQEGLSAFYKGYVPNMLGIIPYAGIDLAVYETLKNAWLQRYATSSA 418
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
+ V L CG V+ T GQ +YPL ++R RMQ A++ V G + M
Sbjct: 419 DPG---VFVLLACGTVSSTCGQLASYPLALVRTRMQ------AEASVEGAPQMT-----M 464
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
F+ V+ EG LY+GL PN +KV+P+++I++V YE +K LGV+ R
Sbjct: 465 SKLFKHIVKTEGAFGLYRGLAPNFMKVIPAVSISYVVYENLKLTLGVQSR 514
Score = 90.5 bits (223), Expect = 5e-17
Identities = 61/226 (26%), Positives = 104/226 (46%)
Frame = -1
Query: 1009 FSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTD 830
F+ E+ G+ W + L AG AG ++ + T P+D R ++ +Q
Sbjct: 221 FTVEEKQTGMWWRH----------------LVAGGGAGAVSRTCTAPLD--RLKVLMQVH 262
Query: 829 KSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSF 650
S + + RE GFR+L++G +VI + P + F YE +K +
Sbjct: 263 ASRSNNMSILGGFTHMIREGGFRSLWRGNGINVIKIAPESAIKFMAYEQIKRII------ 316
Query: 649 GLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQY 470
+N L + R G++AG I Q+ YP++V++ RM + ++ QY
Sbjct: 317 -GSNQETLGIHERFVAGSLAGVIAQSSIYPMEVLKTRMAL--------------RKTGQY 361
Query: 469 NGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKD 332
G++D +K + EG+ A YKG VPN + ++P I YE +K+
Sbjct: 362 QGVLDCGKKILLQEGLSAFYKGYVPNMLGIIPYAGIDLAVYETLKN 407
>sptr|Q9SZI9|Q9SZI9 Putative mitochondrial carrier protein.
Length = 325
Score = 177 bits (450), Expect = 2e-43
Identities = 110/295 (37%), Positives = 161/295 (54%), Gaps = 5/295 (1%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PLER+KIL Q + K G V + I +TEGL G ++GNG + ARIVP +A+ + +Y
Sbjct: 37 PLERIKILFQTRRDE-FKRIGLVGSINKIGKTEGLMGFYRGNGASVARIVPYAALHYMAY 95
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
E+ + I++ + D PLL L AG+ AG A+ TYP+D+VR ++ QT
Sbjct: 96 EEYRRWIIFGF------PDTTRGPLLDLVAGSFAGGTAVLFTYPLDLVRTKLAYQTQVKA 149
Query: 820 YQ-----YRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTN 656
YRG+ YRE G R LY+G PS+ G+ PY GL F YE +K
Sbjct: 150 IPVEQIIYRGIVDCFSRTYRESGARGLYRGVAPSLYGIFPYAGLKFYFYEEMKR------ 203
Query: 655 SFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEAL 476
+ +++ + +L CG+VAG +GQT+ YPLDV+RR+MQ+ + + + +E
Sbjct: 204 --HVPPEHKQDISLKLVCGSVAGLLGQTLTYPLDVVRRQMQV------ERLYSAVKEETR 255
Query: 475 QYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEMR 311
+ G + K R EG L+ GL N +KVVPS+AI F Y+++K L V R
Sbjct: 256 R--GTMQTLFKIAREEGWKQLFSGLSINYLKVVPSVAIGFTVYDIMKLHLRVPPR 308
>sptr|Q9LY28|Q9LY28 Peroxisomal Ca-dependent solute carrier-like
protein.
Length = 479
Score = 177 bits (450), Expect = 2e-43
Identities = 108/291 (37%), Positives = 161/291 (55%), Gaps = 3/291 (1%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK++LQVQ H+ G + +K IWR + L G F+GNG N ++ P SA+KF +Y
Sbjct: 225 PLDRLKVVLQVQRAHA----GVLPTIKKIWREDKLMGFFRGNGLNVMKVAPESAIKFCAY 280
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITV---QTD 830
E I G ED + RL AG AG +A +A YPMD+V+ R+ +
Sbjct: 281 EMLKPMI--------GGEDGDIGTSGRLMAGGMAGALAQTAIYPMDLVKTRLQTCVSEGG 332
Query: 829 KSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSF 650
K+P ++ ++ EG RA YKG PS++G+VPY G++ A YE+LKD L +T
Sbjct: 333 KAPKLWK----LTKDIWVREGPRAFYKGLFPSLLGIVPYAGIDLAAYETLKD-LSRTY-- 385
Query: 649 GLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQY 470
+ D E + +L CG +G +G + YPL V+R RMQ ADS T +E
Sbjct: 386 -ILQDTEPGPLIQLSCGMTSGALGASCVYPLQVVRTRMQ------ADSSKTTMKQE---- 434
Query: 469 NGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVE 317
F T++ EG+ Y+GL+PN +KVVP+ +I ++ YE +K + ++
Sbjct: 435 ------FMNTMKGEGLRGFYRGLLPNLLKVVPAASITYIVYEAMKKNMALD 479
>sptrnew|EAA11419|EAA11419 ENSANGP00000009995 (Fragment).
Length = 524
Score = 177 bits (449), Expect = 3e-43
Identities = 100/292 (34%), Positives = 166/292 (56%), Gaps = 3/292 (1%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+RLK+ LQVQ L+Y+ + G+R L++GN N +I P SA+KF +Y
Sbjct: 258 PLDRLKVFLQVQ----ASKQRISDCLQYMLKEGGVRSLWRGNFINVLKIAPESAIKFAAY 313
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ + I G + Q+T R AGACAG ++ +A YPM++++ R+ + +
Sbjct: 314 EQVKRLI-------RGNDKRQMTIYERFVAGACAGGVSQTAIYPMEVLKTRLAL---RKT 363
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
+Y + A +YR EG R+ Y+G++P+++G++PY G++ AVYE+LK L +
Sbjct: 364 GEYSSILDAASKIYRREGLRSFYRGYIPNMLGIIPYAGIDLAVYETLKKKYLSHH----- 418
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTG---QGKEALQY 470
+ L CG+ + T+GQ +YPL ++R R+Q A ++ G G A++
Sbjct: 419 ETEQPSFWLLLACGSASSTLGQVCSYPLALVRTRLQ------AQAVTIGPNPDGSVAVEP 472
Query: 469 NGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEM 314
N M + F++ ++ EG LY+G+ PN +KV+P+++I++V YE LGV M
Sbjct: 473 N-MTNVFKRIIQTEGPVGLYRGITPNFIKVLPAVSISYVVYEYTSRALGVNM 523
Score = 39.7 bits (91), Expect = 0.095
Identities = 27/95 (28%), Positives = 44/95 (46%)
Frame = -1
Query: 610 LGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRH 431
L G +AG + +T PLD ++ +Q+ S + LQY ++
Sbjct: 242 LAAGGIAGAVSRTCTAPLDRLKVFLQVQASKQRIS-------DCLQY---------MLKE 285
Query: 430 EGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVL 326
GV +L++G N +K+ P AI F YE VK ++
Sbjct: 286 GGVRSLWRGNFINVLKIAPESAIKFAAYEQVKRLI 320
>sw|Q20799|CMC2_CAEEL Probable calcium-binding mitochondrial carrier
F55A11.4.
Length = 588
Score = 171 bits (434), Expect = 2e-41
Identities = 95/287 (33%), Positives = 168/287 (58%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
P +R+K+ LQV + + + G + LK + G++ ++GNG N +I P SA+KF Y
Sbjct: 268 PFDRIKVYLQVNSSKTNRL-GVMSCLKLLHAEGGIKSFWRGNGINVIKIAPESAIKFMCY 326
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
+Q + I +++ G E+ ++ RL AG+ AG I+ S YPM++++ R+ ++ K+
Sbjct: 327 DQLKRLI----QKKKGNEE--ISTFERLCAGSAAGAISQSTIYPMEVMKTRLALR--KTG 378
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
RG+ H +Y +EG R YKG+LP++IG++PY G++ A+YE+LK ++ +
Sbjct: 379 QLDRGIIHFAHKMYTKEGIRCFYKGYLPNLIGIIPYAGIDLAIYETLKRTYVR---YYET 435
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNGM 461
N +E V+ L CG + T GQ +YP ++R R+Q + + + + Q + M
Sbjct: 436 NSSEPGVLALLACGTCSSTCGQLSSYPFALVRTRLQALSIT----------RYSPQPDTM 485
Query: 460 IDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGV 320
F+ +++EGV Y+G+ PN +KV+P+++I++V YE V+ LGV
Sbjct: 486 FGQFKYILQNEGVTGFYRGITPNFLKVIPAVSISYVVYEKVRTGLGV 532
Score = 33.5 bits (75), Expect = 6.8
Identities = 15/53 (28%), Positives = 28/53 (52%)
Frame = -1
Query: 1102 KYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEED 944
KYI + EG+ G ++G N +++P ++ + YE+ G+ + G ED
Sbjct: 490 KYILQNEGVTGFYRGITPNFLKVIPAVSISYVVYEKVRTGLGVPVCSRGGLED 542
>sptr|Q9VTX4|Q9VTX4 CG32103 protein.
Length = 583
Score = 170 bits (430), Expect = 5e-41
Identities = 96/298 (32%), Positives = 167/298 (56%), Gaps = 9/298 (3%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+R+K+ LQVQ G + + + G R +++GNG N +I P +A KF +Y
Sbjct: 305 PLDRIKVYLQVQTQRM----GISECMHIMLNEGGSRSMWRGNGINVLKIAPETAFKFAAY 360
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ + I + + Q++ + R AGA AG I+ + YPM++++ R+ ++
Sbjct: 361 EQMKRLI------RGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTG-- 412
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY G+ A +Y++EG R+ Y+G++P+++G++PY G++ AVYE+LK + +
Sbjct: 413 -QYAGIADAAVKIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHD---- 467
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNG- 464
N+ + + L CG+ + T+GQ +YPL ++R R+Q A++I + K +
Sbjct: 468 NNEQPSFLVLLACGSTSSTLGQLCSYPLALVRTRLQA---QAAETIANQKRKTQIPLKSS 524
Query: 463 --------MIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEM 314
M FRK VR EG+ LY+G+ PN +KV+P+++I++V YE LG++M
Sbjct: 525 DAHSGEETMTGLFRKIVRQEGLTGLYRGITPNFLKVLPAVSISYVVYEYTSRALGIKM 582
Score = 92.4 bits (228), Expect = 1e-17
Identities = 57/195 (29%), Positives = 95/195 (48%)
Frame = -1
Query: 919 LGAGACAGIIAMSATYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWL 740
L AG AG ++ + T P+D ++ + VQT Q G+ + + E G R++++G
Sbjct: 289 LVAGGIAGAVSRTCTAPLDRIKVYLQVQT-----QRMGISECMHIMLNEGGSRSMWRGNG 343
Query: 739 PSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYP 560
+V+ + P FA YE +K + G ++ +V R GA AG I QT+ YP
Sbjct: 344 INVLKIAPETAFKFAAYEQMKRLIR-----GDDGSRQMSIVERFYAGAAAGGISQTIIYP 398
Query: 559 LDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKV 380
++V++ R+ + + QY G+ DA K + EGV + Y+G VPN + +
Sbjct: 399 MEVLKTRLAL--------------RRTGQYAGIADAAVKIYKQEGVRSFYRGYVPNILGI 444
Query: 379 VPSIAIAFVTYEVVK 335
+P I YE +K
Sbjct: 445 LPYAGIDLAVYETLK 459
>sptr|Q9VTX3|Q9VTX3 CG32103 protein (RE56970p).
Length = 363
Score = 170 bits (430), Expect = 5e-41
Identities = 96/298 (32%), Positives = 167/298 (56%), Gaps = 9/298 (3%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
PL+R+K+ LQVQ G + + + G R +++GNG N +I P +A KF +Y
Sbjct: 85 PLDRIKVYLQVQTQRM----GISECMHIMLNEGGSRSMWRGNGINVLKIAPETAFKFAAY 140
Query: 1000 EQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTDKSP 821
EQ + I + + Q++ + R AGA AG I+ + YPM++++ R+ ++
Sbjct: 141 EQMKRLI------RGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTG-- 192
Query: 820 YQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLA 641
QY G+ A +Y++EG R+ Y+G++P+++G++PY G++ AVYE+LK + +
Sbjct: 193 -QYAGIADAAVKIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHD---- 247
Query: 640 NDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYNG- 464
N+ + + L CG+ + T+GQ +YPL ++R R+Q A++I + K +
Sbjct: 248 NNEQPSFLVLLACGSTSSTLGQLCSYPLALVRTRLQA---QAAETIANQKRKTQIPLKSS 304
Query: 463 --------MIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVLGVEM 314
M FRK VR EG+ LY+G+ PN +KV+P+++I++V YE LG++M
Sbjct: 305 DAHSGEETMTGLFRKIVRQEGLTGLYRGITPNFLKVLPAVSISYVVYEYTSRALGIKM 362
Score = 92.4 bits (228), Expect = 1e-17
Identities = 57/195 (29%), Positives = 95/195 (48%)
Frame = -1
Query: 919 LGAGACAGIIAMSATYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWL 740
L AG AG ++ + T P+D ++ + VQT Q G+ + + E G R++++G
Sbjct: 69 LVAGGIAGAVSRTCTAPLDRIKVYLQVQT-----QRMGISECMHIMLNEGGSRSMWRGNG 123
Query: 739 PSVIGVVPYVGLNFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYP 560
+V+ + P FA YE +K + G ++ +V R GA AG I QT+ YP
Sbjct: 124 INVLKIAPETAFKFAAYEQMKRLIR-----GDDGSRQMSIVERFYAGAAAGGISQTIIYP 178
Query: 559 LDVIRRRMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKV 380
++V++ R+ + + QY G+ DA K + EGV + Y+G VPN + +
Sbjct: 179 MEVLKTRLAL--------------RRTGQYAGIADAAVKIYKQEGVRSFYRGYVPNILGI 224
Query: 379 VPSIAIAFVTYEVVK 335
+P I YE +K
Sbjct: 225 LPYAGIDLAVYETLK 239
>sptr|Q8R225|Q8R225 Hypothetical 25.4 kDa protein (Fragment).
Length = 232
Score = 170 bits (430), Expect = 5e-41
Identities = 94/257 (36%), Positives = 148/257 (57%)
Frame = -1
Query: 1081 GLRGLFKGNGTNCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGAC 902
G+R L++GNGTN +I P +AVKF++YEQ Y++ EE +L R +G+
Sbjct: 1 GIRSLWRGNGTNVIKIAPETAVKFWAYEQ--------YKKLLTEEGQKLGTFERFISGSM 52
Query: 901 AGIIAMSATYPMDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGV 722
AG A + YPM++++ R+ V QY G++ + + EGF A YKG++P+++G+
Sbjct: 53 AGATAQTFIYPMEVLKTRLAVAKTG---QYSGIYGCAKKILKHEGFGAFYKGYIPNLLGI 109
Query: 721 VPYVGLNFAVYESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRR 542
+PY G++ AVYE LK + L + N V+ L CGA++ T GQ +YPL ++R
Sbjct: 110 IPYAGIDLAVYELLKSYWLDNFAKDSVNPG---VMVLLSCGALSSTCGQLASYPLALVRT 166
Query: 541 RMQMVGWSHADSIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAI 362
RMQ + T +G L M+ F++ V EGV LY+G+ PN +KV+P++ I
Sbjct: 167 RMQ--------AQATVEGAPQL---SMVGLFQRIVSKEGVSGLYRGITPNFMKVLPAVGI 215
Query: 361 AFVTYEVVKDVLGVEMR 311
++V YE +K LGV +
Sbjct: 216 SYVVYENMKQTLGVAQK 232
Score = 65.1 bits (157), Expect = 2e-09
Identities = 49/173 (28%), Positives = 80/173 (46%), Gaps = 2/173 (1%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVKFFSY 1001
P+E LK L V +Y+G K I + EG +KG N I+P + + Y
Sbjct: 63 PMEVLKTRLAVAKTG--QYSGIYGCAKKILKHEGFGAFYKGYIPNLLGIIPYAGIDLAVY 120
Query: 1000 EQASKGILWAYRQQTGEEDAQLTP--LLRLGAGACAGIIAMSATYPMDMVRGRITVQTDK 827
E +L +Y +D+ + P ++ L GA + A+YP+ +VR R+ Q
Sbjct: 121 E-----LLKSYWLDNFAKDS-VNPGVMVLLSCGALSSTCGQLASYPLALVRTRMQAQATV 174
Query: 826 SPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWL 668
M + +EG LY+G P+ + V+P VG+++ VYE++K L
Sbjct: 175 EGAPQLSMVGLFQRIVSKEGVSGLYRGITPNFMKVLPAVGISYVVYENMKQTL 227
>sw|Q19529|CMC3_CAEEL Probable calcium-binding mitochondrial carrier
F17E5.2.
Length = 565
Score = 169 bits (428), Expect = 8e-41
Identities = 101/305 (33%), Positives = 166/305 (54%), Gaps = 16/305 (5%)
Frame = -1
Query: 1180 PLERLKILLQVQNPH--------------SIKYN--GTVQGLKYIWRTEGLRGLFKGNGT 1049
P +R+K+ LQV H S K N G V + + G++ ++GNG
Sbjct: 281 PFDRIKVYLQVMYLHLLFHIMFLKALQVNSTKTNKLGVVSCVHLLHAEGGIKSFWRGNGI 340
Query: 1048 NCARIVPNSAVKFFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYP 869
N +I P SA+KF Y+Q + W + G A+L+ + RL AG+ AG I+ +A YP
Sbjct: 341 NVIKIAPESAMKFMCYDQIKR---WMQEYKGG---AELSTIERLLAGSSAGAISQTAIYP 394
Query: 868 MDMVRGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVY 689
M++++ R+ ++ ++ +GMFH +Y +EG + YKG+LP+++G++PY G++ VY
Sbjct: 395 MEVMKTRLALR--RTGQLDKGMFHFAHKMYTKEGIKCFYKGYLPNLLGIIPYAGIDLTVY 452
Query: 688 ESLKDWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHAD 509
ESLK + T + E V+ L CG + T GQ +YPL ++R R+Q S
Sbjct: 453 ESLKS--MYTKYY--TEHTEPGVLALLACGTCSSTCGQLASYPLALVRTRLQARAIS--- 505
Query: 508 SIVTGQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDV 329
K + Q + M+ F+ ++ EG LY+G+ PN +KV+P+++I++V YE V+
Sbjct: 506 ------PKNSTQPDTMVGQFKHILQTEGFTGLYRGITPNFMKVIPAVSISYVVYEKVRKQ 559
Query: 328 LGVEM 314
LG M
Sbjct: 560 LGATM 564
Score = 33.1 bits (74), Expect = 8.9
Identities = 24/99 (24%), Positives = 45/99 (45%), Gaps = 7/99 (7%)
Frame = -1
Query: 610 LGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQYN-------GMIDA 452
L G VAG + +T P D I+ +Q++ + + +ALQ N G++
Sbjct: 265 LVAGGVAGAMSRTCTAPFDRIKVYLQVM---YLHLLFHIMFLKALQVNSTKTNKLGVVSC 321
Query: 451 FRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVK 335
G+ + ++G N +K+ P A+ F+ Y+ +K
Sbjct: 322 VHLLHAEGGIKSFWRGNGINVIKIAPESAMKFMCYDQIK 360
>sptr|Q9FU82|Q9FU82 Putative peroxisomal Ca-dependent solute carrier
protein.
Length = 322
Score = 168 bits (426), Expect = 1e-40
Identities = 102/289 (35%), Positives = 156/289 (53%), Gaps = 4/289 (1%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHS----IKYNGTVQGLKYIWRTEGLRGLFKGNGTNCARIVPNSAVK 1013
PL RL IL QV HS +K I R EG +KGN +P SA+
Sbjct: 45 PLARLTILFQVAGMHSDVAALKKYSIWHEASRIVREEGFGAFWKGNLVTIVHRLPYSAIS 104
Query: 1012 FFSYEQASKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQT 833
F+SYE+ K + R +ED+ + RL +G AGI A S TYP+D+VR R+ Q
Sbjct: 105 FYSYERYKK---FLQRVPGLDEDSNYVGVARLLSGGLAGITAASVTYPLDVVRTRLATQ- 160
Query: 832 DKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLKDWLLQTNS 653
K+ Y+G+FHA+ T+ R+EG + LYKG +++GV P + ++F VYESL+ +
Sbjct: 161 -KTTRYYKGIFHAVSTICRDEGVKGLYKGLGATLLGVGPSIAISFTVYESLR------SH 213
Query: 652 FGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVTGQGKEALQ 473
+ + + V L G+++G T +PLD+++RRMQ+ G G ++
Sbjct: 214 WQMERPQDSPAVVSLFSGSLSGIASSTATFPLDLVKRRMQLQG---------AAGTSSVC 264
Query: 472 YNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVL 326
+ + R+ + EG+ Y+G+VP +KVVPS+ IAF+TYE +K +L
Sbjct: 265 KSSITGTIRQIFQKEGLRGFYRGIVPEYLKVVPSVGIAFMTYETLKSLL 313
>sptr|Q8W4M2|Q8W4M2 Hypothetical 39.8 kDa protein.
Length = 365
Score = 168 bits (425), Expect = 2e-40
Identities = 101/297 (34%), Positives = 158/297 (53%), Gaps = 12/297 (4%)
Frame = -1
Query: 1180 PLERLKILLQVQNPHSIKYNGTVQGLKYIW-------RTEGLRGLFKGNGTNCARIVPNS 1022
PL RL IL Q+Q S + IW + EG R +KGN A +P
Sbjct: 89 PLARLTILFQIQGMQS---EAAILSSPNIWHEASRIVKEEGFRAFWKGNLVTVAHRLPYG 145
Query: 1021 AVKFFSYEQA-----SKGILWAYRQQTGEEDAQLTPLLRLGAGACAGIIAMSATYPMDMV 857
AV F++YE+ S +L +Y+ G + + + +G AG+ A SATYP+D+V
Sbjct: 146 AVNFYAYEEYKTFLHSNPVLQSYKGNAGVDIS-----VHFVSGGLAGLTAASATYPLDLV 200
Query: 856 RGRITVQTDKSPYQYRGMFHALGTVYREEGFRALYKGWLPSVIGVVPYVGLNFAVYESLK 677
R R++ Q ++ Y+G+ HA T+ REEG LYKG +++GV P + ++FA YE+ K
Sbjct: 201 RTRLSAQ--RNSIYYQGVGHAFRTICREEGILGLYKGLGATLLGVGPSLAISFAAYETFK 258
Query: 676 DWLLQTNSFGLANDNELHVVTRLGCGAVAGTIGQTVAYPLDVIRRRMQMVGWSHADSIVT 497
+ L N+ + V LGCG+++G + T +PLD++RRRMQ+ G
Sbjct: 259 TFWLS------HRPNDSNAVVSLGCGSLSGIVSSTATFPLDLVRRRMQLEG--------- 303
Query: 496 GQGKEALQYNGMIDAFRKTVRHEGVGALYKGLVPNSVKVVPSIAIAFVTYEVVKDVL 326
G+ + G+ F+ + EG+ LY+G++P KVVP + IAF+T+E +K +L
Sbjct: 304 AGGRARVYTTGLFGTFKHIFKTEGMRGLYRGIIPEYYKVVPGVGIAFMTFEELKKLL 360
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,005,994,181
Number of Sequences: 1272877
Number of extensions: 23087357
Number of successful extensions: 65171
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 55474
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 60622
length of database: 406,886,216
effective HSP length: 125
effective length of database: 247,776,591
effective search space used: 74580753891
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

